Too big to see: the uncovering of large phage genomes in the world's oceans
Published in Microbiology

This post tells the story of how we came about our study published in The ISME Journal about unusually large phage genomes we recovered from metagenomes of seawater collected around the world.
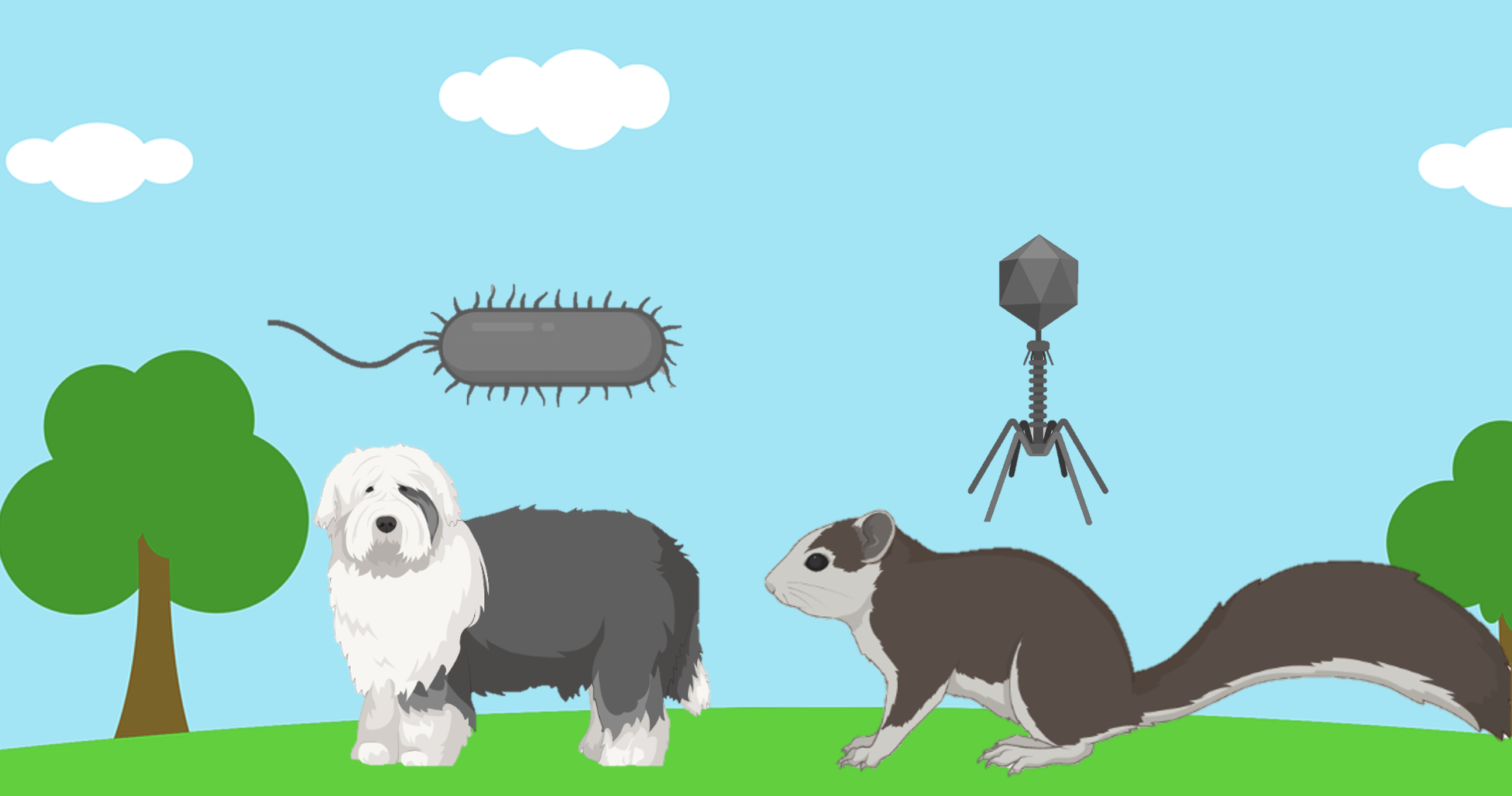
Imagine looking for your sheep dog, in the park and a giant squirrel runs past. Maybe you wouldn't have even noticed this massive squirrel because you were only looking for your dog, and squirrels are never as big as sheep dogs. This can be the case for bacterial and viral genomes in metagenomes - except your dog is bacteria, the squirrel is a virus, and the park is a metagenome.

