Volatile-releasing nanosensors enable monitoring of respiratory disease via breath analysis
Published in Bioengineering & Biotechnology

While breath contains hundreds of volatile organic compounds (VOCs) arising in part from metabolic processes in the body, few VOCs of diagnostic value have been identified. Thus, there are currently only a handful of clinical breath tests that are used to detect and monitor disease. In the Bhatia Lab, we wanted to devise a way to leverage the practicality of breath tests to monitor lung health without being beholden to the VOCs available naturally in breath. Therefore, we sought to engineer breath biomarkers for respiratory disease. Through an established diagnostic platform in the lab – activity-based nanosensors (ABNs) – we have previously demonstrated that engineering urinary biomarkers is possible by leveraging the in vivo activity of proteases in diseased tissues.
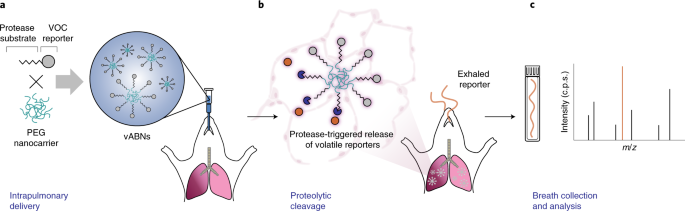
ABNs consist of mass-barcoded peptide substrates tethered to long-circulating nanoparticle carriers. After in vivo delivery, the surface-bound peptides are cleaved by disease-associated proteases, releasing barcoded peptide fragments small enough for renal clearance. The ABN platform has enabled urinary detection of several diseases in preclinical models1–4 and, in recent work, was delivered directly into the lungs to generate a multiplexed urinary signature for lung cancer5. Given this precedence, we envisioned a modified system in which ABNs would be delivered into the lungs to query for pathology-driving protease activity and release bio-orthogonal volatile reporters for detection via a rapid breath test (Figure 1a). While existing breath tests, such as the 13C-urea breath test6, utilize metabolism of small molecule substrates to induce a breath signal for disease, proteases represent a class of enzymes which, to our knowledge, have yet to be harnessed for breath-based diagnostics. This would involve expanding beyond small molecule substrates to macromolecular peptide substrates.
In our pursuit to develop volatile-releasing ABNs (vABNs), we (Melodi Anahtar, a graduate student; Kelsey Hern, a former technician/current graduate student at UC Berkeley; and myself from the Bhatia Lab) teamed up with scientists specializing in chemical detection (Drs. Roderick Kunz and Ta-Hsuan Ong from MIT Lincoln Laboratory). Rod’s group, historically focused on applications in homeland security, introduced us to instrumentation for vapor detection, and provided access to their in-house mass spectrometer7 that enabled the work described in our paper. This instrument provides high-throughput sample analysis via real-time quantification of VOCs. Thanks to this collaboration, we reached two pivotal milestones via a quick succession of experiments: (1) verification that we could engineer volatile release from peptide substrates, which were subsequently nanoformulated into vABNs for intrapulmonary delivery and (2) demonstration that we could achieve differential protease-driven breath signal between healthy and diseased mice within minutes after vABN delivery (Figure 1b). The latter was particularly memorable as we had painstakingly collecting up to 500cc breath per mouse due to concerns with sensitivity only to realize 12cc breath was enough to show elevated breath signal in the diseased cohort. That day, Melodi and I startled our collaborators with our loud cheering in front of the mass spectrometer.
