Urine cell-free DNA multi-omics to detect MRD and predict survival in bladder cancer patients
Published in Cancer
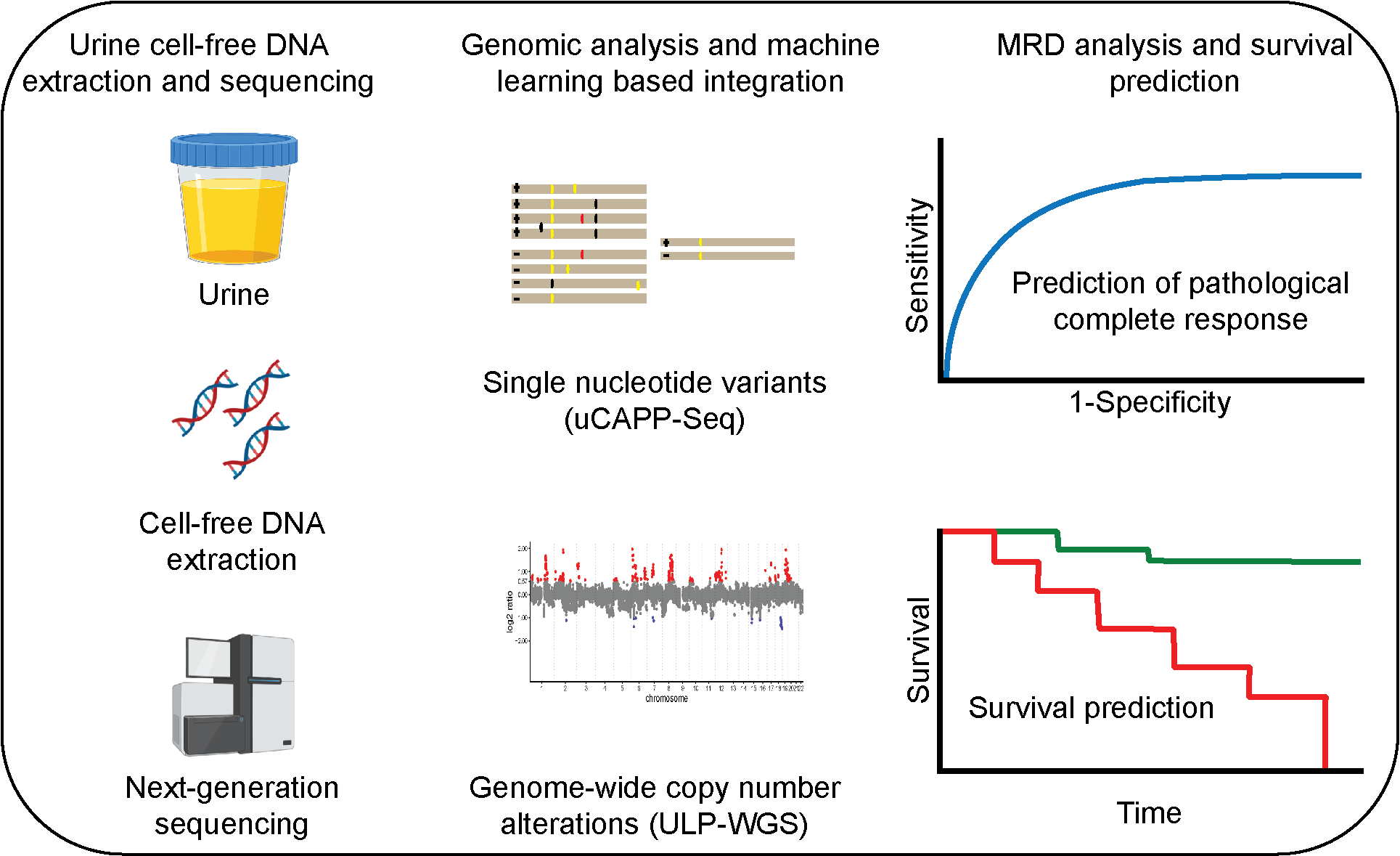
Urine CAPP-Seq applied to muscle-invasive bladder cancer
Liquid biopsy has shown promise as a noninvasive alternative to cystoscopy in the diagnosis and surveillance of localized bladder cancer1-3. Although liquid biopsy can be performed with a multitude of biofluids including plasma, urine is particularly compelling for bladder cancer minimal residual disease (MRD) detection as it is readily obtainable and in direct contact with the tumor. Our group recently showcased the application of a liquid biopsy assay (uCAPP-Seq) based on the detection of single nucleotide variants (SNVs) in urine cell-free DNA from patients with muscle-invasive bladder cancer (MIBC) undergoing radical cystectomy4. Assay positivity corresponded with the presence of residual cancer in the surgical specimen and a higher risk for disease relapse despite curative-intent treatment. Despite these compelling results, it may be challenging to maintain a high enough sensitivity to forego the risk of false negatives in clinical practice using a purely SNV-based approach to detecting tumor DNA in the urine.
Integrative multi-omic analysis of urine cell-free DNA detects MRD and predicts survival
Here, we built upon our previous study by using a multi-omic approach of combinatorial ultra-deep targeted sequencing and ultra-low-pass whole genome sequencing (ULP-WGS) to sensitively detect MRD in urine and predict survival after curative-intent radical cystectomy. We hypothesized that improved residual disease detection and prognostic power gained from urine cell-free DNA would allow for better risk stratification and more precise identification of a subset of lower risk patients who may be able to forego radical cystectomy altogether and instead opt for bladder-sparing treatment in the future.
