The power of air: Airborne environmental DNA plant community monitoring
Published in Ecology & Evolution

Lubbock, Texas is a challenging place to conduct traditional plant community surveys. It seems that just about every plant species you find is trying to stab you, while most of the animal life is trying to bite you. Honey mesquite (Prosopis glandulosa) for example, loves growing in dense patches in this environment while also sporting some rather impressive thorns. In addition to the biota making surveys challenging, the weather with >100 °F daytime high temperatures in the summer and haboob dust storms in the fall also offer unique challenges (Figure 1b). While these challenges may be specific to Lubbock, any location on Earth is going to offer unique challenges to field work and biodiversity monitoring. Despite these challenges, increasing rates of disturbance from climate change, biological invasion, and habitat destruction, make the monitoring of those ecosystems all the more important. However, traditional surveying, in addition to being physically challenging, oftentimes destroys the environment, takes long periods of time, and requires a lot expertise to accurately identify species. These challenges and shortcomings of traditional surveys matched with an urgency to conduct biodiversity monitoring indicates a need to explore other avenues of species detection. That’s why we have turned to the use of cutting edge genetic methods to address some of these concerns.
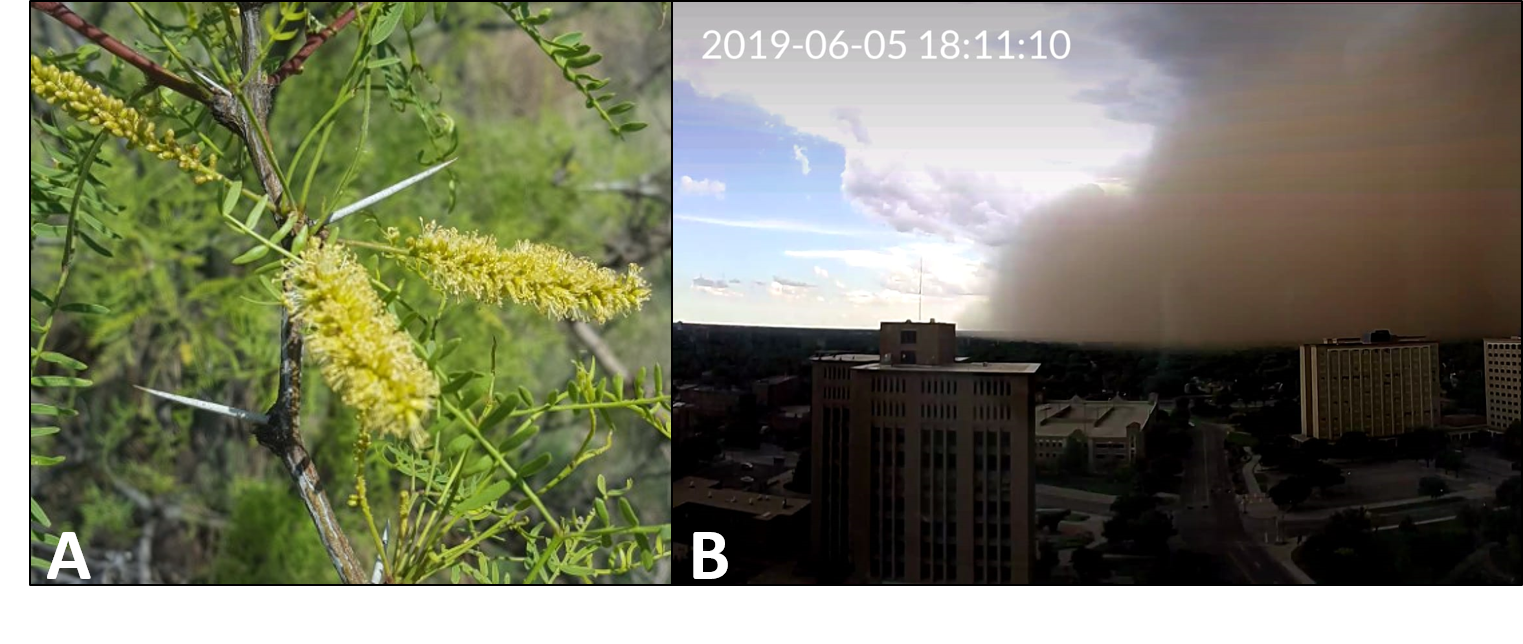 Figure 1. (A) Honey mesquite thorns (photo credit: Dr. Robert Cox) and (B) Haboob dust storm moving across Lubbock Texas (photo credit: Dr. Karin Ardon-Dryer)
Figure 1. (A) Honey mesquite thorns (photo credit: Dr. Robert Cox) and (B) Haboob dust storm moving across Lubbock Texas (photo credit: Dr. Karin Ardon-Dryer)
One of these genetic methods is environmental DNA (eDNA), which refers to the genetic material that is shed from an organism into its environment. Researchers can than collect bulk environmental samples such as water, soil, or air to determine whether that sample contains genetic material from a species of interest. Furthermore, this method has been shown