TransPolymer: a Transformer-based language model for polymer property predictions
Published in Materials
Accurate prediction of polymer properties is of great significance in polymer development and design. Conventionally, laborious experiments or simulations are required to evaluate the function of polymers. Deep learning models have been broadly investigated for property predictions of polymer materials by directly learning expressive representations from data to generate deep fingerprints (FPs), instead of relying on manually engineered descriptors. For instance, Graph neural networks (GNNs), which have outperformed many other models on several molecules and polymer benchmarks, can learn representations from graphs and find optimal fingerprints based on downstream tasks. Despite the numerous success in representation learning from molecules, it's more challenging for GNNs to learn representations from polymers, as the structural and conformational information required by GNNs are not explicitly known for polymers due to the varying polymer structures under different conditions. Moreover, the optimal method of graph representation for polymers is still obscure. The degree of polymerization varies for each polymer, thus using the repeating unit only as a graph is likely to result in missing structural information.
Meanwhile, another plausible approach is to represent polymers as texts and apply language models for representation learning. A few systems have been developed for molecules and polymers like SMILES1 and BigSMILES.2 Chemistry sequences are suggested to have the same structure as a natural language like English, in terms of the distribution of text fragments and molecular fragments.3 This elucidates the development of sequence models similar to those in computational linguistics for extracting information from chemical sequences and realizing the intuition of understanding chemical texts just like understanding natural languages. Recurrent neural networks (RNNs) and their variants have been widely investigated in this case, while RNN-based models rely on previous hidden states for dependencies between words and tend to lose information when they reach deeper steps.
In addition, the acquisition of polymer labels is expensive and time-consuming which greatly limits the performance of deep learning models on novel polymers. Polymer databases are more complicated to build compared with those for molecules: polymer properties would be different even for polymers with the same repeating units if some key factors like the degree of polymerization or molecular weight distribution change. The situation becomes even worse when some of the polymer data sources are not fully accessible. Therefore, it is crucial to enlarge the data space through the augmentation of known data and making use of the abundant unlabeled data.
