Unraveling bacterial interactions during conjugation
Published in Microbiology

I first met Josh in the Spring of 2018. He had just started investigating antibiotic resistance mechanisms in Klebsiella pneumoniae (KP) and was specifically interested in isolates from the globally pervasive sequence type, ST258. These strains employ a combination of strategies which confer resistance to carbapenems, an important last line antibiotic. Josh described how structural modifications which constrict the channel of the major outer membrane (OM) porin, OmpK36, limit carbapenem entry into the cell but come with an associated fitness cost1. ST258 strains also often harbour plasmids encoding enzymes which inactivate carbapenems. One such plasmid is pKpQIL, a conjugative plasmid which is transferred via a system similar to that encoded by the well-known F plasmid.

hunched over the lab transilluminator as she
counts fluorescent transconjugant colonies.
Photo credits to Joshua Wong.
While attempting to conjugate pKpQIL into different strains of KP, he noticed that the plasmid transfer efficiency into strains expressing the constricted isoform of OmpK36 was consistently lower. Understanding why, he suggested, could form the basis of a project. After establishing a system to evaluate conjugation efficiency, I validated his chance observation and found that this reduction in plasmid uptake was more than 100-fold compared to strains expressing the non-constricted OmpK36 isoform from a laboratory strain of KP. This was an exciting finding as the involvement of a recipient OM protein during conjugation had only been shown once before – OmpA during F plasmid conjugation.
In their 1998 paper which served as a guiding light throughout this project, Klimke and Frost showed that OmpA dependency was associated with the F plasmid encoded conjugation protein, TraN2. They did so by substituting the endogenous traN with traN from a related plasmid, R100-1. TraN is required for mating pair stabilization (MPS), a process which leads to the intimate attachment of donor and recipient cells and greatly enhances conjugation efficiency. By performing a similar experiment, we showed that the pKpQIL-encoded TraN mediates the previously observed dependency on OmpK36.