to directly address the limitations of traditional surveys by being faster, more efficient, and less invasive. In our recently released manuscript we were interested in determining if airborne eDNA could be used to survey a whole plant community. Anyone who has suffered from allergies can attest to the fact that there is plant DNA in the air. While that means that there is pollen present, there is also DNA from a variety of other sources such as leaf fragments, flower fragments, and free floating DNA. Furthermore, we know that airborne eDNA is sensitive to the environment and can detect this varied material and acute changes on the landscape. This means that we can expand airborne eDNA analysis past pollen and single species detection to assess all the species on a study site regardless of how they pollinate (wind, insect, animal, etc.) or if they are currently pollinating.
To test this hypothesis, we preformed both a traditional and airborne eDNA plant community survey over the course of a year across the Texas Tech University Native Rangeland (Figure 2A). We revisited the thorns, heat, and snakes twice from June 2018 to June 2019 to conduct our traditional plant survey. The traditional surveys consisted of 9 sampling locations, each consisting of three 100 m transects organized as spokes around a central point. While this was taking place, we also deployed nine Big Spring Number Eight (BSNE) dust traps across the rangeland at the center of each one of our spokes (Figure 2a). Each BSNE trap consisted of two independent triangular metal collectors approximately 0.9 and 0.4 m above the ground (Figure 2b). Each collector included a metal sail that aligned the opening at the front of the collector into the wind to collect dust and particles at the bottom of the trap. The traditional surveys took hundreds of work-hours each to try and complete, with over ten volunteers and plant identification experts all working together to try and find the most diversity possible. On the other hand, the eDNA sampling events took only two hours to complete so we were able to sample eDNA 22 times across the entire year, taking fewer work-hours than a single traditional survey.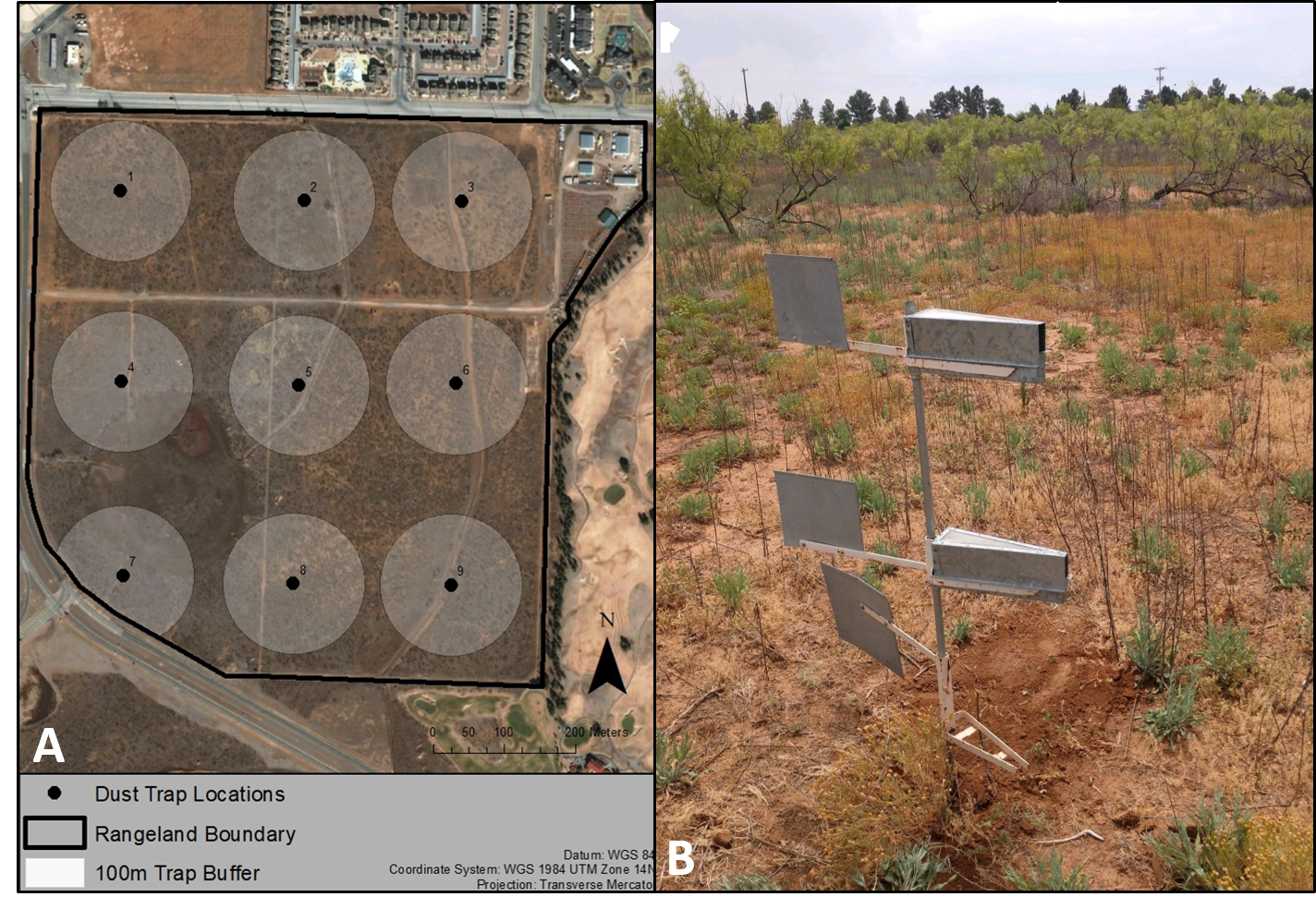
