TP53 and CMML – a rare but consequential combination.
Published in Cancer
Background:
Chronic myelomonocytic leukemia, a chronic yet severe blood cancer, is the most common MDS/MPN overlap neoplasm and is characterized by peripheral blood monocytosis and bone marrow dysplasia 1. Other clinical features can reflect both myelodysplasia (anemia, thrombocytopenia) and myeloproliferative effects (splenomegaly, systemic inflammatory symptoms). CMML can also transform to an acute leukemic phase (blast transformation; 15-30% over 3-5 years) which is challenging to treat. As with MDS and AML, no single genetic abnormality is diagnostic of CMML, however specific clinical and disease-associated genetic findings are used to inform prognosis, and potentially guide therapy. Prognostic scoring systems are derived from factors known to impact survival in patients with CMML – including lower hemoglobin values and platelet counts, increasing white cell count (‘proliferative disease’), immature myeloid cells found in the peripheral blood, increasing numbers of bone marrow blast cells and specific bone marrow chromosomal abnormalities. Newer scores have integrated the prognostic effect of several somatically mutated genes including ASXL1, NRAS, RUNX1 and SETBP1.
Genetic events which inactivate the tumor suppressor gene TP53, located on chromosome 17p are generally considered high-risk findings in MDS and AML and are included in prognostic tools for these diseases. TP53 mutations are enriched in secondary myeloid neoplasms occurring after exposure to prior DNA-damaging therapies (termed ‘therapy-related-’ or ‘-post-cytotoxic therapy’). Additionally, allelic status (i.e the presence or absence of one residual, intact copy of TP53) is important in cases of MDS, and the 5th edition of the WHO classification defines a myeloid neoplasm with biallelic TP53 mutations associated with high risk monosomal karyotypic abnormalities and adverse outcomes 2. Although not previously studied systematically, TP53 alterations are noteworthy for their rarity in reported CMML cohorts and are not considered in current CMML prognostic models. Given this rarity, in our experience, TP53 alterations detected with a clinical presentation of CMML raise some uncertainty as to whether CMML is the correct diagnosis. This led us to examine these cases in a collaborative cohort across two large academic medical centers. We systematically documented the frequency of TP53 alterations (mutations and/or deletions) in patients with a diagnosis of CMML, reviewed the cases for characteristic CMML features and considered their impact on prognosis.
Main Findings:
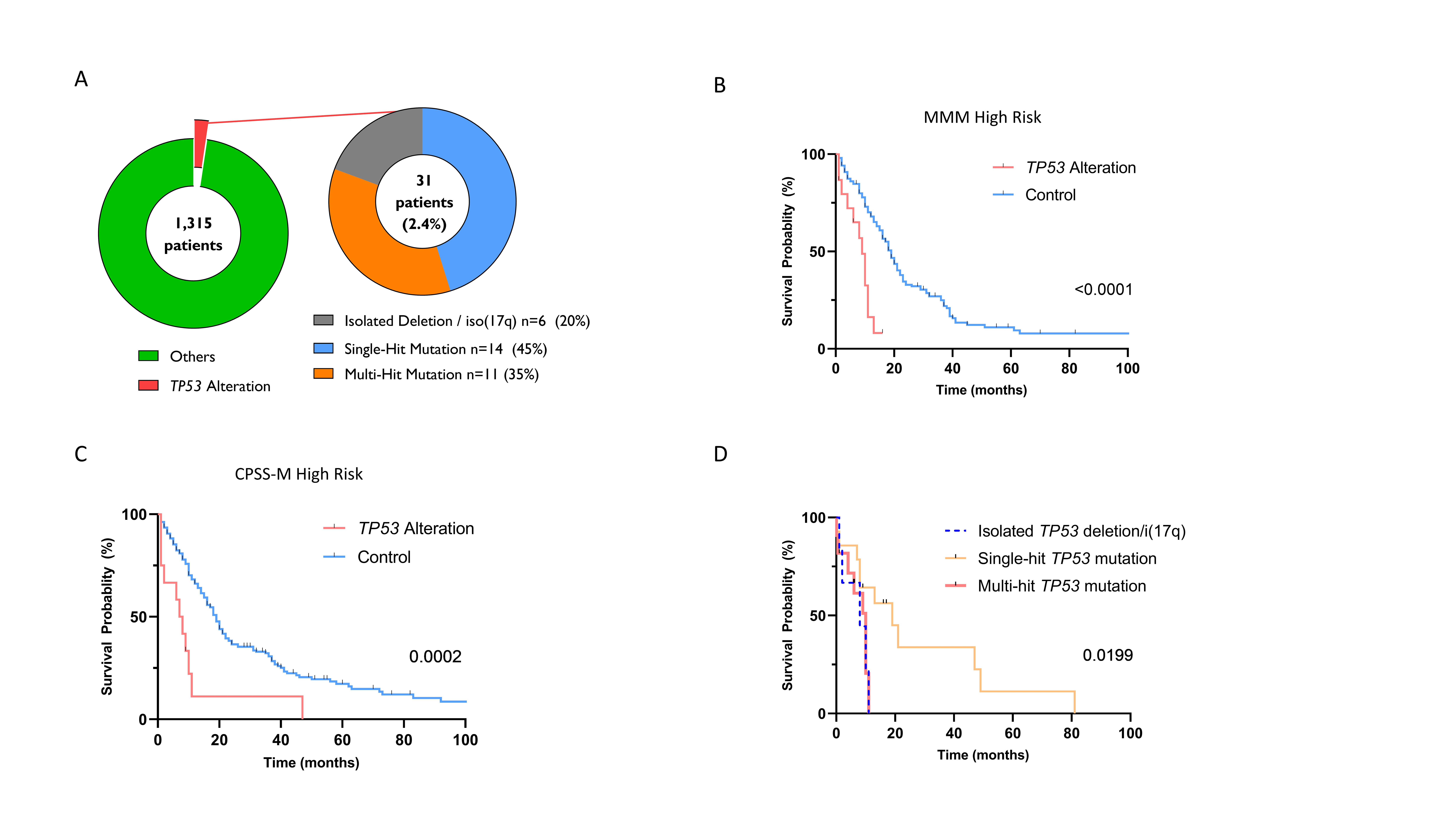
Amongst 1,315 patients with CMML, 1.9% had at least one TP53 mutation, while 0.6% had large structural deletions or rearrangements leading to loss of one copy of TP53 (Figure 1A). We confirmed that these cases met criteria for CMML, with peripheral blood monocytosis, bone marrow dysplasia, and expected rates of BM fibrosis. Cases with TP53 alterations were notable for having a higher proportion of patients with therapy-related CMML and higher bone marrow blast counts, meaning a greater proportion had the more aggressive CMML-2 subtype of the disease.
