What's up your nose?
Published in Microbiology
We are interested in the contents of your nose, not at the level of hair, bogey and the occasional finger, but at the level of the complex microbial community that lives there and how it is associated with respiratory infection. The makeup of this community has been interrogated through sequencing (the airway microbiome) with indications that some bacterial communities may be associated with health and others with disease.
A bacterial diet
However, what the bacteria in the airways eat to survive is less well understood. One tool that may help us to characterise which biochemicals in the airways bacteria can use as food is called metabolomics. This uses liquid chromatography, to separate the biochemicals, and then mass spectrometry, to interrogate what they are. Comparing the mass spectrometry data against a curated library, we can then determine which individual biochemicals are present and their relative abundance. This tool has been used widely to investigate changes in the blood but has not been used much to interrogate the airway.
Blotting paper 2.0
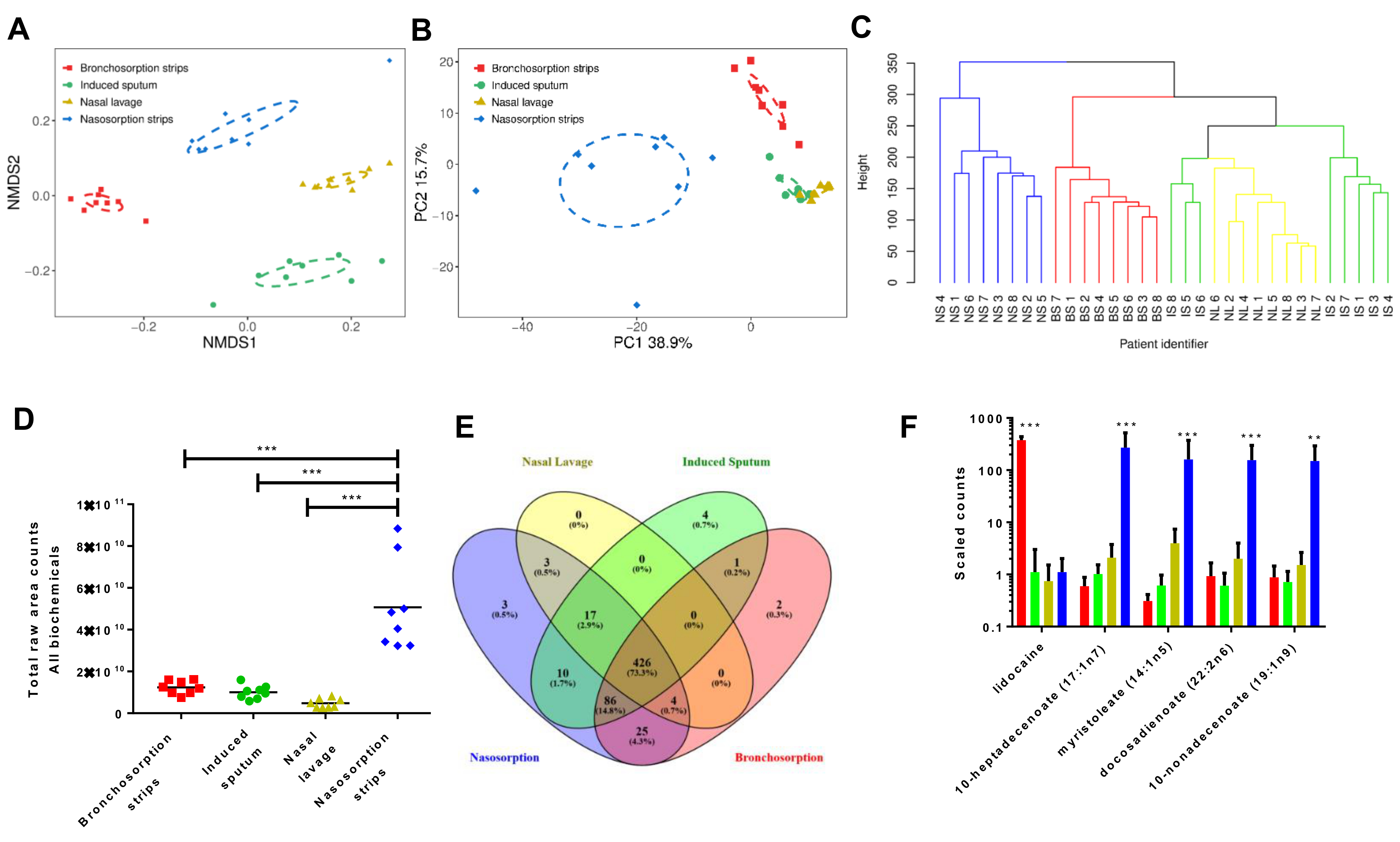
The aim of our recently published study was to compare methods for sampling the airway metabolome. We looked at two standard techniques – nasal lavage (flushing a millilitre of saline through the nose and recovering whatever you can – quite a lot never comes back!) and induced sputum (getting people to breathe in an expectorant and then spit in a cup). We also used a newer technique, that had never been used for metabolomics sampling, called Synthetic Absorptive Matrix (SAM) strips. These are hi-tech blotting paper and have been used to recover other types of sample from the airways, including antibodies and cytokines. You can watch a video of their inventor having them put up his nose here. We looked at the use of these SAM strips in both the upper airway (via the nose) and the lower airway (via a bronchoscope). In the traditions of Barry Marshall (though I doubt I will get a Nobel prize for this), I volunteered to be one of the subjects for the sampling; the nasal wash, induced sputum and upper airway SAM were all fine, but having a bronchoscopy was fairly unpleasant.
It’s good to share
Having collected the samples, we then outsourced the running of the metabolomics to a company, called Metabolon in the US. This choice had mixed reviews, but I think it is ok to outsource, increasingly labs are outsourcing some of the more specialist analysis approaches – sequencing, transcriptomics, metabolomics. This makes sense in terms of time, expertise and access to equipment. Specifically in the case of metabolomics, outsourcing gave us access to a much larger curated library of samples, giving us more information from our samples, the biochemicals were also grouped into families, enabling us to interrogate the data more easily.