Metagenomic data or reads (short sequences of DNA of all organisms in a sample) typically get assembled into contigs and subsequently into bins. Bins are groups of contigs that likely belong to the same genome. We typically only look for viruses at the contig-level because we know they tend to have small genomes, and we look for cellular genomes at the bin-level because we know they have big genomes - bigger than viruses at least… Think again!
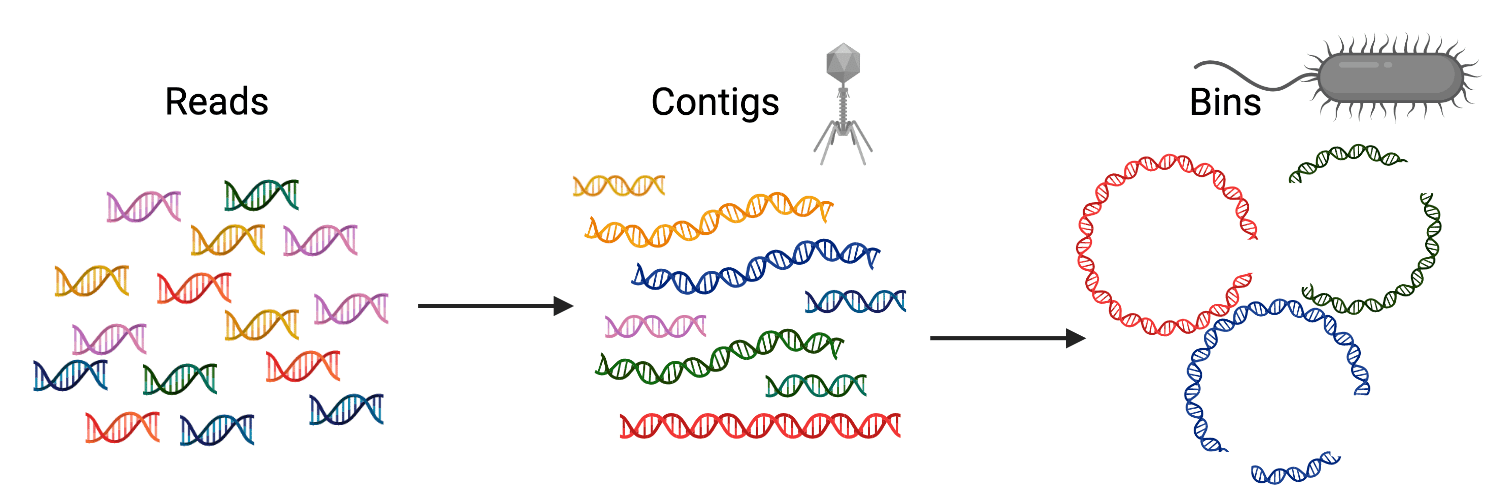
Some viruses can be "jumbo". Believe it or not, "jumbo bacteriophage" is a scientific term. Jumbo bacteriophages, or jumbo phages, are viruses of bacteria that are solely distinguished by having genomes over 200 kilobases in length (~4x the size of a typical phage). They can have large particles, too - as big as small bacteria. They've long been thought to be rare in nature, but this might be because common methods in virology bias toward smaller phages, such as the concentrations of agar used to isolate phages or the way we look for phages in metagenomes.