Having identified a novel TraN-OM protein pair, we surmised that different TraN variants mediate MPS via a different recipient OM protein. To prove this, we sought to identify the receptor for TraN from R100-1. With dozens of potential OM protein candidates, it seemed the only way forward would be to perform an extensive knockout screen. Around the same time, I was assigned an undergraduate student to supervise during a 2-week lab attachment. These lab attachments would have typically lasted for 10 weeks but due to COVID were turned into hybrid projects with a greater emphasis on data analysis. So, I tasked Tatiana with selecting a candidate receptor OM protein to delete and characterize during her short stint in the lab. The fact that she selected, out of all the potential OM proteins, the actual receptor for TraN from R100-1 (OmpW) still feels nothing short of miraculous.
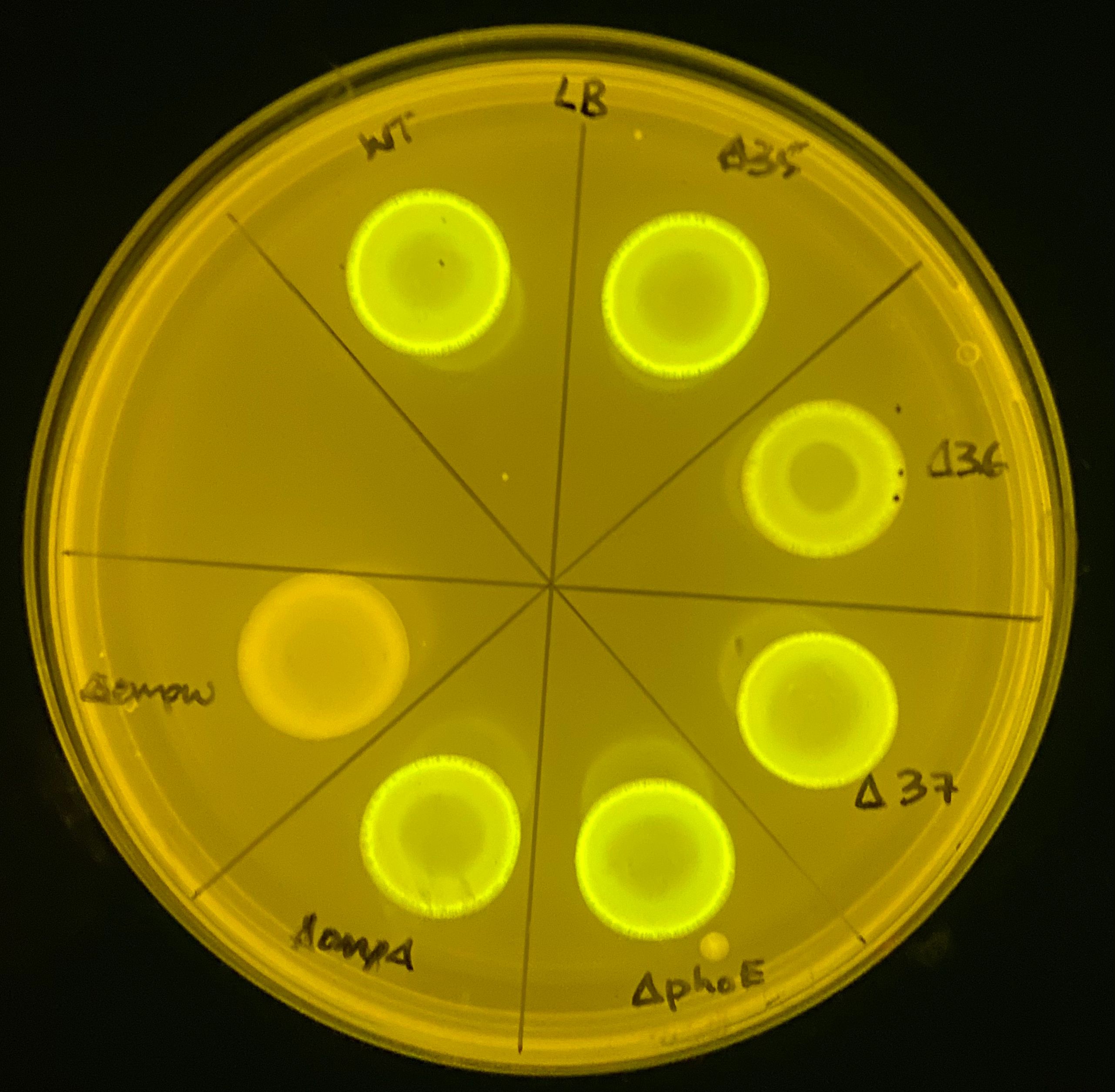
screen for potential TraN receptors. The fluorescent
patches indicate where efficient plasmid transfer has
occurred. Shown here is the initial screen performed to
determine the receptor for TraN from the pSLT plasmid.
Understanding the structural basis of TraN specificity proved to be a larger challenge. For decades, attempts had been made to predict the structure of TraN from F with little success. As fate would have it, just as we were looking to answer this question, news started circulating of a revolutionary new protein structure prediction software3. It is difficult to imagine what this project would have become had AlphaFold not existed. Not only did it provide us with valuable structural insights explaining how different TraN variants recognize different receptor proteins, but it aided in solving the last part of the MPS puzzle.
Despite the genetic evidence suggesting that TraN variants and their respective receptors interact during MPS, there has never been direct evidence to support this. That is, until Chloe managed to purify TraN from pKpQIL in complex with OmpK36, finally elucidating the mechanism behind MPS. Unfortunately, when Leti went on to try and determine the structure by cryo-EM, things were complicated by the fact that the majority of TraN is highly flexible. Luckily, density found within one of the monomers of the OmpK36 trimer aligned perfectly with part of the structure which had been predicted using AlphaFold enabling us to describe how these proteins interact.
The specificity of structurally similar TraNs for different OM receptor proteins was found to result in a species-specific effect during conjugation. Thus, we questioned the implications of this on the real-world distribution of plasmids. Interestingly, we saw that plasmid distribution within different bacterial species closely mirrored the specificity we observed in vitro. This suggests that although MPS is not essential for conjugation to occur, the increased efficiency of transfer that results from a productive interaction may be sufficient to influence plasmid host range.