Sooo much data
From the point of performing the study to publishing it has been a lengthy process. In part this was due to the complexity of the dataset. We had approximately 14,000 data points – which may be small compared to some types of project, but when you are used performing focussed studies on individual mediators it was quite a step change. This was combined with a bewildering list of biochemicals, most of which we had never heard of – 1-stearoyl-2-arachidonyl-GPC anyone? In the end, through the power of the R programming platform and a very talented PhD student, we have ended up with a paper that uses a wide range of graph types, all of which aimed to compress the data into a meaningful form.
Food for the Pseuds
So what did we find? In total, 581 biochemicals were recovered from the airways belonging to a range of different families. When we compared the relative abundance of the these biochemicals between the different sampling techniques, we saw that the SAM strips gave us a much greater recovery of biochemicals than the other approaches. Since we were interested in how the airway metabolome enables bacterial colonisation, we screened some of these biochemicals for their ability to support bacterial growth. 35 of these biochemicals were able to support growth of the opportunistic airway bacteria Pseudomonas aeruginosa, including a number of sugars and amino acids.
A microcosm in a nostril
The airways represent a fascinating ecosystem because they are nutritionally more restricted in terms of the range and specific concentrations of any one biochemical compared to say the gut, but at the same time the nutrients are constantly refreshed. The balance of biochemicals in the airways shapes the bacteria that can live there, and we believe that this could be dysregulated in disease. By developing the tools to sample the airway metabolome, we are now one step closer to understanding how changes in airway biochemistry affects infection.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Very interesting - the study and the methods-Dr Tregoning.
Probably-would be useful in detecting antibodies?
Mala Menon
maladmbcp@yahoo.co.in
You are right. We (and other groups) have used this methodology for nasal antibody sampling:
https://www.ncbi.nlm.nih.gov/pubmed/28647455