The world's longest selection experiment on mice provides unique animal models for exploring the architecture of polygenic traits
Published in Ecology & Evolution
At the Research Institute for Farm Animal Biology (FBN) in Dummerstorf, Germany, a unique set mouse lines were generated over the course of more than 50 years of artificial selection. These lines have evolved impressive phenotypes of high fertility, body size and endurance fitness. To our knowledge, this is the longest selection experiment ever conducted on mice and we were keen on peering into the genomes of these unique animals, in order to uncover known and new loci associated to the traits under selection.
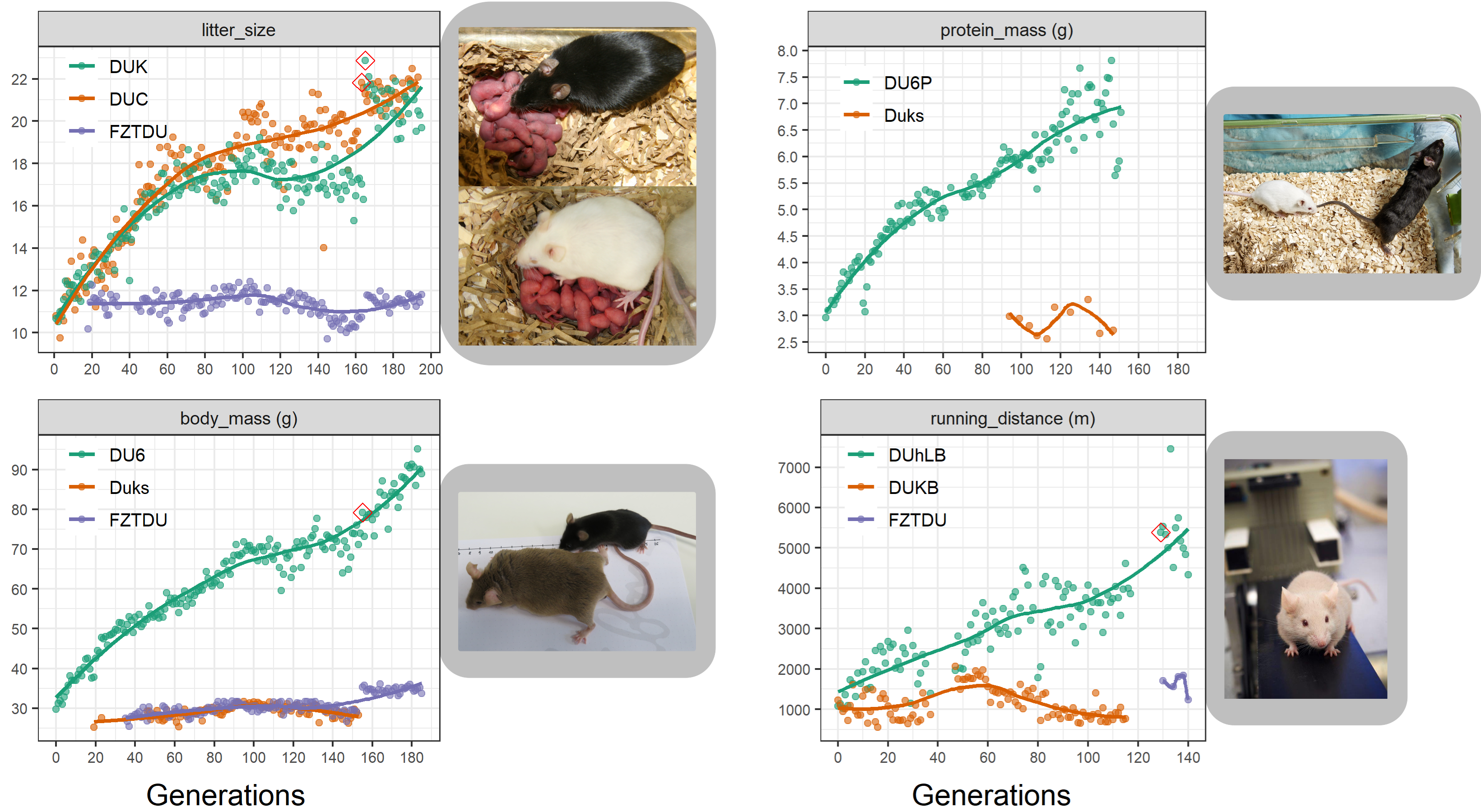
As a population adapts to a selective pressure, the alleles responsible for adaptation become more and more frequent over time. To identify the subtle allele frequency changes underlying complex traits, neutral evolution needs to be rigorously modelled, requiring genetic information not only from the present population but also from ancestral populations, ideally the founders.