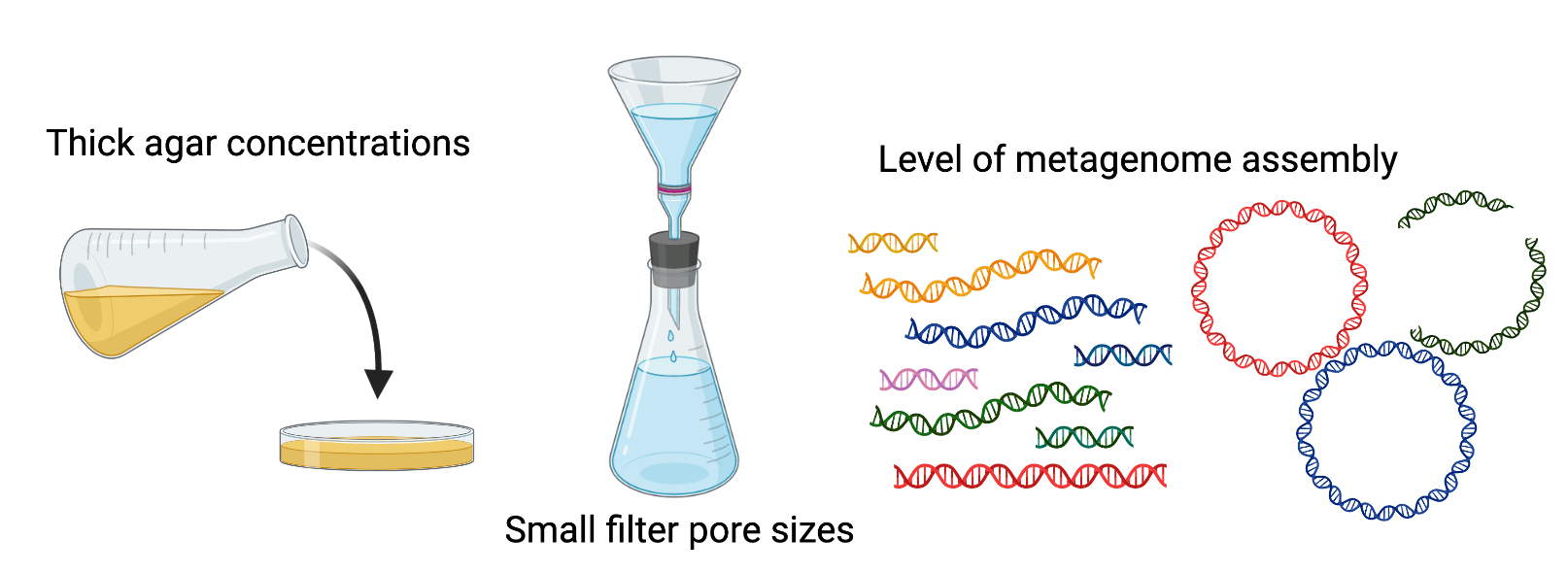
Since jumbo phage genomes can be quite large, some up to 700 kilobases, their genomes might be present in bins, rather than only assembled into contigs. And thus began my quest - to uncover genomes of jumbo phages in bins of metagenomes. Easy!
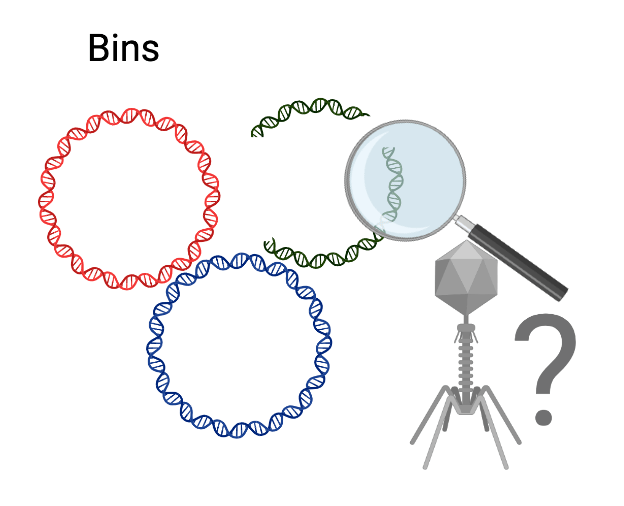
Not so fast! Jumbo phages are a bit unruly, rebellious, anarchic - one might say. Unlike bacteria or other organisms, jumbo phages lack single-copy, core genes that distinguish them from other viruses. In fact, jumbo phages likely evolved independently, multiple times from smaller phages. This presents challenges to detect jumbo phage genomes and estimate their quality and completeness.
To overcome these complications, we developed a robust pipeline that included multiple phage detection tools and comparisons of contig co-abundance profiles to account for genome contamination and completion. As part of this pipeline, we used Tara Oceans metagenomic data, which resulted in focusing our study on marine jumbo phages. Tara Oceans is a global sampling effort that has sailed around the ocean, collecting seawater samples from multiple depths and biomes.