Figure 2 (A) The Texas Tech University Native Rangeland (Lubbock County, TX, United States) study site where both the traditional and metabarcoding plant community surveys were conducted. The points and numbers represent individual Big Spring Number Eight dust trap locations while the buffers around each point represent the 100 m traditional survey extent. (B) The Big Spring Number Eight Dust traps that collected the airborne eDNA.
Upon examining our results, we found that our airborne eDNA survey was actually able to detect more species (N=91) compared to the traditional survey (N=80). In general, our airborne eDNA analysis detected more grasses and forbs with less showy flowers, while the traditional method detected fewer grasses but also detected rarer forbs with large showy flowers (Figure 3). Furthermore, the airborne eDNA survey was much more efficient than the traditional survey allowing us to survey across the entire year and see trends on our study site that the traditional surveys missed. For example, the eDNA survey found that tansy mustard (Descurainia pinnata) was one of the most common species on our study site, but interestingly, the traditional survey rarely found it. The reason for this we realized however, turned out to be a large bloom event that occurred between the two traditional surveys. For a month, our study site was dominated by this species, but by the time the traditional survey took place in May there was little remaining to indicate this. However, since our airborne eDNA methodology was so efficient we could sample more and were able to detect this explosion of tansy mustard. We also found that the eDNA survey detected more invasive species than the traditional survey. While both methods detected two invasive species, the eDNA survey was able to detect a third invasive, tree of heaven (Ailanthus altissima). This is exciting because tree of heaven represents an invasive species that is at the perfect stage for eradication. The most effective time to stop an invasive species is before it becomes established, and that is the stage tree of heaven is in at our study site, where airborne eDNA was the only method able to detect it.
