The perks of growing slowly
Published in Microbiology

The amount of time it takes a cell to undergo one round of division varies dramatically across the bacterial tree of life. Some bacteria have a need for speed, whereas others prefer to stop and smell the nutrient broth. When provided with optimal conditions for growth, Vibrio natriegens can divide once every 16 minutes. Methylomagnum ishizawai, on the other hand, takes 24 hours to complete a single round of division.
To be able to reproduce so quickly, fast growing bacteria maximize their translational speed. Rapid translation allows these bacteria to quickly turn mRNA into the proteins needed to sustain their growth. Functionally, this means that fast-growing bacteria can be sensitive to even slight hiccups in translational speed, especially in the highly expressed proteins most crucial for growth and replication.
Multiple factors can put the brakes on translational speed, including the specific amino acid being added to the polypeptide chain. Proline is the slowest amino acid to form peptide bonds due to its structural rigidity. This rigidity imparts proline with unique properties. Proline rich motifs are over-represented in the binding domains of signaling proteins1 and increased proline content is linked to increased thermostability2. However, sequential stretches of prolines can cause ribosomes to stall, slowing down translation3. Consequently, fast-growing bacteria like V. natriegens (Figure 1, left) and E. coli4 show signs of active selection against these proline motifs, meaning they encode fewer than would be expected by chance. Conversely, slow-growing bacteria like M. ishizawai (Figure 1, right) often encode more proline motifs than expected.
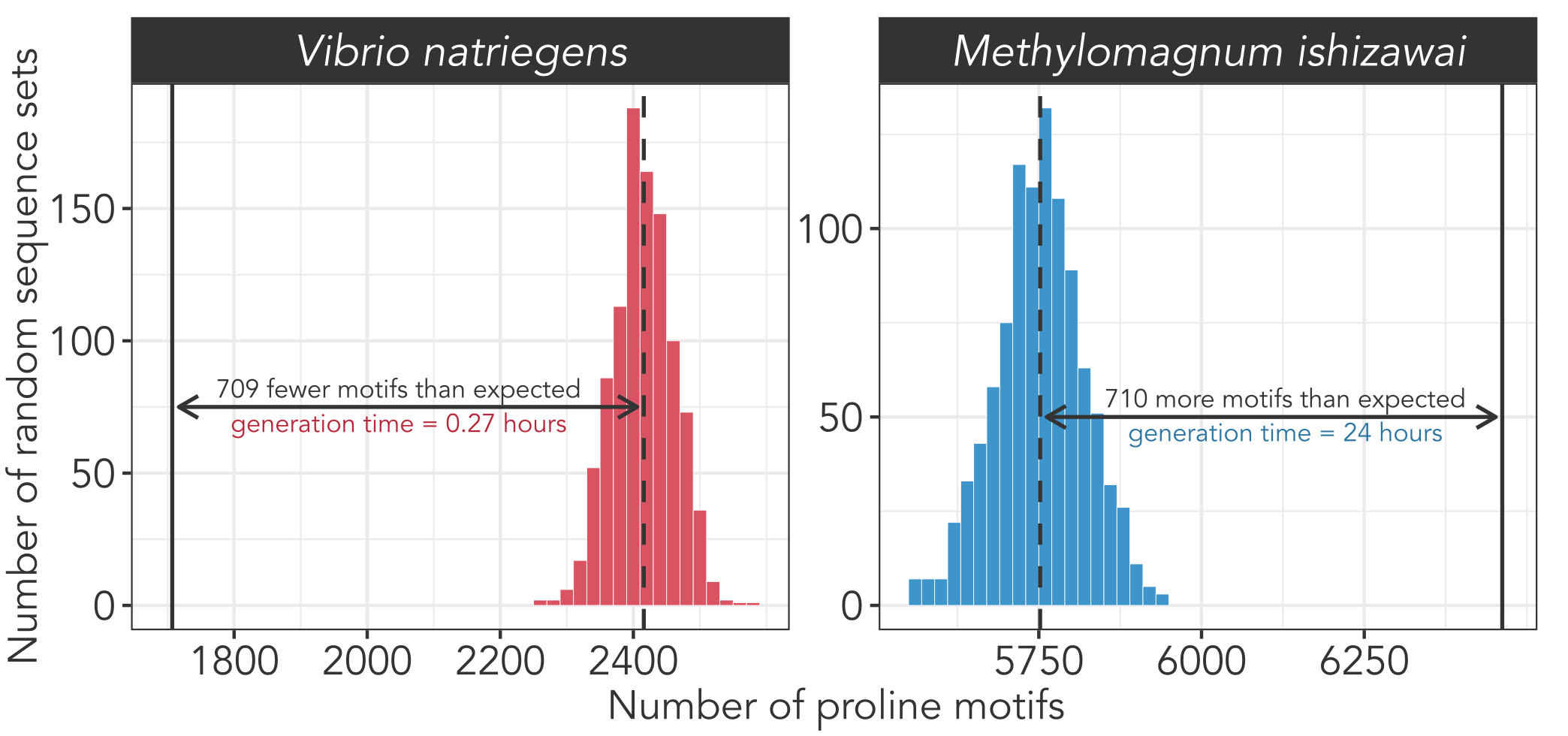
Figure 1. Fast-growing species (like V. natrigiens) tend to encode fewer proline motifs (XXPPX, where X is any amino acid) than expected by chance, while slow-growing species (like M. ishizawai) often encode more proline motifs than expected. The solid vertical line indicates the actual number of proline motifs encoded by each genome. The dashed vertical line indicates the average number of proline motifs that occur when protein sequences are shuffled 1000 times while preserving their overall amino acid composition. The histograms indicate the results of each round of shuffling.
