Using DNA Nanostructures to Organize Insulin Molecules
Published in Bioengineering & Biotechnology

Our team of researchers from the Karolinska Institute in Stockholm, Sweden has recently published an article in Nature Nanotechnology, which showcases our work in producing user-defined, customizable insulin peptide nanoclusters. This paper is the culmination of over four years of work on a project that involved team members with a wide range of scientific backgrounds. It demonstrates for the first time that biological systems’ responses to insulin through the insulin receptor can be tuned through the careful spatial organization of the insulin peptide.
The insulin receptor signalling pathway is a major component of the body’s metabolism, helping to regulate blood sugar and lipid levels, and to stimulate various anabolic, biosynthetic processes 1,2. Misregulation of the pathway is associated with the development of diseases such as diabetes. Our work began with the question of whether or not the insulin receptor has some sort of organization on the surface of the cell. Previous projects in the lab had studied several different membrane proteins 3,4 and found various degrees of clustering and oligomerization. After reading through papers spanning several decades, all hinting at possible insulin receptor cluster formation but lacking the tools to test whether this was in fact the case 5,6, we decided to investigate ourselves, first by looking for insulin receptor clusters with state-of-the-art imaging techniques, and then by probing the insulin receptors in the membrane with tools developed through the techniques of DNA nanotechnology.
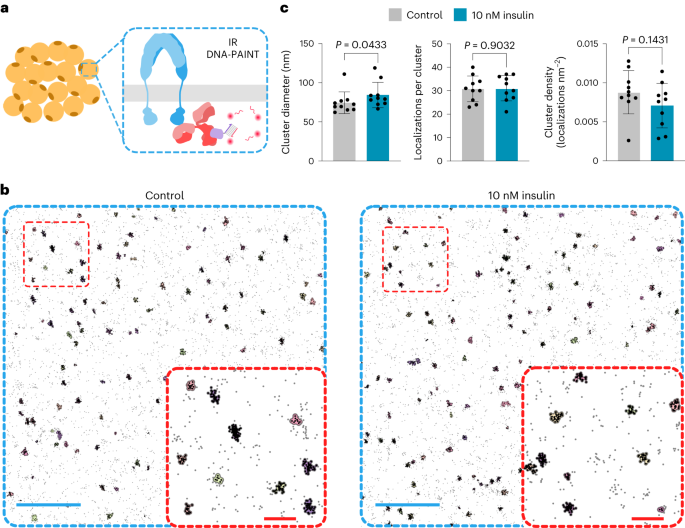
Long days (and nights) spent at the microscope and at the bench led us to our first conclusions. Through DNA-PAINT imaging 7 Georges Kiriako conclusively demonstrated the existence of insulin receptor clusters in the membranes of fat cells with diameters on the order of 10s of nanometres. In parallel, I had developed a protocol for tagging insulin peptides with a DNA strand, and Elena Ambrosetti had shown using SPR that these DNA-tagged insulins could bind the insulin receptor. Having a DNA tag on the insulin peptide opened up a whole new world of possibilities for the generation of insulin nanoparticles, allowing us to take advantage of the nanometre-level precision available in designs based on the DNA origami approach 8.
Several DNA origami nanostructures were constructed and tested, but in the end we settled on a simple rod-like design with dimensions similar to those of the insulin receptor clusters we had observed. This design gave us a ‘molecular pegboard’ to work from, allowing us to position our DNA-tagged insulin at specified sites with nanometre precision. Working from this pegboard, a series of structures was prepared with one to fifteen insulin peptides attached at a range of spacings (Fig. 1A). Using this palette of ‘insulin NanoRods’ as we called them, we aimed to test how binding to and activation of the insulin receptor changed when presented with insulin in various spatial arrangements.
Handing over the structures to Elena Ambrosetti, our SPR specialist, she came back with some interesting trends. Structures with multiple insulins on them bound and stuck around for much longer on the insulin-receptor-coated surface than a single, monomeric insulin peptide did. In fact, our structure with seven insulins on it, with neighbouring insulins spaced approximately 15 nanometres apart, remained bound over 100 times longer than monomeric insulin did (Fig. 1B). Importantly, we also saw a clear and continuous trend of binding strengths as the number of insulins increased, meaning that we could tailor the binding strength simply by tweaking the design of the underlying DNA nanostructure.
Coupled with these assays to test the binding of our structures to the insulin receptor, we also tested if they could activate the insulin receptor signalling pathway. José Dias performed countless western blots to suss out the trends in activation and came back to us with a very reassuring result. The structures that we saw bound best via SPR also most strongly activated the insulin receptor (Fig. 1C). There was a direct match in our trends for binding and for activation.
Excited by these findings, we sought to dig a little further to see what downstream effect our structures were having on cells. Were they activating the same pathways that normal, monomeric insulin was? Enya Engström took charge at this point, performing a series of RNA sequencing experiments to determine the transcriptional output of cells that had been treated with our structures. Analyzing the output showed that 91.5% of the genes that insulin stimulation normally affected were also affected by our seven-insulin-containing structure. This big overlap in the sets of affected genes indicated to us that our structures were not having spurious side-effects. They were targeting the same pathways that standard, monomeric insulin was. Importantly though, it took a much smaller concentration of insulin to achieve these changes in transcription with our structures compared to standard, monomeric insulin. They induced a much stronger transcriptional response at low insulin concentrations.