Considering the number of TP53 mutations present, the variant allele frequency of the mutation detected, and/or concurrent loss of one TP53 allele through deletion or monosomy 17, 11 cases were considered ‘multi-hit’ (implying complete loss of TP53 activity), while 14 cases were considered single-hit mutations (mutation on one allele only) 3. Multi-hit TP53 mutations were usually the dominant clone and associated with monosomal (high-risk) karyotypic abnormalities. Isolated deletion of 17p without concurrent TP53 mutation occurred in 6 cases, always due to formation of an isochromosome 17q, a finding which has been previously described as high risk and may be the basis of a future genetically defined myeloid neoplasm subtype.
The presence of TP53 alterations negatively impacted CMML prognosis, including for patients already deemed to have high risk disease by the commonly applied prognostic scores tested (Mayo molecular model and CPSS-molecular model, Figure 1 B-C). Cases with multi-hit mutations (median survival of 10 months), or isochromosome 17q (median 8 months) had an inferior prognosis relative to cases with single-hit TP53 mutations alone (median 19 months, Figure 1D).
Conclusions and next steps:
Establishing the prognostic impact of rare genetic abnormalities in myeloid neoplasms is challenging. Large collaborative research efforts have been key to understanding the nuances of TP53 alterations occurring in MDS and AML 4,5. The prognostic impact of these events cannot be assumed however as was recently highlighted by the apparent lack of impact of TP53 alterations on outcomes in patients with myeloid neoplasms with an associated germline DDX41 variant. Given the rarity of TP53 alterations in CMML, this investigation was also dependent on a large cohort with detailed follow up, which has only been achievable through enthusiastic collaboration. Applying the approach to allelic status (single and multi-hit mutations) that was defined in other myeloid neoplasms – provided a coherent structure and evidence base with which to explore this issue in CMML.
While not a formal integration of TP53 to either of the prognostic models examined, we believe these findings are important. We confirm that cases can concurrently meet criteria for CMML, and harbor TP53 alterations with independent prognostic impact. Our clinical report does not explain the biological basis for the rarity of TP53 mutations in CMML relative to other myeloid disorders. We briefly explore some possibilities in the manuscript – there is a clear genotype-phenotype association for biallelic TET2 inactivation, and TET2/SRSF2 co-mutation in generating the monocyte skewed hematopoiesis that is fundamental to CMML, without any defined role for TP53 alterations. Alternatively, TP53 mutated clones have been associated with abnormalities of the erythroid lineage, and loss of differentiation capability in other contexts. Ultimately, further study is required to confirm these mechanisms.
The main take-away point of this report is that although representing a rare finding in CMML, TP53 alterations retain their negative prognostic impact, and principles of allelic status defined in MDS can be applied to refine this impact further.
References:
1 Patnaik MM. How I diagnose and treat chronic myelomonocytic leukemia. Haematologica 2022; 107: 1503–1517.
2 Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia 2022; 36: 1703.
