An enzymatic pathway in the human gut microbiome that converts A to universal O type blood
Published in Microbiology
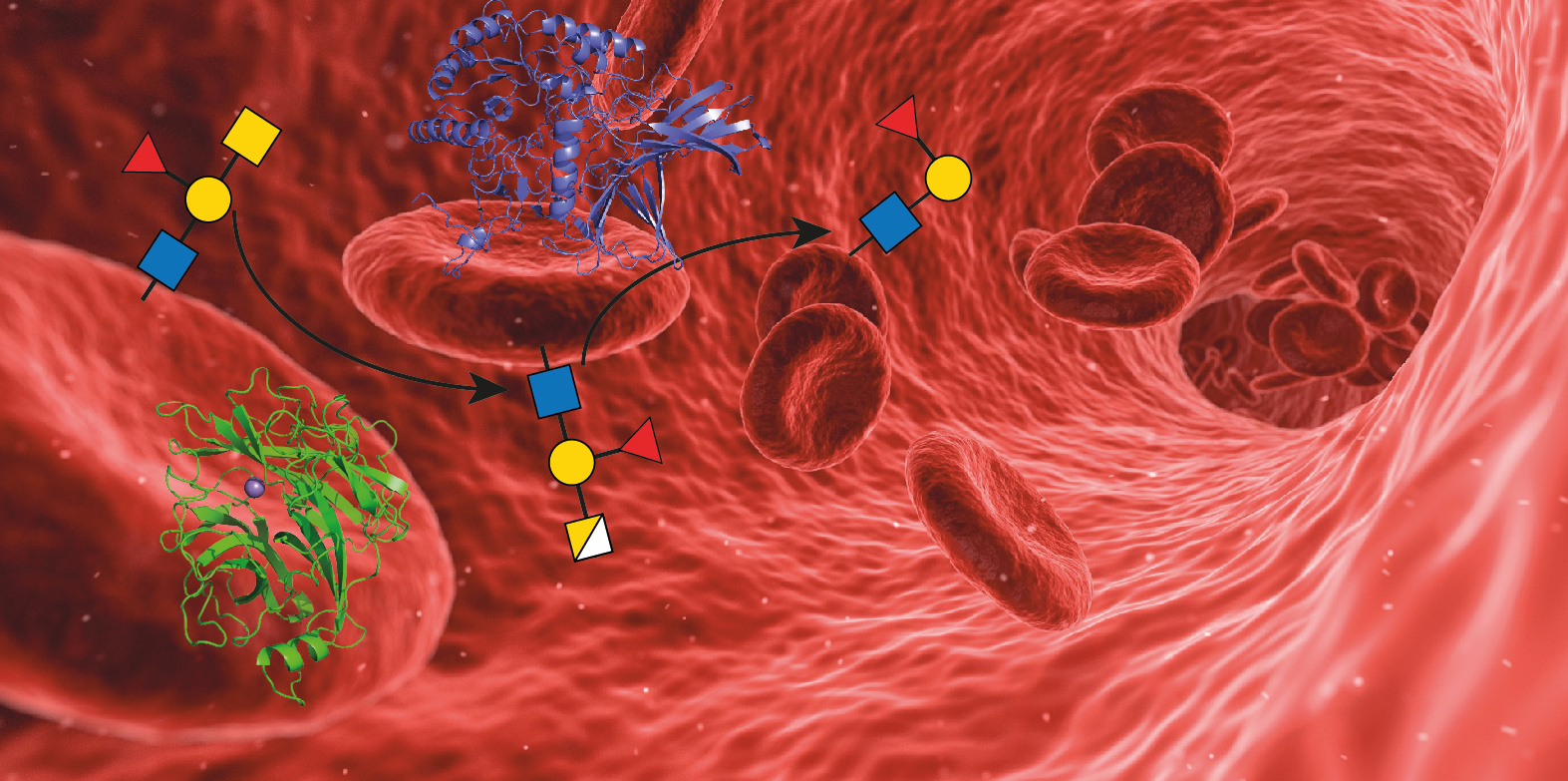
Enzymes can be used to convert A and B type red blood cells (RBC) into O type, broadening the supply. However the enzymes available to date have not worked well on whole blood, thus new enzymes are needed. We in the Withers Laboratories asked: Where do we find those enzymes?
Interestingly the answer to this question was, within us! In recent years the research community has started to realize the importance of the human microbiome in the context of human health. However, it may prove to be even more important since the microorganisms within us also harbor enzyme activities we do not even known about, yet. The human gut is covered with mucins, large glycoproteins presenting a variety of glycan structures on their surface, including those of the blood group antigens. Bacteria within the gut derive some of their energy by foraging these glycans, so must produce enzymes to cut them off. We investigated this community by use of functional metagenomics - a technique that allows the screening of enzyme activity without culture bias. By screening a library of gut microbiome enzymes using fluorogenic substrates that mimic the blood antigen carbohydrates, we identified a set of enzymes expressed by a particular bacterium, Flavonifractor plautii, that are able to cleave the A antigen very efficiently.
An important factor in the efficient cleavage of substrates bound to cell surfaces, such as the A antigens on the RBCs, is the ability of the enzyme to associate with the cell surface. Some enzymes do this by acquiring a net charge that is complementary to that of the cell surface. In our case the enzymes carry carbohydrate binding modules (CBMs) that interact with other cell surface glycans allowing them to rapidly cleave the nearby A antigens. The combination of high substrate activity and affinity for the RBC surface makes our enzymes very efficient in the removal of A antigens from RBCs in whole blood samples.
In the future we hope these enzymes will be widely deployed for the production of enzymatically converted universal donor blood (ECO O Type RBCs) directly after blood donation. In addition to their use in RBC conversion we plan to test the use of these and related enzymes in the removal of antigens from other important cell surfaces and tissues. Such approaches could widen the availability of good “matches” in organ and stem cell transplantation's.
For me it was very interesting to see what our gut microbiome had to offer and I am keen to see what kind of other activities will be discovered within the human gut microbiome in the future.
For more information check out the full manuscript: https://doi.org/10.1038/s41564-019-0469-7
Illustration was designed by Andreas Geissner.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in