We were pleased with our results yet we were still left with a big question. Would our structures actually have an effect in a real, living organism? To test this we turned to Christina Kolonelou, our resident expert working with zebrafish larvae, a perfect model system with which we could test our structures. Working with a line that had had its pancreatic beta-cells destroyed (and hence its free glucose levels left uncontrolled), she injected our structures to see if they could control the rising glucose levels. The answer in short was yes, our structure with seven insulins on it could tame the rising levels of glucose just as normal, monomeric insulin did (Fig. 1D). So even with all the extra challenges that a living organism presented, our insulin-DNA nanostructures did the job, functioning as we hoped they would. For us this was really the cherry on top of over four years of hard work!
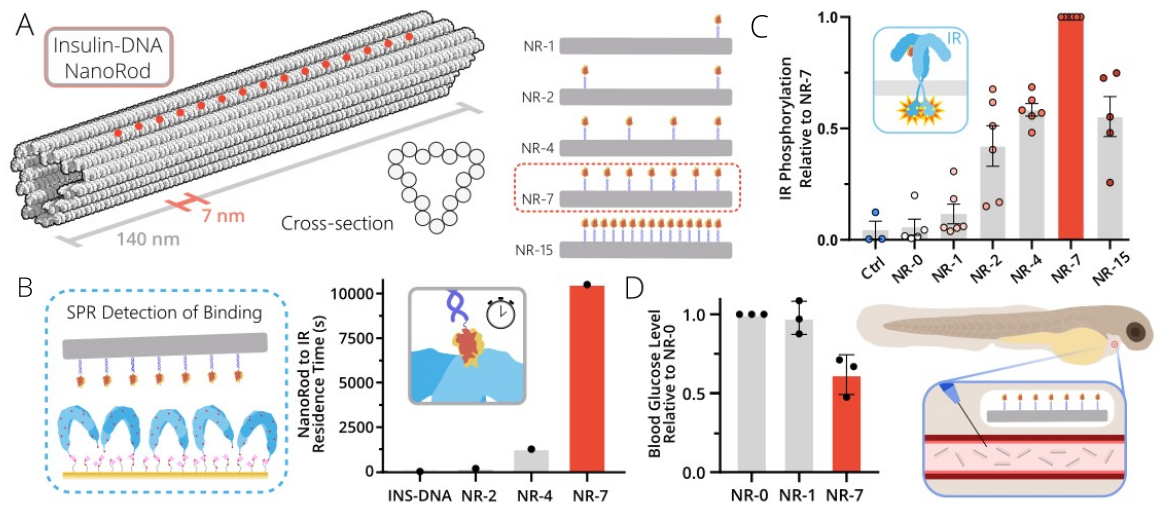
Fig. 1 | Multivalent insulin receptor activation using insulin-DNA origami nanostructures. A) A rod-like DNA origami was designed, allowing for the placement of insulin peptides at specific sites along the structure (red circles). B) Binding of the structures to the insulin receptor was investigated using SPR. Residence times of the structures to the insulin-receptor-coated surface increased with increasing numbers of insulin attached. C) Activation of the insulin receptor was determined via western blot. The highest activation was seen with our seven-insulin-containing structure. D) The structures were injected into zebrafish larvae which had had their pancreatic beta-cells destroyed. Our seven-insulin structure was able to significantly decrease the free glucose levels in the fish.
Taken together our results suggest that we can control insulin-receptor-mediated responses, not by altering the amount or chemical nature of the insulin peptide, but by controlling the spatial organization of insulin nanoclusters using an underlying DNA origami scaffold. We see this paving the way for new avenues of research on insulin receptor signalling and as the starting point in the development of new drugs for the treatment of diabetes.
References
-
Sims, E. K., Carr, A. L. J., Oram, R. A., DiMeglio, L. A. & Evans-Molina, C. 100 years of insulin: celebrating the past, present and future of diabetes therapy. Nat. Med. 27, 1154–1164 (2021). -
James, D. E., Stöckli, J. & Birnbaum, M. J. The aetiology and molecular landscape of insulin resistance. Nat. Rev. Mol. Cell Biol. 22, 751–771 (2021). -
Verheyen, T. et al. Spatial organization-dependent EphA2 transcriptional responses revealed by ligand nanocalipers. Nucleic Acids Res. 48, 5777–5787 (2020). -
Fang, T. et al. Spatial regulation of T-cell signaling by programmed death-ligand 1 on wireframe DNA origami flat sheets. ACS Nano 15, 3441–3452 (2021). -
Jarett, L. & Smith, R. M. The natural occurrence of insulin receptors in groups on adipocyte plasma membranes as demonstrated with monomeric ferritin-insulin. J. Supramol. Struct. 6, 45–59 (1977). -
Gustavsson, J. et al. Localization of the insulin receptor in caveolae of adipocyte plasma membrane. FASEB J. 13, 1961–1971 (1999). -
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014). -
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Follow the Topic
-
Nature Nanotechnology
An interdisciplinary journal that publishes papers of the highest quality and significance in all areas of nanoscience and nanotechnology.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in