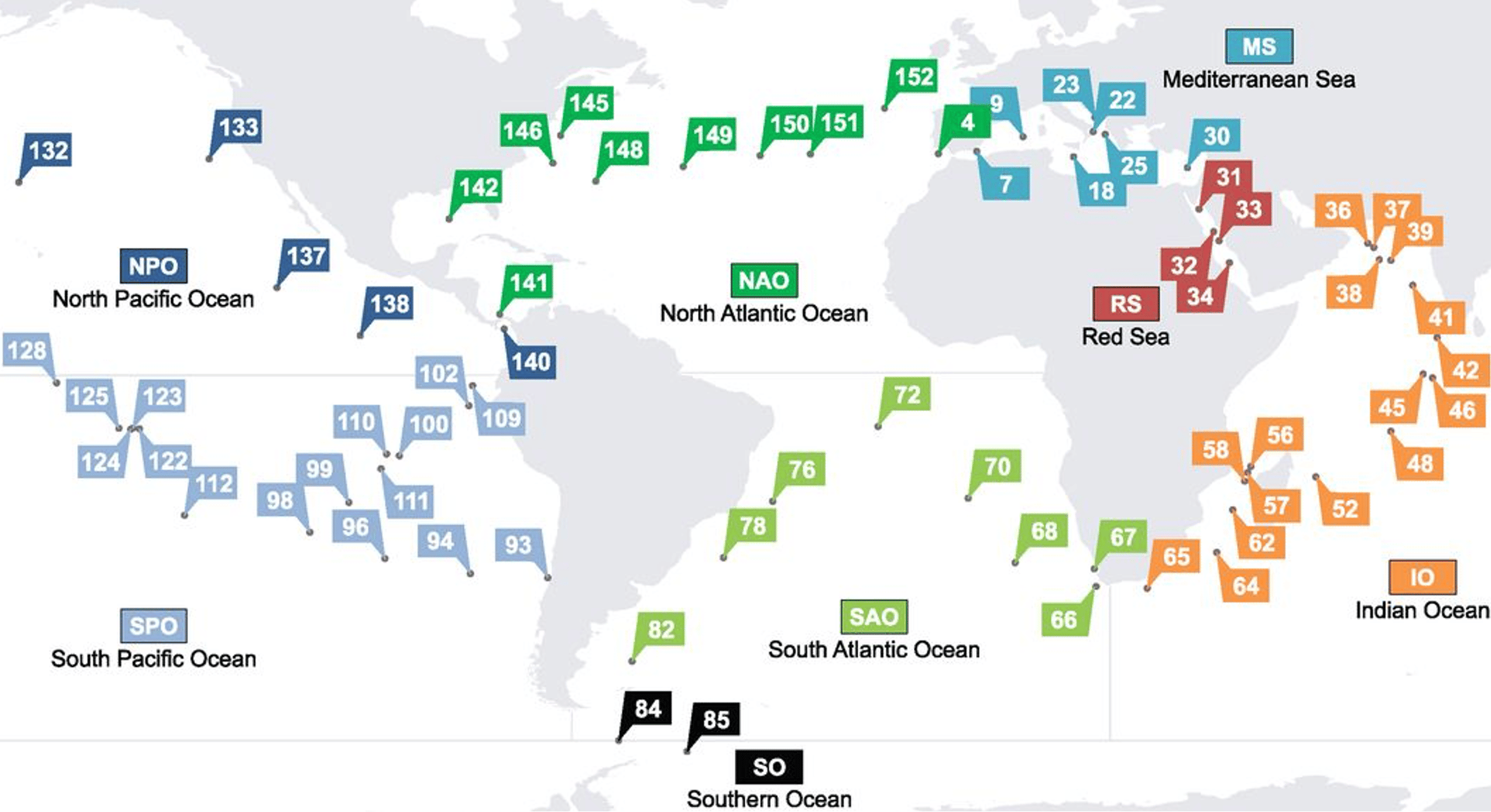
With this approach, we successfully recovered 85 bins of high quality jumbo phage genomes present in the ocean. We then grouped these jumbo phages based on gene content with reference jumbo phages, and we found that certain groups of jumbo phages are more prevalent in surface waters than deeper waters and vice versa. Our study shows that jumbo phages are not only diverse, cosmopolitan members of marine ecosystems, but also potentially overlooked members of other ecosystems due to lack of binning. You can read all about how we developed a pipeline to detect bins of jumbo phages and the distribution of these marine viruses in our paper here.
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in