I’m interested in how growth rates affect other aspects of bacterial existence, from genomes to lifestyles. If strong pressure on maximizing translational speed forces fast-growing bacteria to limit the number of proline motifs they encode, this could present an interesting tradeoff between rapid growth and the specific activity of proline motifs. After all, the biochemical activity conferred by proline motifs cannot always be replicated by other amino acids5.
To investigate the connection between growth rate and the abundance of proline motifs in bacterial genomes, I compiled a set of over 3000 genomes from 35 phyla. Using statistical tests which take shared phylogeny into account, I found that traits associated with slow growth were significantly correlated with increased amounts of proline motifs. This suggests that high pressure on maximizing translational speed does limit the inclusion of slow-to-translate proline sequences within the proteins of fast-growing species.
I also found two groups of bacteria which encoded significantly more proline motifs than their genomic characteristics would otherwise predict: thermophilic and multicellular bacteria. Both groups have reduced pressure on maximizing translational speed compared to fast-growing bacteria. Thermophiles are thought to benefit from enhanced enzymatic speeds purely because catalysis proceeds at higher rates at higher temperatures6, while multicellular bacteria are generally slow growing. Myxococcus xanthus, the prototypical multicellular species, takes 3.3 hours to undergo a single round of division.
These findings raise questions about whether strong pressure on growth rates, and therefore on maximizing translational speed may limit the evolutionary paths available to fast-growing bacteria. To me, this is especially interesting considering most microbiological studies tend to focus on a few fast-growing model organisms such as E. coli and B. subtilis. As a result, traditional microbiology is rooted in the physiology of fast-growing bacteria, despite their relative rarity in environmental habitats7. The increased abundance of proline motifs in the genomes of slow-growing bacteria may indicate that some paths are uniquely accessible to slow-growing organisms, and that slow-growth can be an evolutionary asset.
Check out the full paper here.
References
1) Mandal A, Mandal S, Park MH. Genome-wide analyses and functional classification of proline repeat-rich proteins: potential role of eIF5A in eukaryotic evolution. PLoS One 2014; 9: e111800-13.
2) Watanabe K, Suzuki Y. Protein thermostabilization by proline substitutions. J. Mol. Catal. B Enzym. 1998; 4: 167–180.
3) Woolstenhulme CJ, Guydosh NR, Green R, Buskirk AR. High-precision analysis of translational pausing by ribosome profiling in bacteria lacking EFP. Cell Rep. 2015; 11: 13–21.
4) Qi F, Motz M, Jung K, Lassak J, Frishman D. Evolutionary analysis of polyproline motifs in Escherichia coli reveals their regulatory role in translation. PLoS Comput. Biol. 2018; 14: e1005987-19.
5) Starosta AL, Lassak J, Peil L, Atkinson GC, Woolstenhulme CJ, Virumäe K, et al. A conserved proline triplet in Val-tRNA synthetase and the origin of elongation factor P. Cell Rep. 2014; 9: 476–483.
6) Vieira-Silva S, Rocha E. The systemic imprint of growth and its uses in ecological (meta)genomics. PLoS Genet. 2009; 6: 1–15.
7) Gibson B, Wilson DJ, Feil E, Eyre-Walker A. The distribution of bacterial doubling times in the wild. Proc Biol Sci. 2018; 285(1880): 20180789

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
This research is fascinating and I really enjoyed the write up. This is such a great example of good science communication. Thanks for sharing!