A whole-cell biocatalyst for degrade the high-crystallinity poly(ethylene terephthalate) (hcPET)
Published in Bioengineering & Biotechnology
Poly(ethylene terephthalate) (PET) is a kind of plastic widely used in clothing, packaging, medicine, and other fields. Due to its strong physical and chemical stability, PET is highly resistant to natural degradation. A large amount of nondegradable PET waste accumulated in the environment and PET microplastics derived from larger one have posed a serious threat to global environmental security and human health. Therefore, more and more attention has recently been paid to the recycling of PET. The enzymatic hydrolysis of PET is a relatively new and developing technology, which combines some advantages of traditional physical and chemical degradation. Its core idea is to decompose PET into reusable industrial raw materials, to reduce or get rid of the use of raw materials made from petroleum. At present, many countries in the world have made a lot of strategic investments in this technology. Biodegradation of low crystallinity PET by biological enzymes has made great progress, and the application of some catalytic enzymes has reached the semi-industrial level. However, for the degradation of high crystallinity PET (hcPET) (used to make food-grade containers such as cola bottles and mineral water bottles), most of the degradation enzymes found so far are powerless.
Recently, a novel PET-hydrolyzing enzyme named PETase was identified from the bacterium Ideonella sakaiensis. This enzyme can degrade hcPET at a moderate temperature (30 °C). Once discovered, PETase has become a new research hot spot. A variety of PETase mutants have been designed to increase its degradation performance through different strategies. But so far, the degradation capacity of these mutants for hcPET is still relatively low. How to further increase the degradation efficiency of PETase towards hcPET? Recently, we constructed a whole-cell biocatalyst by displaying PETase on the surface of yeast cells to degrade hcPET. We, therefore, consider that it is possible to increase the degradation efficiency of the whole-cell biocatalyst by introducing an external adhesive unit into the surface display system to control the process of PETase-displaying cells adsorbing on the PET surface.
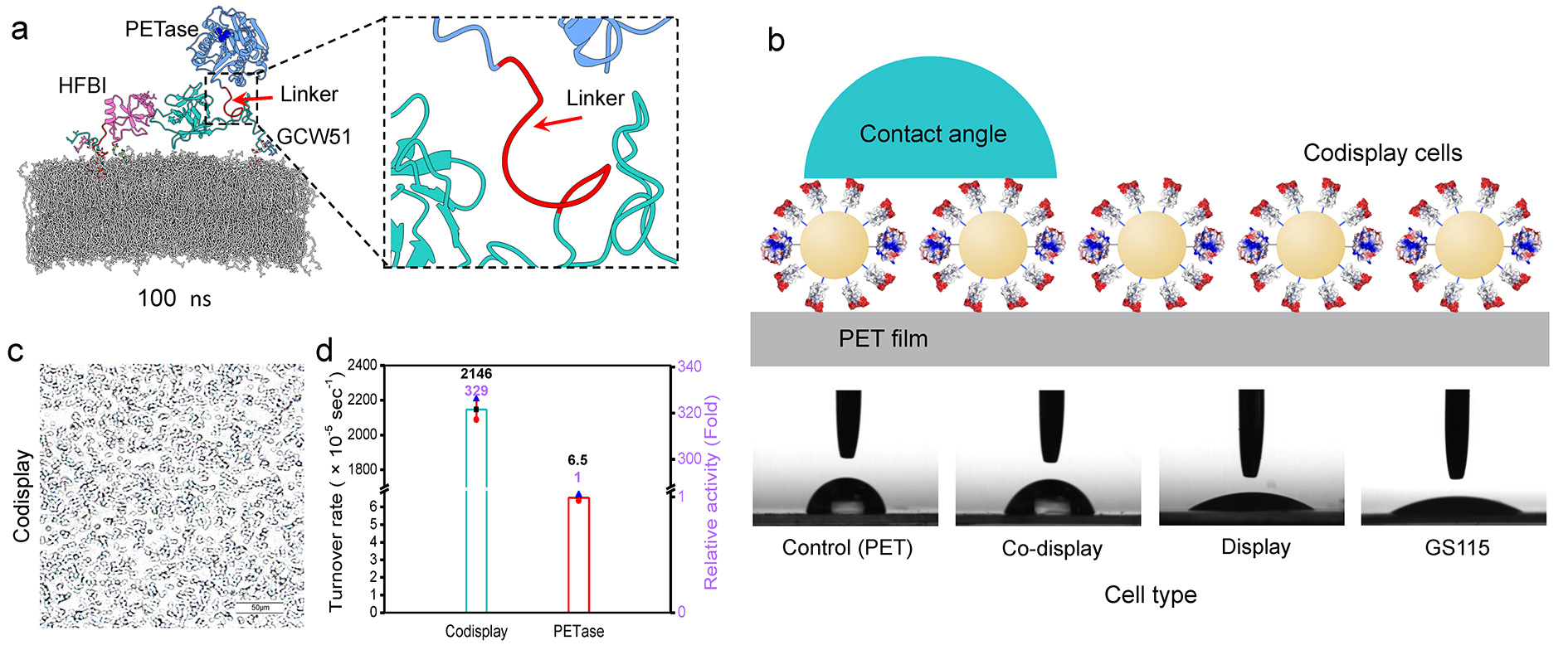
Figure 1. The yeast whole-cell biocatalyst. (a) Molecular dynamics simulation diagram of adhesive unit hydrophobin HFBI and degradation unit PETase codisplayed to yeast cell surface at 100 ns. (b) Schematic diagram of the WCA experiment and image of the WCA experiment. (c) The adsorption of the codisplay cells on the hcPET surface. (d) Comparison of the turnover rate of the codisplay cells and purified PETase under the optimal conditions using PET as a substrate.
