Accelerating Synthetic Biotic Development with Gut-on-a-chip Technology
Published in Bioengineering & Biotechnology
The tools of synthetic biology enable probiotic bacteria to be engineered to perform therapeutic functions within the human body (Charbonneau et al., 2020). These engineered live bacterial therapeutics are a new class of medicines that could address patients’ needs across a broad spectrum of indications and may have the potential to augment human physical and cognitive performance. The low cost of DNA synthesis, combined with high throughput screening of engineered bacterial clones, enables rapid construction of prototype strains. However, advancing candidates into the clinic is low throughput since investigational products for human studies must adhere to strict safety regulations and clinical studies are inherently costly and time-consuming. Therefore, there is strong incentive to develop translational strategies for evaluating strain candidates preclinically. In our article, “Characterization of an engineered live bacterial therapeutic for the treatment of phenylketonuria in a human gut-on-a-chip,” we describe the results of a collaboration between the U.S. Air Force Research Laboratory (AFRL) and Synlogic, a clinical stage biotechnology company, to apply human gut-on-a-chip technology and mathematical models to predict the function of an engineered bacterial therapeutic in vivo (Nelson et al., 2021: (https://www.nature.com/articles/s41467-021-23072-5)).
A cross-department effort by the U.S. Department of Defense, referred to as Synthetic Biology for Military Environments (SBME), was established in 2017 to advance synthetic biology technology to support and/or enhance performance in U.S. Warfighters. Engineered bacterial therapeutics could provide real-time surveillance and remediation of environmental stressors to enable maximal Warfighter performance and readiness. Organ-on-a-chip technology was identified early in the SBME program as a potential solution for screening synthetic live bacterial interventions (biotics)It in the context of human tissues. As a contributor to SBME, the AFRL 711th Human Performance Wing, located at Wright-Patterson Air Force Base, established a gut-on-a-chip microfluidic testing platform (gut-chip). These gut-chips are constructed using enterocyte-like Caco-2 and goblet cell-like HT-29 MTX cells interfaced with microvascular endothelial cells and undergo spontaneous morphogenesis, forming macrovillus structures and a functional gut-blood barrier (Figure 1).
Figure 1. Confocal Z-stack reconstruction of a human gut-chip.
Synlogic, based in Cambridge, MA, is pioneering the development of Synthetic Biotic medicines for the treatment of metabolic and inflammatory diseases. One such indication is Phenylketonuria (PKU), a rare inherited metabolic disorder that causes an inability to degrade dietary phenylalanine (Phe) and an accumulation of systemic Phe, leading to severe neurological complications. Synlogic’s clinical candidate, SYNB1618, is engineered to consume Phe within the gastrointestinal tract, with the objective of lowering Phe levels in the blood.
In this article, we describe our collaborative effort to combine the platform developed by AFRL with Synlogic’s engineered bacterial therapeutic development expertise to characterize the function of a bacterial strain related to SYNB1618, called SYN5183, using human gut-chip technology. SYN5183 is engineered to be auxotrophic for diamino-pimelic acid (DAP), and in the absence of exogenously applied DAP, SYN5183 does not replicate. We found that a single dose of SYN5183 cleared the gut-chips within several hours, allowing us to examine the kinetics of strain and metabolite transit within the gut-chips, as a function of dose level. This feature distinguishes gut-chips from static cell cultures and provides an opportunity to model clinical dosing regimens and tissue exposure to engineered bacterial therapeutics. In the gut-chip model, we observed dose-dependent production of the strain-specific biomarker, trans-cinnamic acid (TCA), as well as depletion of blood Phe levels after co-administration of SYN5183 and Phe in the gut compartment (Figure 2). These findings are consistent with our previous reports of SYNB1618 function in Non-Human Primates (NHP), healthy subjects, and PKU patients (Isabella et al., 2018; Puurunen et al., 2021).
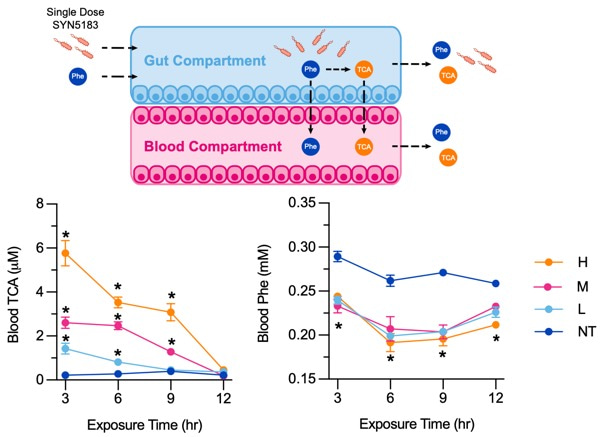
Figure 2. SYN5183 lowers dietary Phe uptake in a gut-chip model.
