Two of a kind: a snapshot of microsporidia ribosomes.
Published in Cell & Molecular Biology
Earlier this year I found myself speaking to a group of secondary school children about our research on microsporidia as part of an outreach project combining art and science (see Fig. 1 for some of their fabulous creations!). The artist running the workshop suggested reading a description of the parasites to the children and getting them to recreate what they imagined they looked like. As I was observing the children’s interpretations, it struck me again just how intriguing these parasites are and how much more there is to learn about them.

Figure 1. Microsporidia-inspired art from children in Years 7-11 from St. James School in Exeter, UK from a workshop run with artist Anna Boland.
What are microsporidia?
Microsporidia are single-celled eukaryotic parasites. Closely related to fungi, there are over 1700 known species of microsporidia and they infect all types of animals ranging from insects to fish to humans. They cause the disease microsporidiosis in humans which is of particular concern for people with a severely weakened immune system and results in chronic diarrhoea and wasting. The parasites are also of economic importance globally and have knock-on effects on food security, as they cause major infections in aquaculture, and within honeybee populations.
Microsporidia are all around us; in the soil and the water. They are found in the environment as dormant spores – tiny, egg-shaped spores of only 1 µm in length (you could fit 1000 of them lined up along a 1 mm mark on a ruler). Animals can ingest these spores by eating or drinking contaminated food or water. This triggers an “awakening” of the spores and initiates a chain reaction of some truly extraordinary biology. When a spore reaches the gut of the animal, it “germinates”, meaning that it rapidly ejects a long, thin tube. Initially coiled up like a spring inside the spore, this polar tube rapidly everts from the spore, like a finger of a glove being turned inside out (Fig. 2). The tube connects with and pierces a host cell, and then all the infectious contents of the spore are squeezed down this tube and into the host cell. Once inside, the parasite feeds on the cell and reproduces within it to form more copies of itself, and eventually more spores. As more and more spores crowd the cell, they finally cause it to burst open with the release of the spores. These spores can then infect neighbouring cells or are passed through the host digestive system until they are expelled back into the environment.

Figure 2. Microsporidia-inspired artwork showing a) a secondary school pupil’s interpretation of a spore in clay photographed from different angles. The sculpture shows the polar tube coiled in the spore like a spiral. b) Another student’s clay sculpture showing a cross-section through a polar tube within a spore. c) Artist Jacky Oliver’s interpretation of polar tube germination and host cell infection. The metal artwork entitled “Coil and Spring” is installed prominently in our home institution - the Living Systems Institute at the University of Exeter. The inset shows a closeup of an ungerminated spore.
Looking into germination
The entire germination process is very quick. It only takes the spore around a quarter of a second to fire the polar tube and inject its infectious contents into the host cell. Our group decided to look into this process by electron cryo-tomography (cryoET), a cutting-edge bioimaging technique that uses powerful electron microscopes to image cells in 3D and at high resolution. Our microsporidian species of choice was Spragua lophii which infects monkfish but not humans, and is readily available from local fishmongers. It did cause a few funny looks when doing the weekly shop though, as we checked the monkfish out for the telltale signs of infection! We purified the spores from the fish and put them on an electron microscopy sample support grid before adding a solution of hydrogen peroxide or alternatively, human saliva, to germinate the spores. We then plunged these grids into liquid ethane. At a temperature of -160 °C, liquid ethane freezes biological material instantly, which prevents the formation of ice crystals, converting the sample into a “vitreous”, glass like state.
By recording multiple series of tilted views of these samples in the electron microscope, we were able to reconstruct 3D images, called tomograms, of the polar tubes. Looking through these tomograms, we could see that they contained cellular content in the process of transport. Some tubes were packed with ribosomes - the molecular machines that make proteins within cells. This allowed us to carry out sub-tomogram averaging where we could select all the ribosomes, average them, and generate a 3D map. We were excited to see that most of the ribosomes within the tube were found in pairs – joined together by a bridge of RNA and protein – and almost looking like they were holding hands (Fig. 3). We realised that this was a very important discovery, as such ribosome pairs have been seen before in bacteria, but not in eukaryotic organisms.
Intrigued about this finding, we set to solving the structure of this ribosome dimer. By combining sub-tomogram averaging with single particle analysis of ribosomes isolated from S. lophii spores, we were able to build an atomic model of the complex. In this way we could see where the interaction occurred between the two ribosomes within the pair, which also turned out to be interesting as it was different to any of the bacterial ribosome pairs previously seen. In our ribosome pairs, they interacted via the ribosome’s “beaks”, the function of which have so far been somewhat enigmatic.