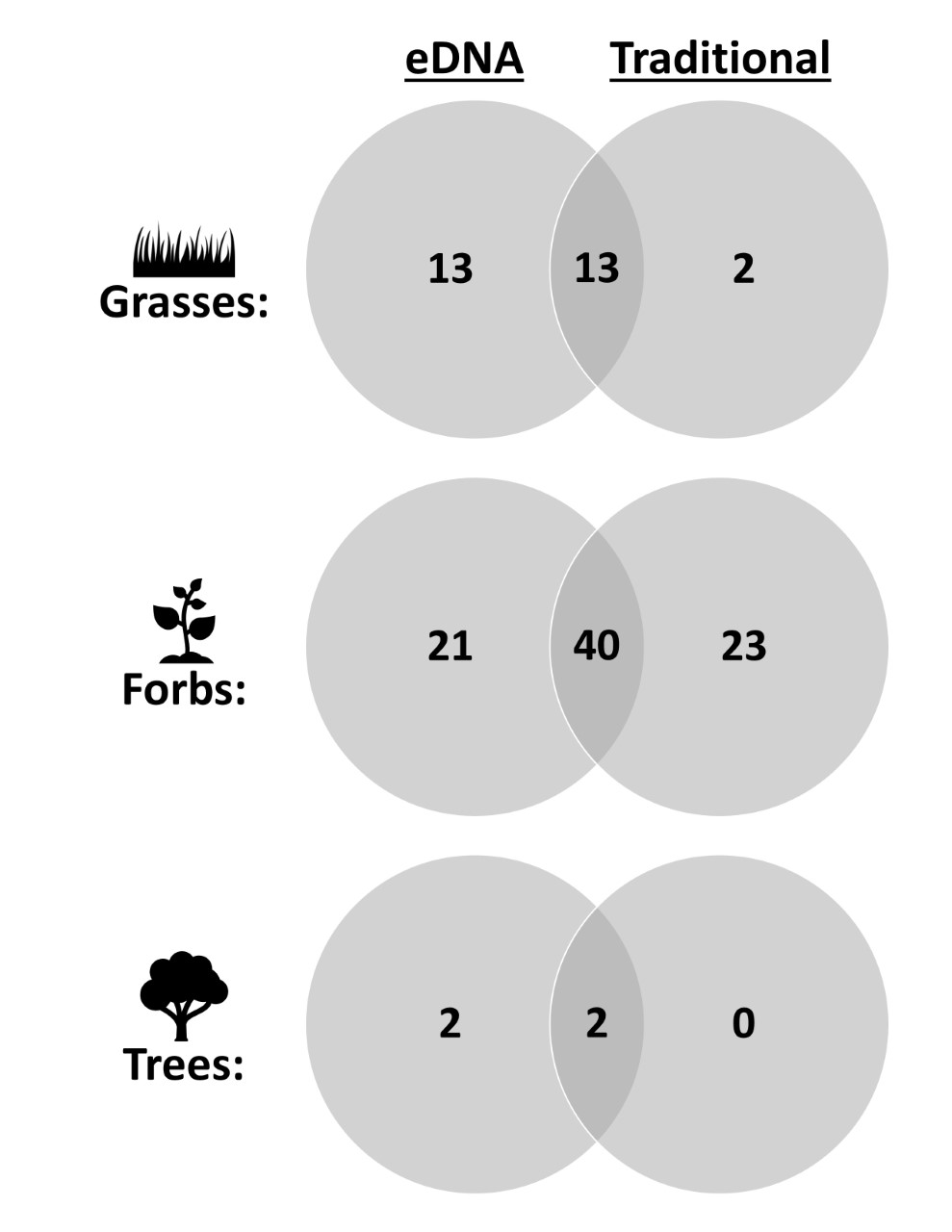
Figure 3. Venn diagram displaying the number of (A) grasses, (B) forbs, (C) and trees that were found by the eDNA and traditional methods alone and together
Detecting and analyzing airborne eDNA has the potential to revolutionize the way plant communities are surveyed. Furthermore, we were able to detect various forms of species, regardless of pollination syndrome, indicating that airborne eDNA is not beholden to the presence of pollen or just wind pollinated species. With much less effort, we were able to detect more species than a traditional survey, including more invasive species. While more research is needed into the ecology of airborne eDNA (origin, state, transport, and state), airborne eDNA in general could help to detect invasive species, endangered species, track climate change, and monitor biodiversity. Thinking back to the challenges of monitoring the Lubbock plant communities (and other plant communities across the world), airborne eDNA could help to address these challenges while providing a plethora of information that could be used to better understand this changing world.