Despite the fact that the Dummerstorf mouse lines are outbred mouse populations, we found high levels of inbreeding within each line, resulting from genetic drift after a severe population bottleneck in 2011. Consequently, selected mice are highly homozygous, while at the same time, lines are genetically uniform and distinct from each other.
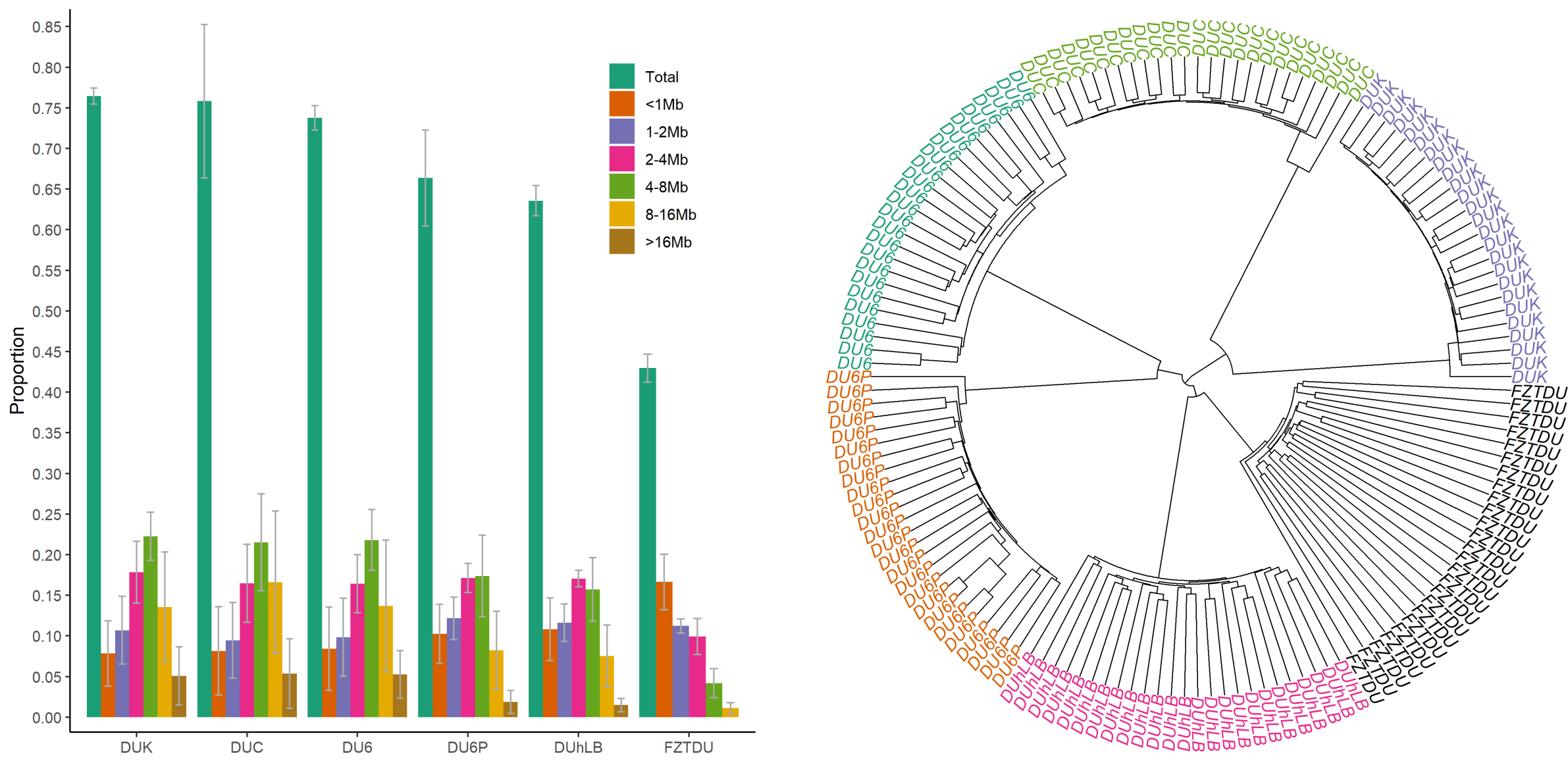
Since genetic information from ancestral generations was not available (this experiment was started in 1969, before the genomic era) and pedigrees were incomplete, detection of signatures of selection was practically impossible. It was also unfeasible to know if response to selection resulted from large- or small-effect alleles, though the complex nature of the selected traits led us to infer that the former is rather unlikely.