Recently, Transformer, which has exhibited superior performance in many natural language processing (NLP) tasks, relies on attention mechanism for learning relationships between tokens in the sequence.4 Various Transformer-based models have shown great potential in pretraining on abundant unlabeled data in a self-supervised manner for enhancement in model performance and generalization in downstream tasks. However, such Transformer-based models have not been investigated for polymer property predictions. Herein, a Transformer-based language model named TransPolymer is proposed for polymer property predictions in this paper.
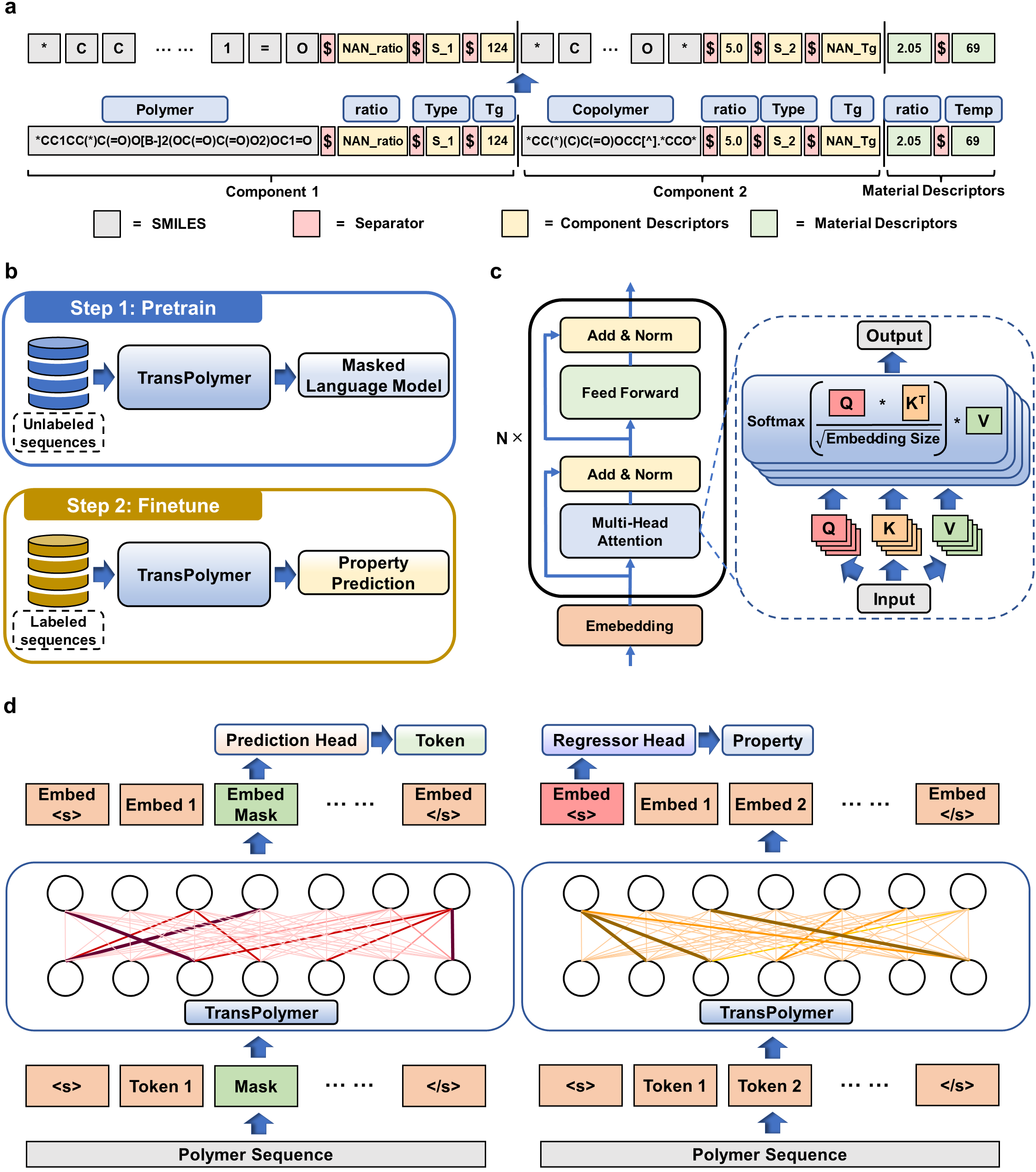
The TransPolymer framework consists of tokenization, Transformer encoder, pretraining, and finetuning. Each polymer data is first converted to a string of tokens through tokenization. Our proposed polymer tokenizer with chemical awareness enables learning representations from polymer sequences, shown in Figure 1(a). The repeating units of polymers are embedded using SMILES and additional descriptors (e.g., degree of polymerization, polydispersity, and chain conformation) are included to model the polymer system. Plus, copolymers are modeled by combining the SMILES of each constituting repeating unit along with the ratios and the arrangements of those repeating units. Moreover, materials consisting of mixtures of polymers are represented by concatenating the sequences for each component as well as the descriptors for the materials. Besides, each token represents either an element, the value of a polymer descriptor, or a special separator. Therefore, the tokenization strategy is chemical-aware and thus has an edge over the tokenizer trained for natural languages which tokenizes based on single letters.
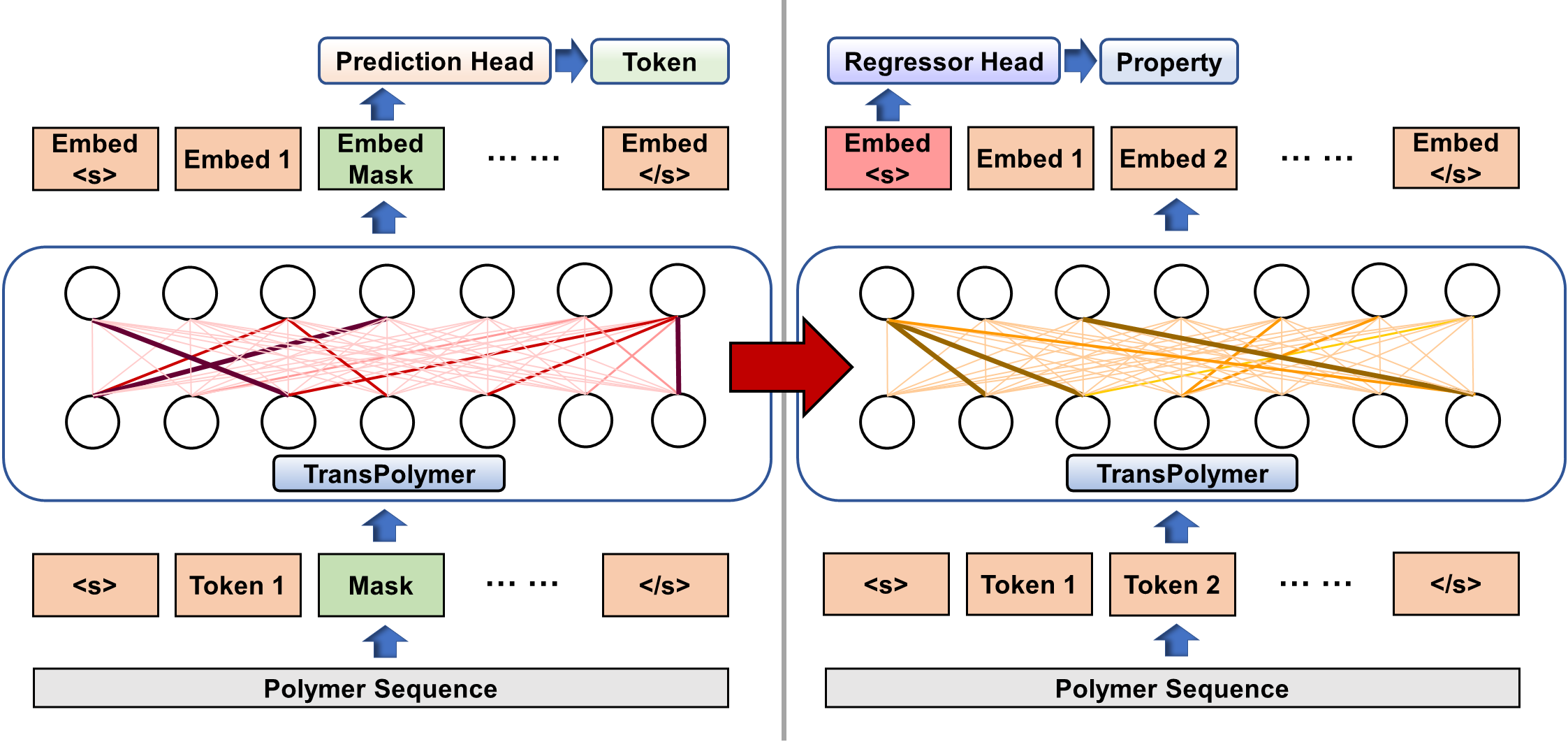
Transformer encoders are built upon stacked self-attention and point-wise, fully connected layers, shown in Fig. 1(c). Unlike RNN or CNN models, Transformer depends on the self-attention mechanism that relates tokens at different positions in a sequence to learn representations. Scaled dot-product attention across tokens is applied which relies on the query, key, and value matrices. To learn better representations from large unlabeled polymer data, the Transformer encoder is pretrained via Masked Language Modeling (MLM), a universal and effective pretraining method for various NLP tasks. As shown in Fig. 1(d) (left), 15% of tokens of a sequence are randomly chosen for possible replacement, and the pretraining objective is to predict the original tokens by learning from the contexts. The pretrained model is then finetuned for predicting polymer properties with labeled data. Particularly, the final hidden vector of the special token '<s>' at the beginning of the sequence is fed into a regressor head which is made up of one hidden layer with SiLU as the activation function for prediction as illustrated in Fig. 1(d) (right).