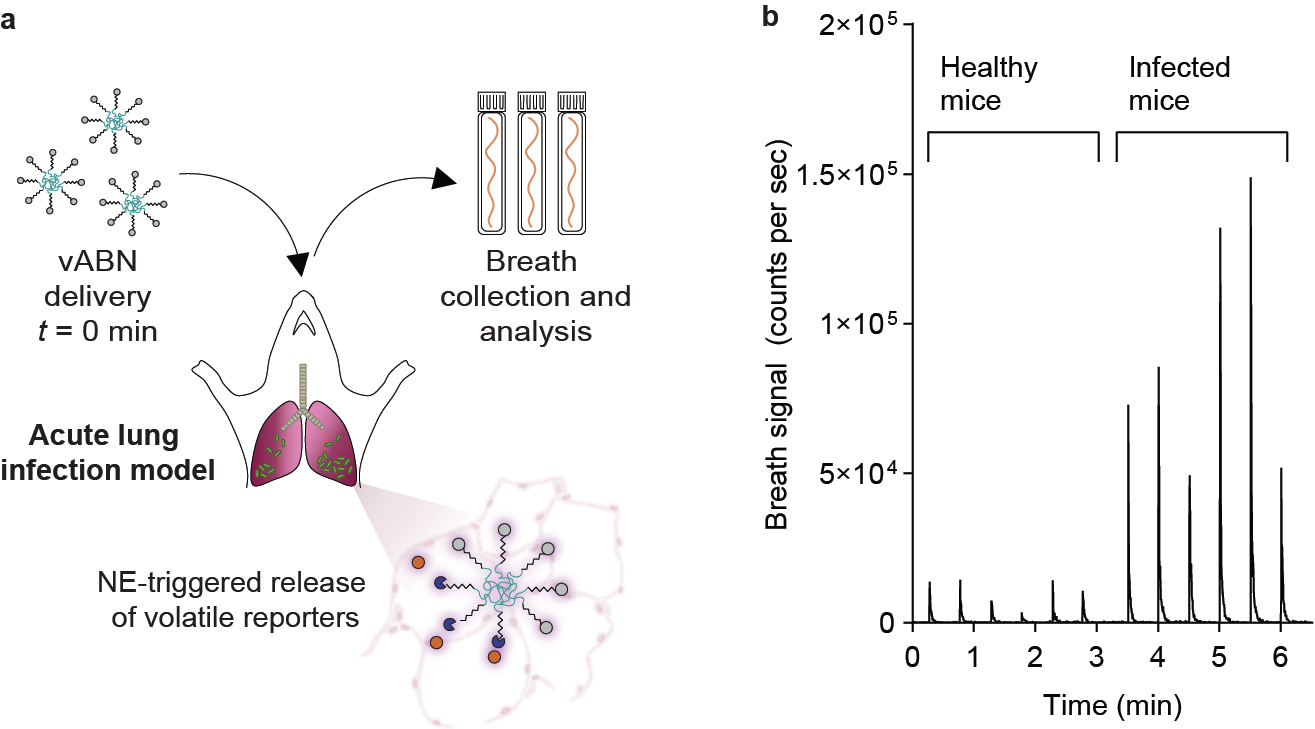
Figure 1. (a) Schematic of our approach. In our proof-of-principle, vABNs were engineered to detect neutrophil elastase (NE) activity, a protease secreted by neutrophils during lung infection. After intrapulmonary administration, vABNs are cleaved by NE, triggering the release of volatile reporters (orange) which are expelled in breath for subsequent quantification via mass spectrometry. (b) Quantification of reporters in 12cc breath from healthy and infected mice collected 10 min after vABN administration. Each signal peak represents one mouse (n = 6 per experimental group).
Reaching these milestones allowed us to then test vABN utility for monitoring lung disease. Our initial vABNs were designed for neutrophil elastase (NE), an inflammation-associated protease. We successfully used vABN-derived breath signal as a pharmacodynamic biomarker for protease inhibitor therapy against destructive NE activity in alpha-1 antitrypsin deficiency (mice were a gift from Christian Mueller’s Lab at U Mass). We are now expanding the vABN toolbox to detect other disease-relevant proteases and working on lower-cost downstream detection. Through this highly interdisciplinary work bringing together bioengineers, chemists, biologists and physicians, we have demonstrated that there are new avenues to engineer breath biomarkers which have the potential to be transformative for point-of-care diagnostics.
References
-
Kwong, G. a et al. Mass-encoded synthetic biomarkers for multiplexed urinary monitoring of disease. Nat. Biotechnol. 31, 63–70 (2013).
-
Kwon, E. J. & Dudani, J. S. Ultrasensitive tumour-penetrating nanosensors of protease activity. Nat Biomed Eng 1, 1–10 (2017).
-
Buss, C. G., Dudani, J. S., Akana, R. T. K., Fleming, H. E. & Bhatia, S. N. Protease activity sensors noninvasively classify bacterial infections and antibiotic responses. EBioMedicine 38, 248–256 (2018).
-
Loynachan, C. N. et al. Renal clearable catalytic gold nanoclusters for in vivo disease monitoring. Nat. Nanotechnol. 14, 883–890 (2019).
-
Kirkpatrick, J. D. et al. Urinary detection of lung cancer in mice via noninvasive pulmonary protease profiling. Sci. Transl. Med 12, 262 (2020).
-
Graham, D. Y. et al. Campylobacter pylori detected noninvasively by the 13C-urea breath test. Lancet 329, 1174–1177 (1987).
-
Ong, T. et al. Use of mass spectrometric vapor analysis to improve canine explosive detection efficiency. Anal. Chem. 89, 6482–6490 (2017).
Follow the Topic
-
Nature Nanotechnology
An interdisciplinary journal that publishes papers of the highest quality and significance in all areas of nanoscience and nanotechnology.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in