The historical resources at our disposal were thus limited, so we tried to come up with an alternative approach that would at least give us an idea of which genes could be involved in the evolution of the selected traits. First, we obtained the genomewide genetic differentiation of each selected line relative to the control line (the control line as a proxy of the founder population, exposed only to genetic drift). We then looked for regions of line-specific high genetic differentiation, each one corresponding to a genomic location in which only the target line is highly differentiated from the control line, while all remaining lines are undifferentiated. This approach revealed a number of candidate genes, some of which are already known to be relevant in the biology of fertility, body mass, muscle growth, and endurance.
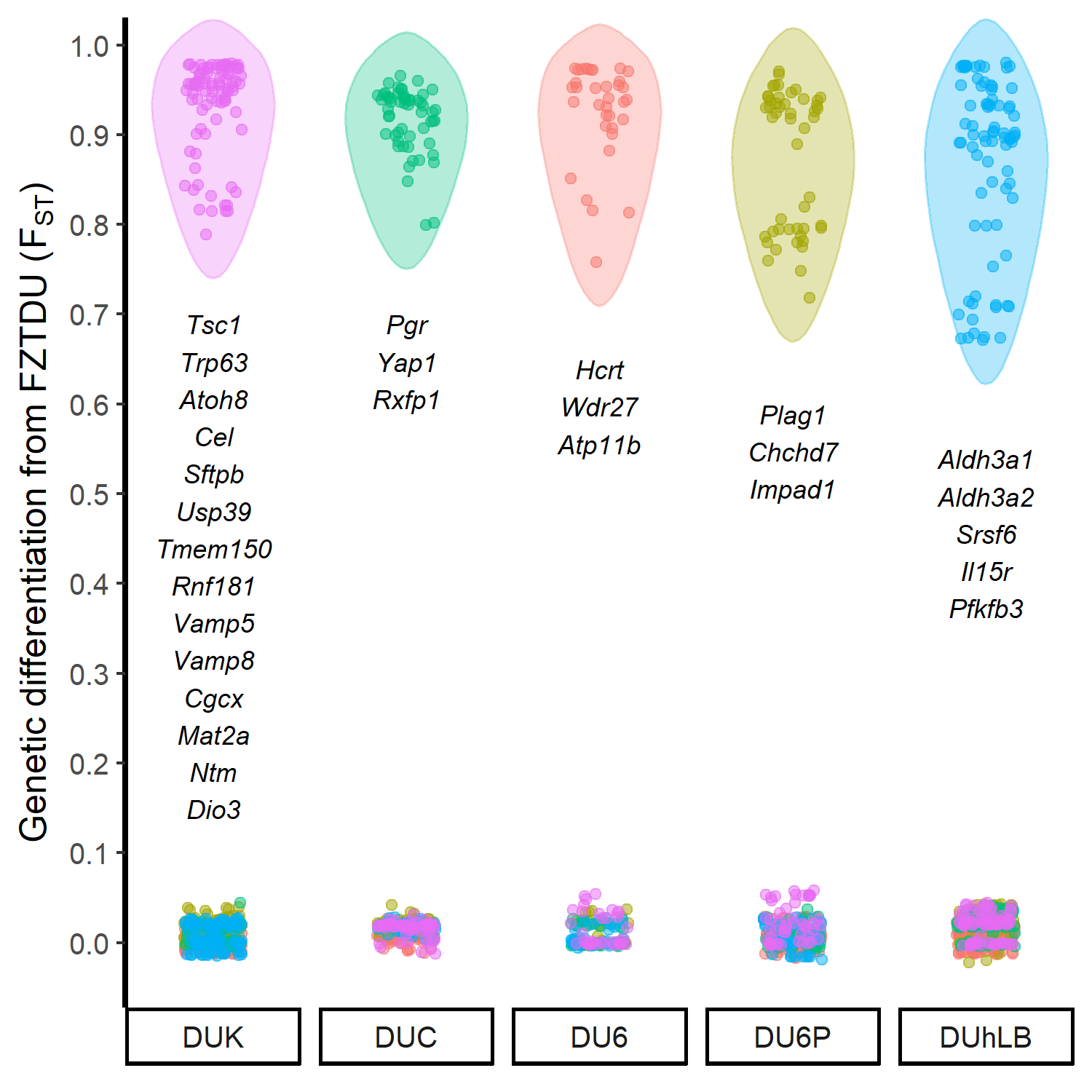
Though we understand that the lack of a proper neutral model prevents us from taking these results as final, these are promising discoveries awaiting validation. We hope that within the next years, the hidden trait-associated alleles within the genomes of these unique mouse models can be uncovered.
