Steering ecological-evolutionary dynamics to improve artificial selection of microbial communities
This is a video walkthrough of our recent theoretical work on how to improve the efficiency of artificial community selection.
Published in Ecology & Evolution
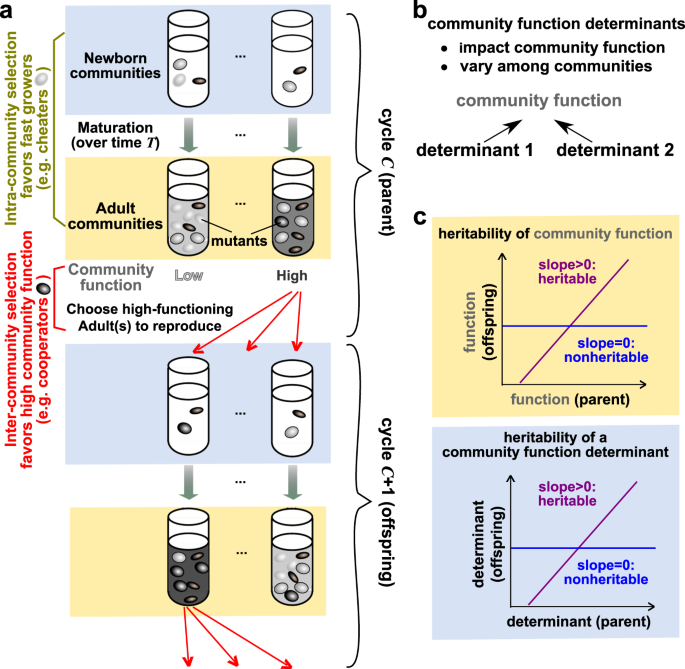
Microbial communities often perform important functions that depend on inter-species interactions. To improve community function via artificial selection, one can repeatedly grow many communities to allow mutations to arise, and “reproduce” the highest-functioning communities by partitioning each into multiple offspring communities for the next cycle. Since improvement is often unimpressive in experiments, we study how to design effective selection strategies in silico. In this video, we try to walk readers through our recent theoretical/simulation paper on how to improve the efficiency of artificial community selection. We will discussed how intracommunity ecological and evolutionary processes affect the heritability of community function, which in turn determines the dynamics and outcome of community selection. We then devise experimentally-implementable manipulations to enhance community function heritability, which speeds up community function improvement under various scenarios.
Follow the Topic
Ecology
Life Sciences > Biological Sciences > Ecology
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
A selection of recent articles that highlight issues relevant to the treatment of neurological and psychiatric disorders in women.
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
With this cross-journal Collection, the editors of Communications Biology, Nature Biomedical Engineering, Nature Sensors, Nature Communications, and Scientific Reports welcome the submission of primary research Articles focusing on the development of engineered biosensing devices with the potential to be applied in biomedical research and in the management of disease conditions.
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in