Rigorous experiments on ten polymer property prediction benchmarks demonstrate the superior performance of TransPolymer. Moreover, we show that TransPolymer benefits from pretraining on large unlabeled datasets via masked language modeling. Experimental results further manifest the important role of self-attention in modeling polymer sequences. You can find more details of the results and discussion using the article link attached to this post.
In summary, we have proposed TransPolymer, a Transformer-based model with MLM pretraining, for accurate and efficient polymer property prediction. Given the desirable model performance and outstanding generalization ability out of a small number of labeled downstream data, we anticipate that TransPolymer would serve as a potential solution to predicting newly designed polymer properties and guiding polymer design. For example, the pretrained TransPolymer could be applied in the active-learning-guided polymer discovery framework, in which TransPolymer serves to virtually screen the polymer space, recommend the potential candidates with desirable properties based on model predictions, and get updated by learning on data from experimental evaluation. In addition, the outstanding performance of TransPolymer on copolymer datasets compared with existing baseline models has shed light on the exploration of copolymers. In a nutshell, even though the main focus of this paper is placed on regression, TransPolymer can pave the way for several promising (co)polymer discovery frameworks.
Article Information:
Xu, C., Wang, Y. & Barati Farimani, A. TransPolymer: a Transformer-based language model for polymer property predictions. npj Comput Mater 9, 64 (2023).
Article Link:
https://www.nature.com/articles/s41524-023-01016-5
References
- Weininger, D.: Smiles, a chemical language and information system. 1. introduction to methodology and encoding rules. J. Chem. Inf. Comput. 28(1), 31–36 (1988)
- Lin, T.-S., et al.: Bigsmiles: a structurally-based line notation for describing macromolecules. ACS Cent. Sci. 5(9), 1523–1531 (2019)
- Cadeddu, A., et al.: Organic chemistry as a language and the implications of chemical linguistics for structural and retrosynthetic analyses. Angew. Chem. Int. Ed. 53(31), 8108–8112 (2014)
- Vaswani, A., et al.: Attention is all you need. Advances in neural information processing systems 30 (2017)
Follow the Topic
-
npj Computational Materials
This journal publishes high-quality research papers that apply computational approaches for the design of new materials, and for enhancing our understanding of existing ones.
Related Collections
With Collections, you can get published faster and increase your visibility.
Recent Advances in Active Matter
Publishing Model: Open Access
Deadline: Sep 01, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in