References
1. Palma-Vera SE, Reyer H, Langhammer M, Reinsch N, Derezanin L, Fickel J, et al. Genomic characterization of the world’s longest selection experiment in mouse reveals the complexity of polygenic traits. BMC Biol. 2022;20:52.
Follow the Topic
-
BMC Biology
This is an open access journal publishing outstanding research in all areas of biology, with a publication policy that combines selection for broad interest and importance with a commitment to serving authors well.
Related Collections
With Collections, you can get published faster and increase your visibility.
Cancer metabolism
BMC Biology is calling for submissions to our Collection on Cancer metabolism. Cancer metabolism is a developing field that explores the biochemical and physiological changes that occur in cancer cells, which often exhibit distinct metabolic pathways compared to normal cells. Alterations such as the Warburg effect, where cancer cells preferentially utilize glycolysis for energy production even in the presence of oxygen, play a pivotal role in tumor progression and survival. Understanding the intricacies of cancer metabolism provides insights into how tumors adapt to their microenvironments and highlights potential therapeutic targets for intervention.
Future research in cancer metabolism promises to yield transformative insights that could reshape therapeutic approaches and improve precision medicine. The continued focus on metabolic pathways may lead to the identification of new biomarkers for cancer diagnosis and prognosis, as well as novel strategies to enhance the sensitivity of cancer cells to treatment. As we deepen our understanding of the metabolic landscape of tumors, we may uncover innovative strategies that exploit these vulnerabilities, ultimately leading to novel and more effective cancer treatments, as well as improved patient outcomes.
Recent advancements in the field, including the identification of metabolic reprogramming strategies and the influence of diet on tumor growth, have opened new avenues for research. Investigations into the roles of lipids, fatty acids, and dietary interventions, such as ketogenic diets, are revealing potential methods for manipulating tumor metabolism and enhancing the efficacy of existing treatments. We invite researchers to submit their work to this Collection, which aims to showcase groundbreaking research and technologies addressing cancer metabolism and support the advancement of this field, encompassing a wide array of topics related to metabolic pathways and their implications for cancer biology and therapy.
Potential topics include but are not limited to:
- Metabolic reprogramming in cancer cells
- The Warburg effect and its implications
- Role of mitochondria in cancer metabolism
- Impact of dietary interventions on tumor metabolism
- Glycolysis and lipid metabolism in cancer
- Animal models
- Imaging and method developments
- Metabolic engineering
This Collection supports and amplifies research related to SDG 3 (Good Health and Well-being).
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Oct 30, 2026
Organelles and molecular assemblies in intercellular signalling
BMC Biology is calling for submissions to our Collection on Organelles and molecular assemblies in intercellular signalling. This Collection aims to highlight the diverse roles of organelles and molecular assemblies in driving cellular and developmental processes. From intracellular organization to tissue-level coordination, these structures are essential for maintaining homeostasis and responding to physiological and pathological cues.
We welcome research that explores how organelles and molecular assemblies contribute to intercellular communication, development, tissue dynamics, and disease—across all biological systems and model organisms.
We welcome studies on exosomes, migrasomes, blebbisomes, mechanosignalling, and the role of organelles—such as mitochondria—in extracellular and intercellular communication. Studies describing the role of cell adhesion molecules, gap junctions, synapses and cell surface receptors in intercellular communication are also encouraged. Studies may also focus on the disruption of secretory pathways and other organelle-mediated processes during disease onset, and their potential as a therapeutic target. Modelling and simulations that investigate and provide significant insights in intercellular communications and emergent properties in physiological contexts are also welcome.
Submissions on novel approaches in imaging, omics-based analyses, and high-throughput analysis to study organelle function and dynamics are particularly welcome.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Jul 18, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in