Why the bad is not always bad
Published in Microbiology

Only recently, we have recognised the importance of microbe-to-host interactions. While before, bacterial, fungal and viral interactions with our cells were not really taken into account, nowadays it is seen as common knowledge that we are not sterile organisms. Indeed, bacteria was already found in our blood and only recently it was hypothesised to be potentially found in the brain.1
Many microbes were associated with certain diseases but now we know that it is not the number of bacteria which differentiates the healthy from the non-healthy but the shift of a prevalence of certain strains. Therefore, we understood that treating these diseases with antibiotics can cause more harm than good. Usually, we cannot select for those we want to keep and those we want to get rid of and we end up with killing all. Also, the wrong use of antibiotics by its patients has led to the occurrence of antibiotic resistant strains which will survive any treatment and therefore become the prevalent strain.
In this sense, it seems a promising approach to identify a healthy mixture and use it to engraft on the host. The same mechanism was already successfully implemented in our gut. Take the bacteria of an “healthy” donor and transplant it in an “unhealthy” one. This approach got famous under the term “fecal microbiome transplantation” - in short FMT. Also, recently it was suggested that there might be a direct link between ageing and gut-microbiome composition. Researchers found that verrucomicrobia Akkermansia muciniphila, found in centenarians, had a lifespan-enhancing effect when transplanted into a mouse-model.2 Another example describing the importance of interactions to the gut microbiome was the finding that Veillonella atypica abundance is responsible for the enhancement of performance in athletes due to its ability to metabolize lactate into propionate.3
So, why not using this transplantation approach on another site on our human body, for example the skin? We swapped multiple patients and we identified their skin microbial community, mainly focusing on Cutibacterium acnes- a major skin commensal, living in the sebaceous glands of the human body. This bacterium was negatively associated with the skin condition acne vulgaris and was long seen as a pathogen. Today, we understand the necessity of C. acnes on the healthy human skin and so we tried to modulate a recipient microbiome with a donor microbiome. We saw that with a certain combination of strains we could reach transplantation significant engraftment on the human recipient. This modulation would stabilise during application and return slowly to its original state when the application was stopped.
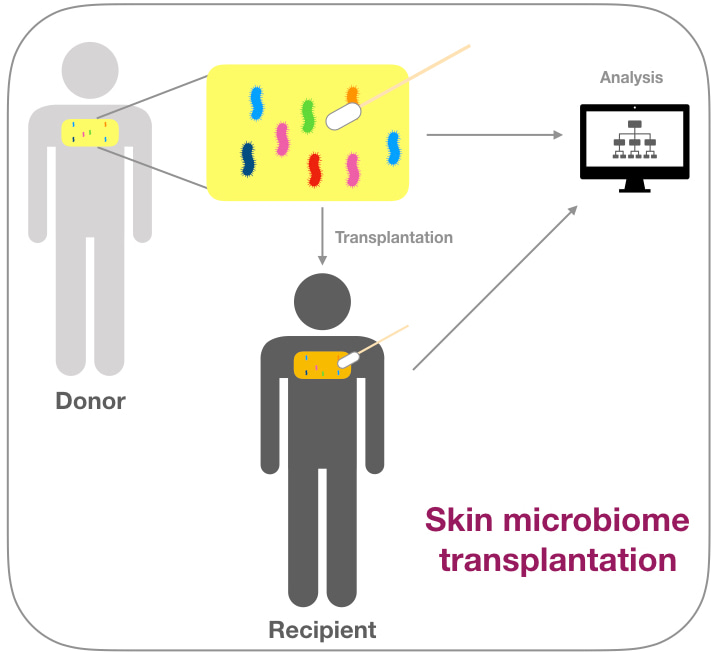
It is interesting to see how different populations can influence the health of the host. We should reconsider the way we see the human body and approach it as a whole and embrace the fact that we are inhabited by a community of organism which can contribute to our well-being.
2. Bárcena, C. et al. Healthspan and lifespan extension by fecal microbiota transplantation into progeroid mice. Nat. Med. (2019). doi:10.1038/s41591-019-0504-5
Follow the Topic
-
Microbiome
This journal hopes to integrate researchers with common scientific objectives across a broad cross-section of sub-disciplines within microbial ecology. It covers studies of microbiomes colonizing humans, animals, plants or the environment, both built and natural or manipulated, as in agriculture.
Related Collections
With Collections, you can get published faster and increase your visibility.
Animal Gut Nutrition and Greenhouse Gas Mitigation
Animal Microbiome, Journal of Animal Science and Biotechnology and Microbiome call for submissions to the collection on Animal Gut Nutrition and Greenhouse Gas Mitigation.
Efforts to reduce greenhouse gas emissions from livestock systems increasingly hinge on innovations in animal gut nutrition. The dynamic relationship between the gut microbiome and nutrient utilization plays a pivotal role in shaping methane output, feed efficiency, and overall sustainability. Advances in microbial ecology—particularly in understanding the role of gut microbiome in nutrient metabolism—are opening new pathways for mitigating emissions while enhancing productivity. These developments support the implementation of climate-smart agricultural strategies to address climate change and its impacts.
Looking ahead, continued research in this field has the potential to yield innovative solutions such as targeted probiotic supplementation, which could further optimize gut function and enhance nutrient absorption. These advancements may lead to reduced greenhouse gas emissions while improving animal health and productivity. By deepening our understanding of the animal gut microbiome, we can contribute significantly to sustainable agricultural practices that benefit both the environment and food security.
We invite researchers to contribute to this special Collection on Animal Gut Nutrition and Greenhouse Gas Mitigation. Topics of interest include but are not limited to:
- Animal Gut Microbiome and Feed Efficiency
- Greenhouse Gas Mitigation Strategies
- Rumen Fermentation Dynamics
- Nutrient Utilization in Livestock
- Probiotic Supplementation Effects
- Sustainable Livestock Production Practices
- Climate-Smart Agriculture Innovations
This Collection supports and amplifies research related to SDG 13, Climate action.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Animal Microbiome fees, Journal of Animal Science and Biotechnology fees, Microbiome fees). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Publishing Model: Open Access
Deadline: Sep 04, 2026
Oncobiome
This collection of papers delves into the burgeoning field of oncobiome research, exploring the intricate relationship between cancer and the microbiome. The oncobiome encompasses the diverse microbial communities residing in and on the human body, which influence cancer development, progression, and treatment responses. By examining these interactions, our aim is to unravel the complex mechanisms through which the microbiome impacts oncogenesis and therapeutic outcomes.
This compilation highlights cutting-edge research, offering insights into potential diagnostic markers and novel therapeutic strategies, thereby advancing our understanding of cancer biology and paving the way for innovative, microbiome-targeted cancer treatments.
This is a cross-journal collection between:
Experimental Hematology and Oncology
Articles will undergo the standard peer-review process of the journal to which they are submitted and are subject to either the BMC editorial policies or those of BJC Reports. Articles will be added to the Collection as they are published. The Editors have no competing interests with the submissions which they handle through the peer review process. The peer review of any submissions for which the Editors have competing interests is handled by another Editorial Board Member who has no competing interests.
Publishing Model: Open Access
Deadline: Ongoing





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in