Follow the Topic
-
BMC Ecology and Evolution
An open access, peer-reviewed journal interested in all aspects of ecological and evolutionary biology.
Related Collections
With Collections, you can get published faster and increase your visibility.
Evolutionary biomechanics
Evolutionary biomechanics is an interdisciplinary field that integrates biology, physics, and engineering to understand the mechanical principles governing movement, function, and adaptation across diverse species. By examining how anatomical structures evolve in response to mechanical demands, researchers can uncover the adaptive strategies that enable species to survive and thrive in their environments.
This Collection will highlight research on the biomechanics of movement in extant and extinct animals, from invertebrates to large vertebrates, across land, water and air. By linking biomechanics with ecological interactions and evolutionary processes, studies in this field provide key insights into locomotion, musculoskeletal function, movement ecology, and the energetic costs of life in different environments.
Topics of research may include but are not limited to:
•
Musculoskeletal biomechanics and locomotion
•
Comparative biomechanics
•
Biomechanics of soft-bodied and flexible organisms
•
Movement ecology and migration
•
Locomotion in complex environments
•
Predator-prey interactions and functional morphology
•
Insect biomechanics
•
Adaptation and evolutionary biomechanics
•
Bio-inspired robotics and engineering
•
Advances in biomechanical methodologies
This Collection supports and amplifies research related to SDG 15: Life on Land and SDG 14: Life Below Water.
Publishing Model: Open Access
Deadline: Jul 13, 2026
Ecology of soils
BMC Ecology and Evolution invites researchers to submit their work on soil ecosystems and their implications for environmental sustainability. Soils are living, dynamic systems that support a vast array of organisms, from tiny microbes to plants and animals. They play a vital role in keeping our planet healthy by recycling nutrients, storing carbon, and helping regulate water and climate. Understanding how soil life functions and adapts is essential for tackling major environmental challenges like climate change, habitat loss, and soil degradation.
This collection of articles aims to showcase the latest research in soil ecology, emphasizing the interactions among soil organisms, biogeochemical processes, and ecosystem functions across various landscapes. We invite submissions that explore the following topics:
•Microbial and faunal diversity in soils: Patterns, drivers, and functional roles of bacteria, fungi, protists, and invertebrates
•Soil biogeochemistry and ecosystem functioning: Nutrient cycling, carbon sequestration, and soil health in both natural and managed ecosystems
•Soil-plant interactions: Rhizosphere processes, plant-microbe symbioses, and their effects on vegetation dynamics
•Land use and climate change impacts on soil communities: Responses of soil biodiversity and functions to agricultural intensification, urbanization, pollution, and climate shifts
•Soil metagenomics, phylogenetics, and functional ecology: Advances in molecular approaches for studying soil microbial and faunal communities
•Conservation and restoration of soil ecosystems: Strategies for maintaining soil biodiversity and ecosystem services in degraded landscapes
All manuscripts submitted to BMC Ecology and Evolution, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
This Collection supports and amplifies research related to SDG 15: Life on Land.
Publishing Model: Open Access
Deadline: Jun 02, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in