It has been reported that circulating Phe can enter the gastrointestinal lumen through a process termed enterorecirculation (Chang et al., 1995). Of particular interest to us was to use the gut-chip model to determine whether gut-resident SYN5183 could deplete high circulating Phe levels, as seen in PKU patients. By administering SYN5183 to the gut compartment and Phe to the blood compartment, we observed a dose-dependent lowering of plasma Phe in the gut-chip model (Figure 3). Lastly, we used the data from these models to calibrate a mathematical framework of SYN5183 Phe consumption in the gastrointestinal tract. Strikingly, we found that these mathematical models could accurately recapitulate the effects of SYNB1618 administration on plasma Phe in NHP.
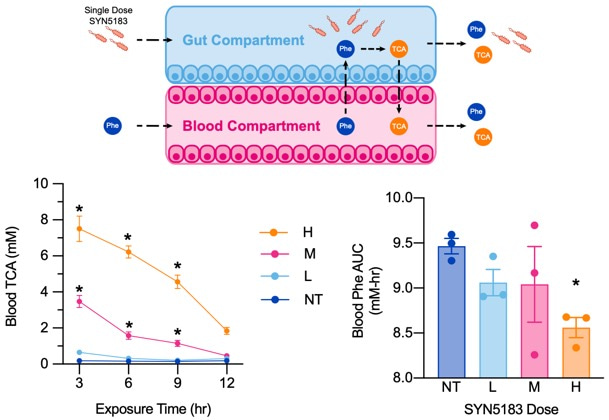
Figure 3. SYN5183 in the gut lowers blood Phe in a gut-chip model.
There are notable limitations to the gut-chip approach, including that implementation of anaerobic conditions and diverse microbial communities within the simulated gut lumen remains a significant technical challenge. In addition, the coarse-grained mathematical modeling approach applied here could be extended to consider bacterial gene expression, metabolism, and/or viability. Nevertheless, this relationship between a DoD laboratory and a public biotechnology company exemplifies the synergistic nature of scientific collaboration, and our findings suggest that translational gut-chip model systems, together with mathematical frameworks that describe strain function in the body, can serve as powerful tools for screening engineered bacterial strain prototypes and predicting their effects on human biology. Synlogic and the AFRL, along with the MIT Christopher Voigt Lab, will continue to work together through an ongoing effort to develop engineered bacterial therapeutics and other novel investigational medicines to address battle fatigue.
About Synlogic
Synthetic biology brings engineering principles to science, enabling the design of new biological systems, genetic circuits and molecular components that can be used to engineer medicines that can have a meaningful impact for patients. Synlogic, a clinical stage biotechnology company, is delivering the power of synthetic biology to medicine, combining the precision of engineering with rational drug development. With a premiere synthetic biology platform that leverages a reproducible, modular approach to microbial engineering, Synlogic designs Synthetic BioticTM medicines to treat disease in new ways.
Link to full manuscript: (https://www.nature.com/articles/s41467-021-23072-5)
References
Chang, T. M., Bourget, L. & Lister, C. A new theory of enterorecirculation of amino acids and its use for depleting unwanted amino acids using oral enzyme-artificial cells, as in removing phenylalanine in phenylketonuria. Artif Cells Blood Substit Immobil Biotechnol 23, 1–21 (1995).
Charbonneau, M. R., Isabella, V. M., Li, N. & Kurtz, C. B. Developing a new class of engineered live bacterial therapeutics to treat human diseases. Nature Communications 11, 1738 (2020).
Isabella, V. M. et al. Development of a synthetic live bacterial therapeutic for the human metabolic disease phenylketonuria. Nat. Biotechnol. 36, 857–864 (2018).
Nelson M.T., Charbonneau M.R., Coia H.G., Castillo M.J., Holt C., Greenwood E.S., Robinson P.J., Merrill E.A., Lubkowicz D., & Mauzy C.A. Characterization of an engineered live bacterial therapeutic for the treatment of phenylketonuria in a human gut-on-a-chip. Nat. Comms. 12, 2805 (2021).
Puurunen, M. K. et al. Safety and pharmacodynamics of an engineered E. coli Nissle for the treatment of phenylketonuria: a first-in-human Phase 1/2a study. Nature Metabolism. In press (2021).
CLEARED for PUBLIC release by the 711th Human Performance Wing, CASE Number: AFRL-2021-2310, CLEARED on 11 AUG 2021
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in