We thus prospectively profiled urine samples from 74 patients with localized bladder cancer, of whom 78% had MIBC and 16 (22%) harbored treatment-refractory high-risk NMIBC. Patients without pathologic complete response (no pCR) had significantly higher variant allele frequency (VAF), inferred tumor mutational burden (iTMB), and copy number-derived tumor fraction levels (TFx) in urine cell-free DNA (cfDNA) compared to patients with pCR on pathological analysis of the final surgical specimen, which included bladder, prostate, and pelvic lymph nodes. Our study further confirmed that mutant DNA levels are enriched in urine compared to plasma. Strikingly, when performing CAPP-Seq on paired urine and plasma, tumor detection was more sensitive and specific from urine for detecting MRD and predicting pCR. This is in line with some of our other work suggesting that more proximal biofluids are enriched for tumor DNA compared to more distal biofluids5. Genome-wide analysis of urine cfDNA revealed focal copy number alteration of genes previously reported by The Cancer Genome Atlas (TCGA) to be recurrently altered in MIBC, with PPARG, ZNF703 and E2F3 being the most frequently amplified6,7. Further, uCAPP-Seq analysis of SNV data from our 74-patient cohort revealed that the TERT promotor and TP53 were the most commonly mutated genes in our study, again consistent with prior tumor next-generation sequencing data6-8.
To further assess the predictive power of these urine cfDNA-derived metrics, we integrated VAF, iTMB and TFx with pretreatment clinical variables within a random forest machine learning model that was subsequently validated via leave-one-out cross validation (LOOCV). The combinatorial urine tumor DNA metric, included in the model as the root product of maximum VAF, iTMB and TFx, was by far the most predictive feature, and area under the curve (AUC) for predicting pathologic complete response was 0.80 (P < 0.0001), demonstrating strong model performance. We obtained similarly strong results when training and cross-validating a random forest model composed of only urine cfDNA metrics. Strikingly, patients predicted by our model to harbor MRD also had significantly worse progression-free survival (HR = 3.00, P = 0.01) and overall survival (HR = 4.81, P = 0.009), comparable to the presence of residual disease in the radical cystectomy specimen itself. Univariate and multivariate Cox proportional hazards models confirmed the significance of our MRD predictions. Our integrative urine cell-free DNA approach also predicted survival outcomes in different patient subgroups and in a held-out validation cohort, further demonstrating its prognostic and predictive potential.
Take-home message and future directions
As we embark into an era of personalized precision medicine, it is imperative that we are able to risk-stratify bladder cancer patients accurately and offer customized treatments with the potential to lower treatment morbidity while maximizing survival. With this in mind, the outcomes of our study suggest that urine liquid biopsy using an integrative approach could be used to not only detect MRD with high sensitivity, but also to precisely risk-stratify patients before and after radical cystectomy. Our technology could form the basis for bladder-sparing treatment in select patients, allowing them to forego the morbidity of radical cystectomy altogether. Our integrative liquid biopsy approach could also potentially be applied to bladder cancer patients treated nonoperatively with chemoradiation, to improve patient selection and response monitoring. Future work should focus on proving these concepts and testing multi-omic urine liquid biopsy approaches in multi-institutional prospective clinical trials to more precisely guide clinical decision-making.
References
- Chaudhuri, A.A. et al. Emerging Roles of Urine-Based Tumor DNA Analysis in Bladder Cancer Management. JCO Precis Oncol 4(2020).
- Dudley, J.C. et al. Detection and Surveillance of Bladder Cancer Using Urine Tumor DNA. Cancer Discov 9, 500-509 (2019).
- Christensen, E. et al. Early Detection of Metastatic Relapse and Monitoring of Therapeutic Efficacy by Ultra-Deep Sequencing of Plasma Cell-Free DNA in Patients With Urothelial Bladder Carcinoma. J Clin Oncol 37, 1547-1557 (2019).
- Chauhan, P.S. et al. Urine tumor DNA detection of minimal residual disease in muscle-invasive bladder cancer treated with curative-intent radical cystectomy: A cohort study. PLoS Med 18, e1003732 (2021).
- Pellini, B. et al. ctDNA MRD Detection and Personalized Oncogenomic Analysis in Oligometastatic Colorectal Cancer From Plasma and Urine. JCO Precis Oncol 5(2021).
- Cancer Genome Atlas Research, N. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315-22 (2014).
- Robertson, A.G. et al. Comprehensive Molecular Characterization of Muscle-Invasive Bladder Cancer. Cell 174, 1033 (2018).
- Pietzak, E.J. et al. Next-generation Sequencing of Nonmuscle Invasive Bladder Cancer Reveals Potential Biomarkers and Rational Therapeutic Targets. Eur Urol 72, 952-959 (2017).
Follow the Topic
-
npj Precision Oncology
An international, peer-reviewed journal committed to publishing cutting-edge scientific research in all aspects of precision oncology from basic science to translational applications to clinical medicine.
Related Collections
With Collections, you can get published faster and increase your visibility.
Minimal Residual Disease and Circulating Tumor DNA Dynamics in Personalized Cancer Treatment
Publishing Model: Open Access
Deadline: Mar 12, 2027
Genomic Instability
Publishing Model: Open Access
Deadline: Jun 24, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in