Assessment of three antibiotic combination regimens against Gram-negative bacteria causing neonatal sepsis in low- and middle-income countries
Published in Microbiology

The work presented in this article is part of a large global, prospective, hospital-based observational study (NeoOBS) across four continents, including LMIC settings. The clinical findings revealed:
- a high mortality among infants with culture positive sepsis (almost 1 in 5);
- wide variations in standard of care for sepsis in neonates and young infants, with more than 200 different antibiotic combinations, significant divergence from WHO-recommended regimens and
- frequent switching of antibiotics.
Along with these findings, the analyses performed on the invasive bacterial isolates from these patients showed
- high genomic diversity, not only between different countries but even within the same hospital, and
- high levels of resistance to the first and second-line antibiotic treatments recommended by WHO for treatment of this life-threatening condition.
We focused on the Gram-negative bacteria as they are often the culprits of neonatal sepsis and are known to be resistant to many antibiotics. Our objective was, by using drug-susceptibility testing, genetic-diversity analyzes and hollow fibre infection model (HFIM), to assess the potential activity of the novel combination antibiotic regimens that can help to treat and/or prevent the spread of neonatal sepsis.
Identifying potential novel antibiotic regimens is a complex process with multiple steps, out of which drug-susceptibility testing is the first one, followed by whole genome sequencing (WGS). As not all bacterial strains are identical and some may have unique genetic mutations that can affect the effectiveness of antibiotics, with WGS you can identify these mutations and even more important, to determine if the identified antibiotic regimens can still fight the different strains.
Results
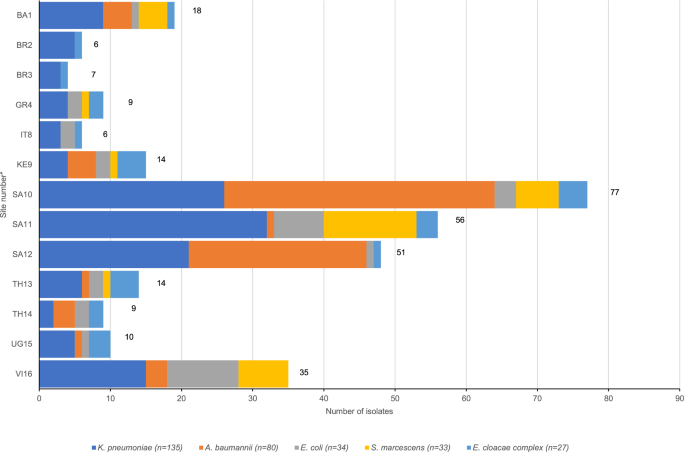
In our study of 420 Gram-negative bacterial isolates, Klebsiella pneumoniae, Acinetobacter baumannii, Escherichia coli, Serratia marcescens, and Enterobacter cloacae complex (ECC) were the most common species identified. These isolates displayed significant resistance to traditional antibiotics such as ampicillin/gentamicin and cefotaxime. K. pneumoniae exhibited high resistance rates due to ESBLs and AMEs, with 30% showing carbapenem resistance primarily through NDM-like and OXA-48-like genes.
Antibiotic susceptibility testing revealed that new combinations like flomoxef-amikacin and fosfomycin-amikacin showed superior efficacy against ESBL-producing isolates compared to traditional regimens. Resistance mechanisms were closely linked to genetic profiles, with a strong correlation between phenotypic and genotypic resistance patterns. The presence of specific resistance genes, such as those encoding ESBLs, carbapenemases, and aminoglycoside-modifying enzymes, was a key determinant of antibiotic resistance.
K. pneumoniae isolates were genetically diverse, with high prevalence of ESBL genes like blaCTX-M-15 and significant geographic variation in MDR strains. A. baumannii isolates, primarily from South Africa, displayed extensive drug resistance, including prevalent blaOXA-23 and blaNDM-1 genes. ECC isolates showed resistance largely due to intrinsic AmpC genes, while S. marcescens and E. coli also harbored multiple resistance genes, including ESBLs and carbapenemases.
Conclusions
These findings highlight the urgent need for novel treatment strategies and continuous surveillance of antibiotic resistance in Gram-negative bacteria. It is therefore important to continue conducting clinical trials such as the NeoSep1 antibiotic trial (ISRCTN48721236) which aims to identify novel first and second-line empiric antibiotic regimens for neonatal sepsis and to validate these regimens ensuring their safety and effectiveness for newborns.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in