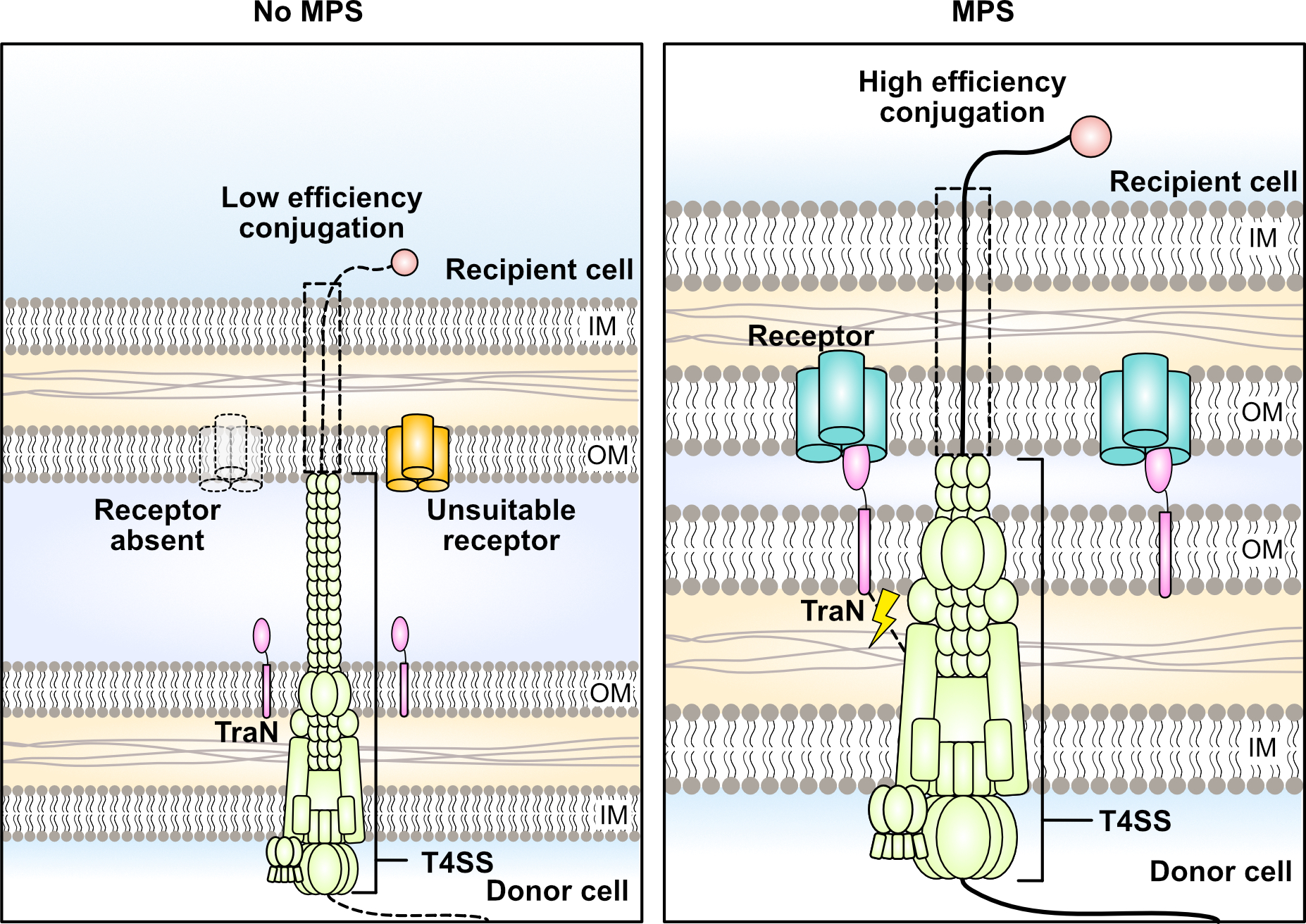
This begs the question, is the specificity associated with TraN a strategy used by conjugative plasmids to enter a preferred recipient population? Does this mean that plasmids encoding TraNs which facilitate efficient conjugation into a broad range of recipient species are disadvantaged by their own promiscuity? On the flipside, are the recipients being driven to evolve to influence their chances of picking up a particular plasmid? Species specificity certainly is not limited to conjugative elements which express TraN. Indeed, recent work is beginning to uncover alternative mechanisms which appear to influence the transfer range of other mobile genetic elements4,5. These are all exciting developments in the field of conjugation and promise to make clearer this complex and highly important process in bacterial evolution.
References
- Wong, J. L. C. et al. OmpK36-mediated Carbapenem resistance attenuates ST258 Klebsiella pneumoniae in vivo. Nat. Commun. 10, 3957 (2019).
- Klimke, W. A. & Frost, L. S. Genetic analysis of the role of the transfer gene, traN, of the F and R100-1 plasmids in mating pair stabilization during conjugation. J. Bacteriol. 180, 4036–4043 (1998).
- Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nat. 2021 5967873 596, 583–589 (2021).
- Allard, N., Neil, K., Grenier, F. & Rodrigue, S. The type IV pilus of plasmid TP114 displays adhesins conferring conjugation specificity and is important for DNA transfer in the mouse gut microbiota. Microbiol. Spectr. 10, e02303-21 (2022).
- Bean, E. L., Herman, C., Anderson, M. E. & Grossman, A. D. Biology and engineering of integrative and conjugative elements: Construction and analyses of hybrid ICEs reveal element functions that affect species-specific efficiencies. PLOS Genet. 18, e1009998 (2022).
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in