A multi-faceted effector: The RNA-dependent RNA polymerase of dengue virus
Published in Microbiology
Age cannot wither her, nor custom stale
Her infinite variety.
- William Shakespeare, Antony and Cleopatra, Act II, scene 2
The dengue virus (DENV) possesses an RNA-dependent RNA polymerase (NS5) that replicates its RNA genome besides capping the nascent RNA to enhance stability. Given its centrality to the propagation of DENV, it presents an attractive target for drug development and is also known to be highly immunogenic in humans. However, replication of the viral genome is by no means its sole function; it is also likely involved in mediating virus-host interactions, specifically by modulating host immune responses. The moonlighting functions of NS5 have been known for some time1-3, as has its nuclear localization in the case of DENV serotypes 2 & 44-6. Many research groups have sought to determine the human proteins that interact with NS5 using a variety of experimental approaches, which have been compiled in a review we published in 20217.
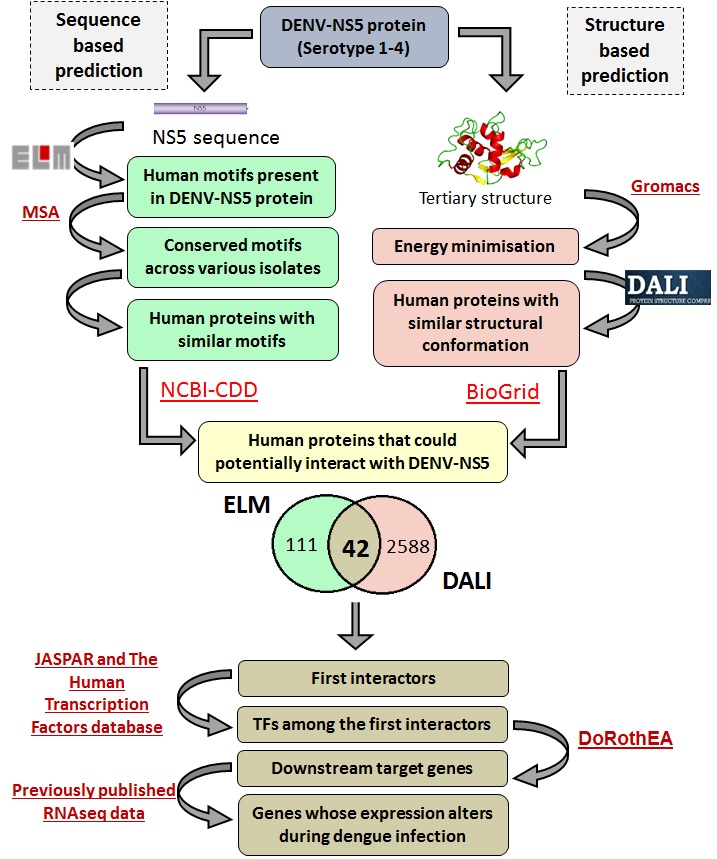
Given these antecedents, it was tempting to for us to make an attempt to computationally predict interactions with human proteins that could form the basis of testable hypotheses leading to a better understanding of the mechanisms underlying observed immunological effects of dengue infection. In our work published in 3 Biotech we have tried to make precisely such predictions starting with a conservative set of 42 human interactors that we identified8. Pathway analysis enabled us to focus on the subsequent interactors that were immunologically relevant. While predictions and speculations may seem trite, prior work, particularly the online availability of raw experimental data from RNA-seq experiments enabled us to validate some of our predictions. In other words, some of our predictions have already been borne out by experiment, giving us hope that other inferences we have made would be of value as well for other researchers, particularly immunologists.
DENV belongs to the family Flaviviridae that comprises arthropod-borne, enveloped, single-stranded RNA viruses such as Zika, Japanese encephalitis and yellow fever among others. As the most conserved of non-structural protein across all flaviviruses9, it seems likely that NS5 is a multi-functional effector that has many surprises in store for those seeking to combat vector-borne tropical viral diseases.
Note: Poster image reproduced from Fig. 1 of Bhatnagar et al8.
References:
1. Ashour et al. (2009). NS5 of Dengue Virus Mediates STAT2 Binding and Degradation. J Virol 83(11):5408 - 5418. doi: 10.1128/jvi.02188-08.
2. Petit et al. (2021). Nuclear dengue virus NS5 antagonizes expression of PAF1-dependent immune response genes. PLOS Pathogens 17(11): e1010100. doi: 10.1371/journal.ppat.1010100
3. Ledesma et al. (2023). Dengue virus NS5 degrades ERC1 during infection to antagonize NF-kB activation. Proc Natl Acad Sci USA 120(23):e2220005120. doi: 10.1073/pnas.2220005120
4. R. Bulich, J. G. Aaskov (1992). Nuclear localization of dengue 2 virus core protein detected with monoclonal antibodies. Microbiology 73(11): 2999-3003. doi: 10.1099/0022-1317-73-11-2999.
5. Tadano et al. (1989). Detection of Dengue 4 Virus Core Protein in the Nucleus. I. A Monoclonal Antibody to Dengue 4 Virus Reacts with the Antigen in the Nucleus and Cytoplasm. Microbiology 70(6): 1409-15. doi: 10.1099/0022-1317-70-6-1409.
7. Bhatnagar et al. (2021). Dengue Virus Non-Structural Protein 5 as a Versatile, Multi-Functional Effector in Host–Pathogen Interactions. Front Cell Infect Microbiol 11:574067. doi: 10.3389/fcimb.2021.574067.
8. Bhatnagar et al. (2023). Prediction of human protein interactome of dengue virus non-structural protein 5 (NS5) and its downstream immunological implications. 3 Biotech 13(6): 180. doi: 10.1007/s13205-023-03569-0
9. Grant et al. (2016). Zika Virus Targets Human STAT2 to Inhibit Type I Interferon Signaling. Cell Host Microbe 19(6): 882-890. doi: 10.1016/j.chom.2016.05.009.
Disclaimer: The opinions expressed herein do not represent the views of the TERI School of Advanced Studies or TERI.
Follow the Topic
-
3 Biotech
This is a hybrid journal that publishes results of the latest research related to the study and application of biotechnology to Medicine and Biomedical Sciences, Agriculture and the Environment.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in