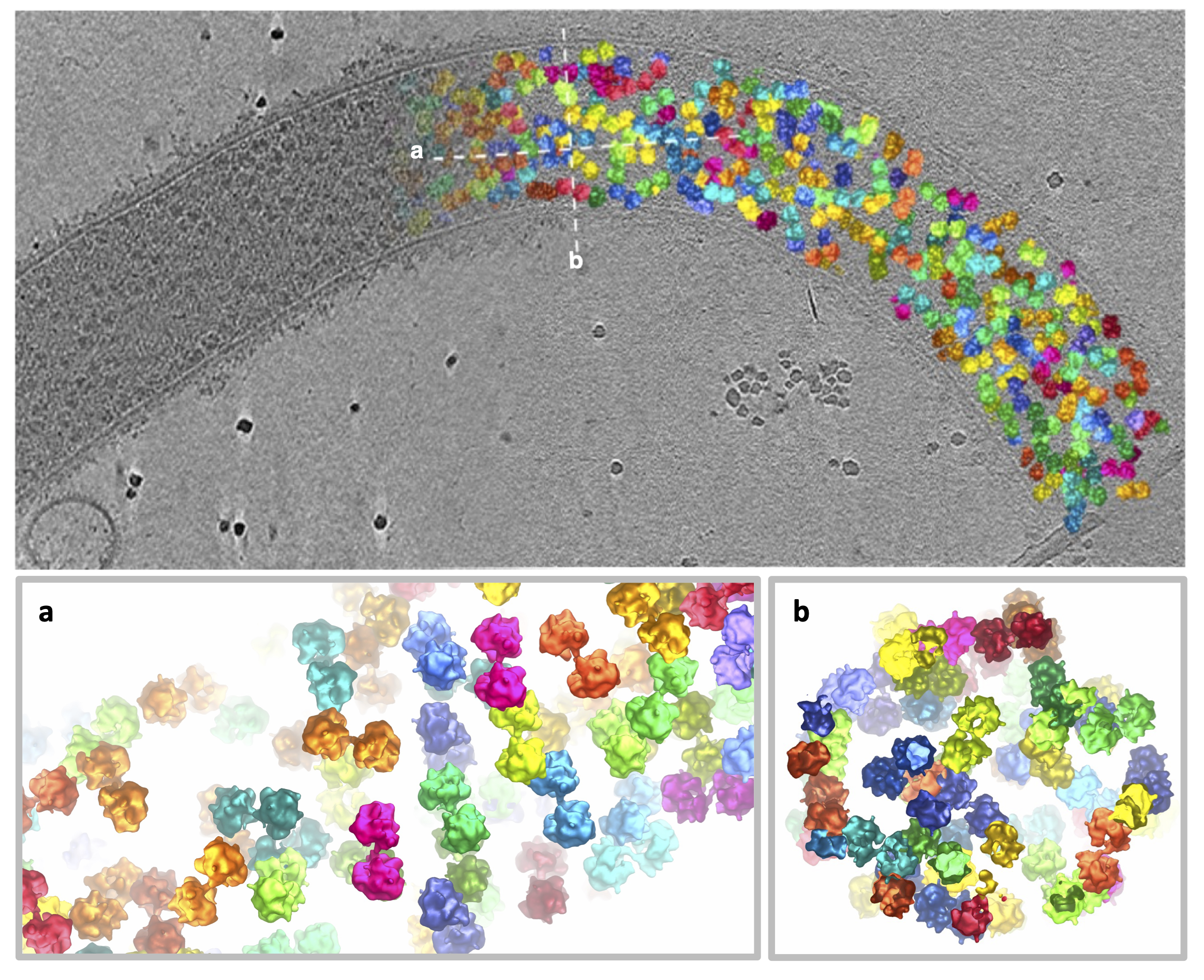
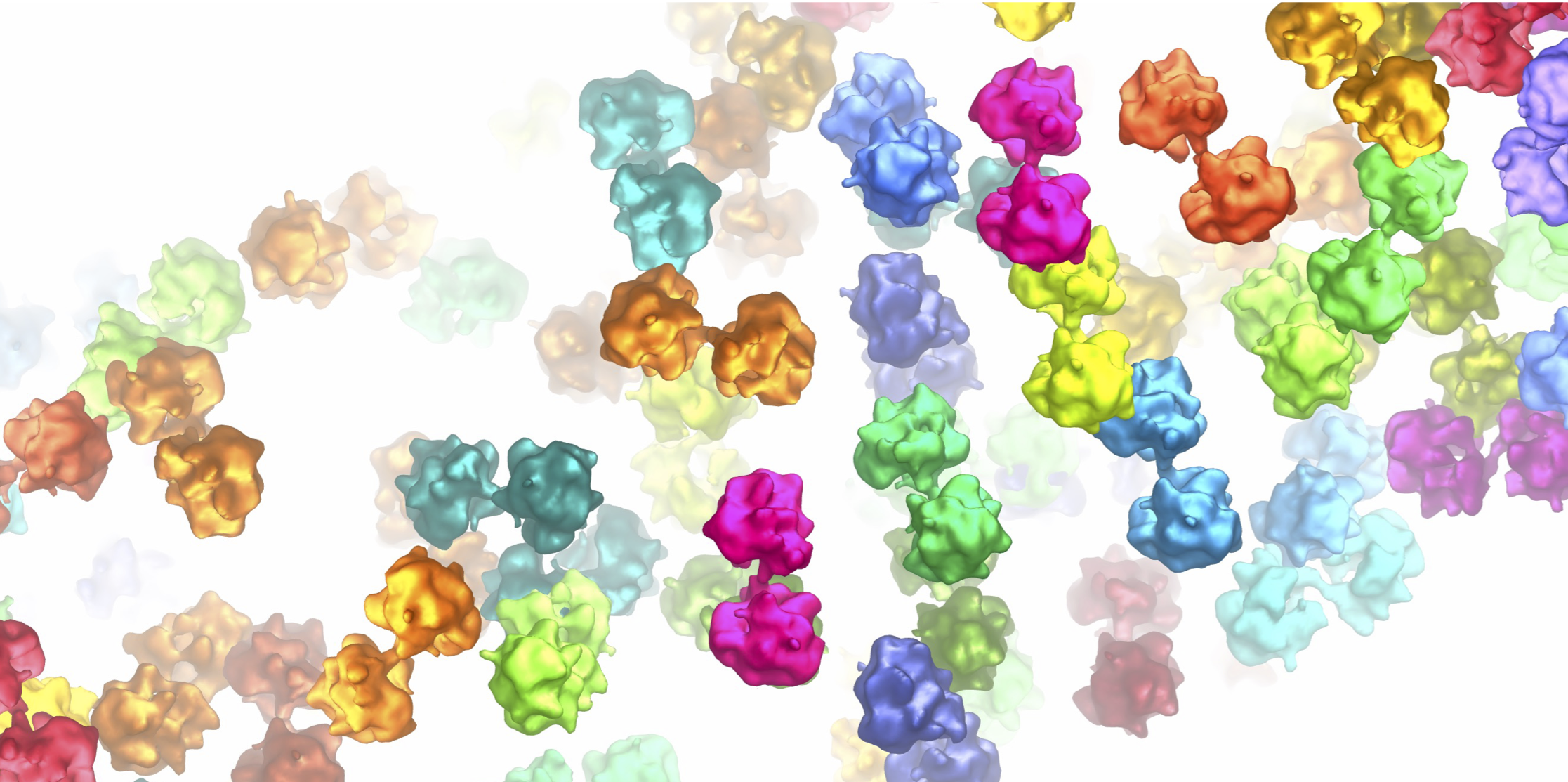
Figure 3. Ribosome pairs observed within the polar tube following germination of the spore. The top panel shows the original image (left-hand side) and has ribosomes coloured in on the right-hand side. The insets a and b show cross-sections through the polar tube as labelled in the top panel. Ribosomes are coloured in pairs.
Why pair up?
Why is it beneficial for ribosomes within microsporidian spores to form pairs? Ribosomes are found in pairs in bacteria when they are under stress conditions such as when there is a lack of nutrients. When ribosomes are in pairs they are inactive; they cannot carry out their usual function of making new proteins, but they remain fully assembled. This pairing up makes sense for the dormant spores, as they would not want to waste energy making proteins until required to, but they would then want to immediately start making proteins as soon as they infect a host cell. For this reason, we think that once the contents of the spores pass through the polar tube and into the host cell, the ribosome pairs split apart, so that the ribosomes can start making all the proteins that are required for the parasite to divide and make new spores. We found that 20 per cent of the ribosomes in the polar tubes do not form pairs. We think that these could either be the active ribosomes that maintain a baseline protein production activity necessary to keep the spores alive, or ribosomes that have already split from their partner, ready for action.
Microsporidia have much smaller ribosomes than other eukaryotes. Like many parasites, natural selection favoured those microsporidia that jettisoned all unnecessary ballast. The fact that microsporidia have retained the ability to create ribosome pairs suggests that this mechanism of silencing protein production is an essential one for the parasite, and provides further evidence of the importance of ribosome pairing as a key mechanism for controlling protein production in eukaryotes and bacteria.
Read our paper here for more detail.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in