In our study, we developed a codisplay system to mimic the natural two-step gradation process of PET by a whole-cell biocatalyst (Figure 1a). We found that both PETase and HFBI were functionally displayed on the yeast cell surface. The displayed HFBI could profoundly increase the hydrophobicity of the yeast cells (Figure 1b), and the attachment of codisplayed cells onto the PET surface was improved (Figure 1c). This codisplay system showed approximately 328.8-fold higher degradation efficiency than that of the native PETase toward the hcPET (crystallinity of 45%) at 30 ℃ (Figure 1d). The corresponding conversion level for hcPET was approximately 10.9% at 30 ℃ within 10 days. Finally, based on the molecular dynamics simulation, we proposed the molecular mechanism of the whole-cell biocatalyst to degrade hcPET, that is, the introduction of the adhesion module is the key to the efficient degradation of hcPET in this system (Figure 2).
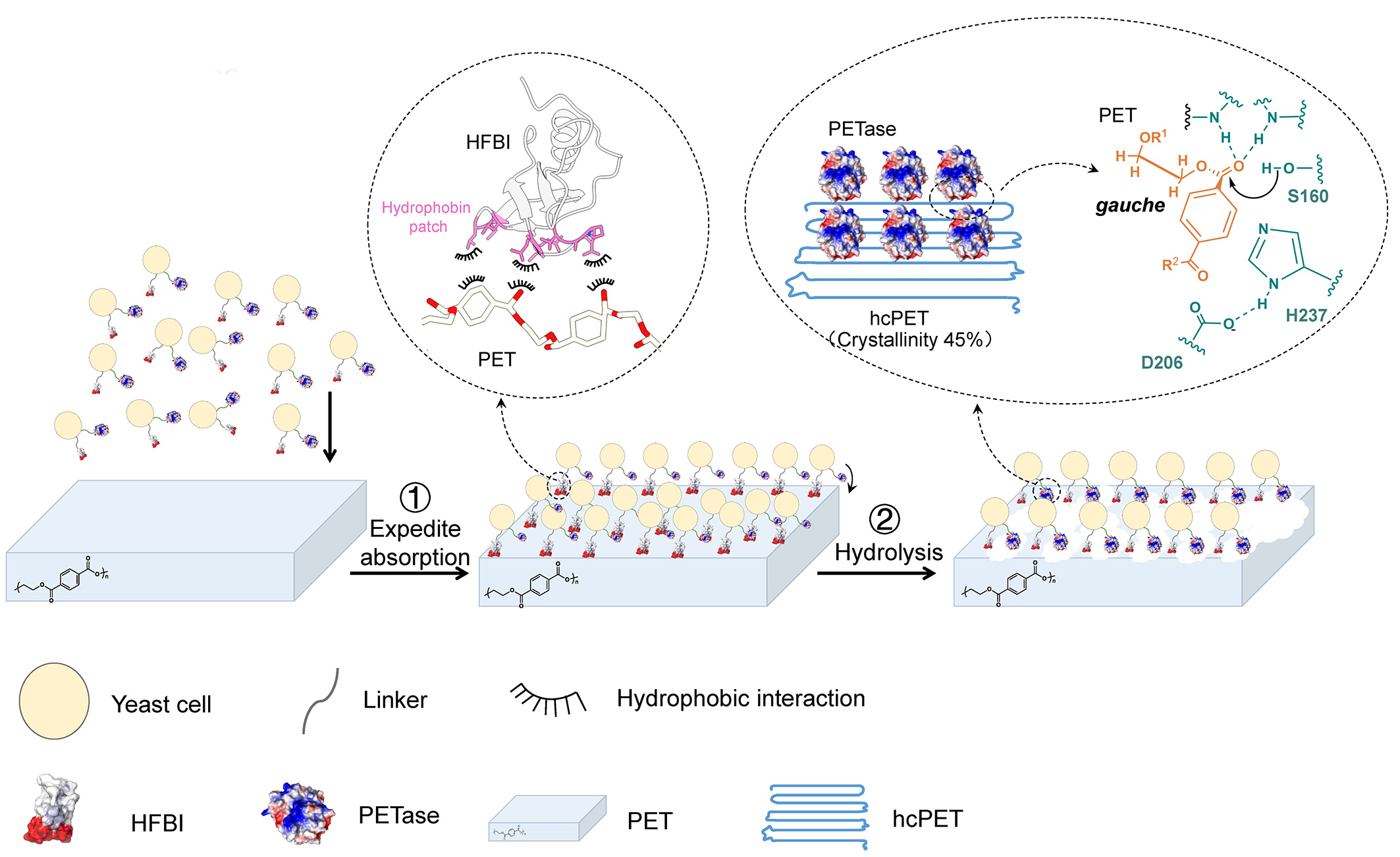
Figure. 2 Schematic diagram of hydrolysis of hcPET by codisplay systerm. Firstly, due to the presence of HFBI, codisplayed cells quickly adsorbed to the hcPET surface, and the adsorption rate on the hcPET surface was close to 100%. Secondly, PETase contacts the surface of high crystallinity PET, and then hydrolyzes the PET chains, thus achieving the effect of efficient hydrolysis of high crystallinity PET. Different figures are used in the illustration to represent different elements.
Our results not only provide an efficient strategy for efficiently biodegrading hcPET but also demonstrate the plasticity of the surface display system. Our study provides a rational organization of different functional units on the microbial surface for enhanced biocatalytic activity, which could find more applications in biocatalysis, biosensing, and bioenergy. You can read more about our approach and methods in the paper in Nature Communications, https://www.nature.com/articles/s41467-022-34908-z.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in