3 Hasserjian RP, Orazi A, Orfao A, Rozman M, Wang SA. The International Consensus Classification of myelodysplastic syndromes and related entities. Virchows Archiv 2023; 482: 39–51.
4 Bernard E, Nannya Y, Hasserjian RP, Devlin SM, Tuechler H, Medina-Martinez JS et al. Implications of TP53 allelic state for genome stability, clinical presentation and outcomes in myelodysplastic syndromes. Nature Medicine 2020 26:10 2020; 26: 1549–1556.
5 Hiwase D, Hahn C, Tran ENH, Chhetri R, Baranwal A, Al-Kali A et al. TP53 mutation in therapy-related myeloid neoplasm defines a distinct molecular subtype. Blood 2023; 141: 1087–1091.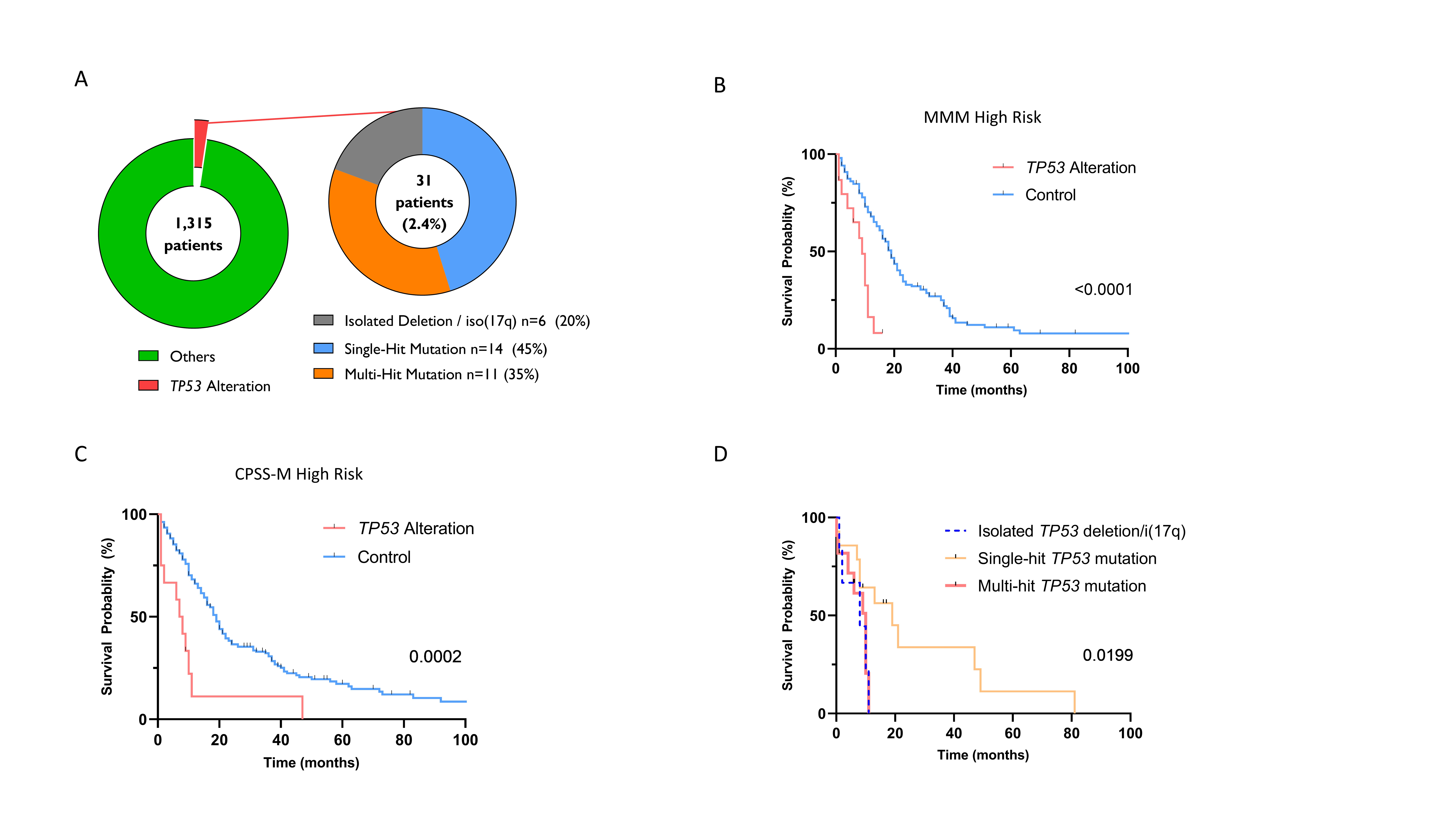
Follow the Topic
-
Leukemia
This journal publishes high quality, peer reviewed research that covers all aspects of the research and treatment of leukemia and allied diseases. Topics of interest include oncogenes, growth factors, stem cells, leukemia genomics, cell cycle, signal transduction and molecular targets for therapy.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in