Bacterial Metacommunities: playing the game of selection
Published in Microbiology

The metacommunity paradigm provides an adequate framework to answer this question1,2. Under this approach, communities are spatially connected in a network by dispersal, and the processes that determine their structure are the result of the interplay between local factors and regional dynamics. Recently, Mark Vellend3 proposed a conceptual framework in which the structure of metacommunities can be explained by four "high-level processes": selection, dispersal, ecological drift, and diversification. These processes are universally present across ecological communities, but the strength in which they rule the assembly may vary across different communities and the environmental heterogeneity.
In bacteria from freshwater ecosystems, several studies have found evidence of the pivotal role of selection in determining the community structure, and a decrease in its relative importance as dispersal rates increase and the systems become more homogeneous 4-12 . These attempts have been spotted considering only the spatial scale, but not considering the temporal contrasting scenarios of environmental heterogeneity.
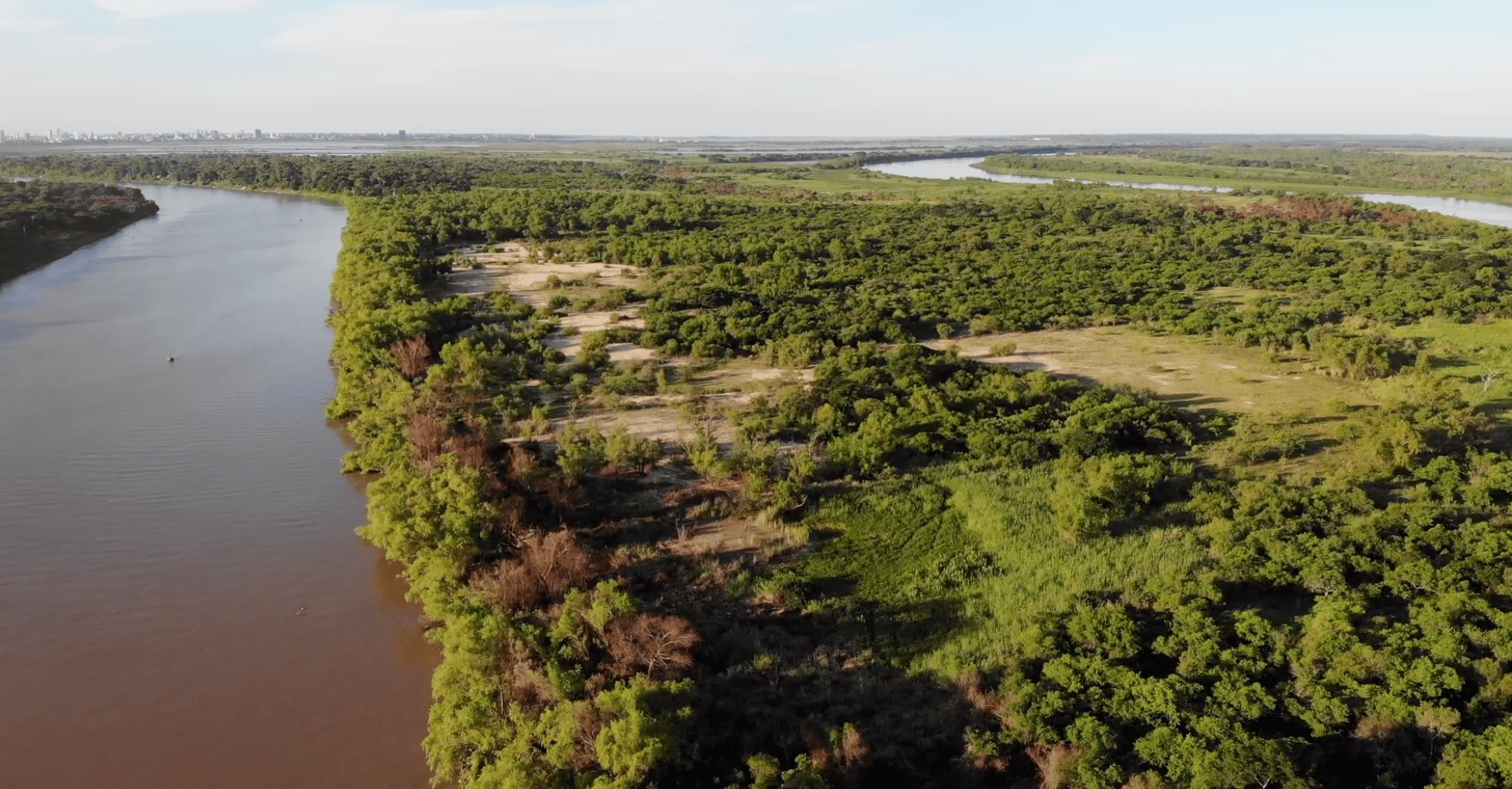
To address the full scope of this issue, we carried out a spatiotemporal survey in the complex floodplain system of the Paraná River (the second largest in South America), which is characterized by a wide range of temporal and spatial heterogeneity mediated by irregular hydrological fluctuations13.
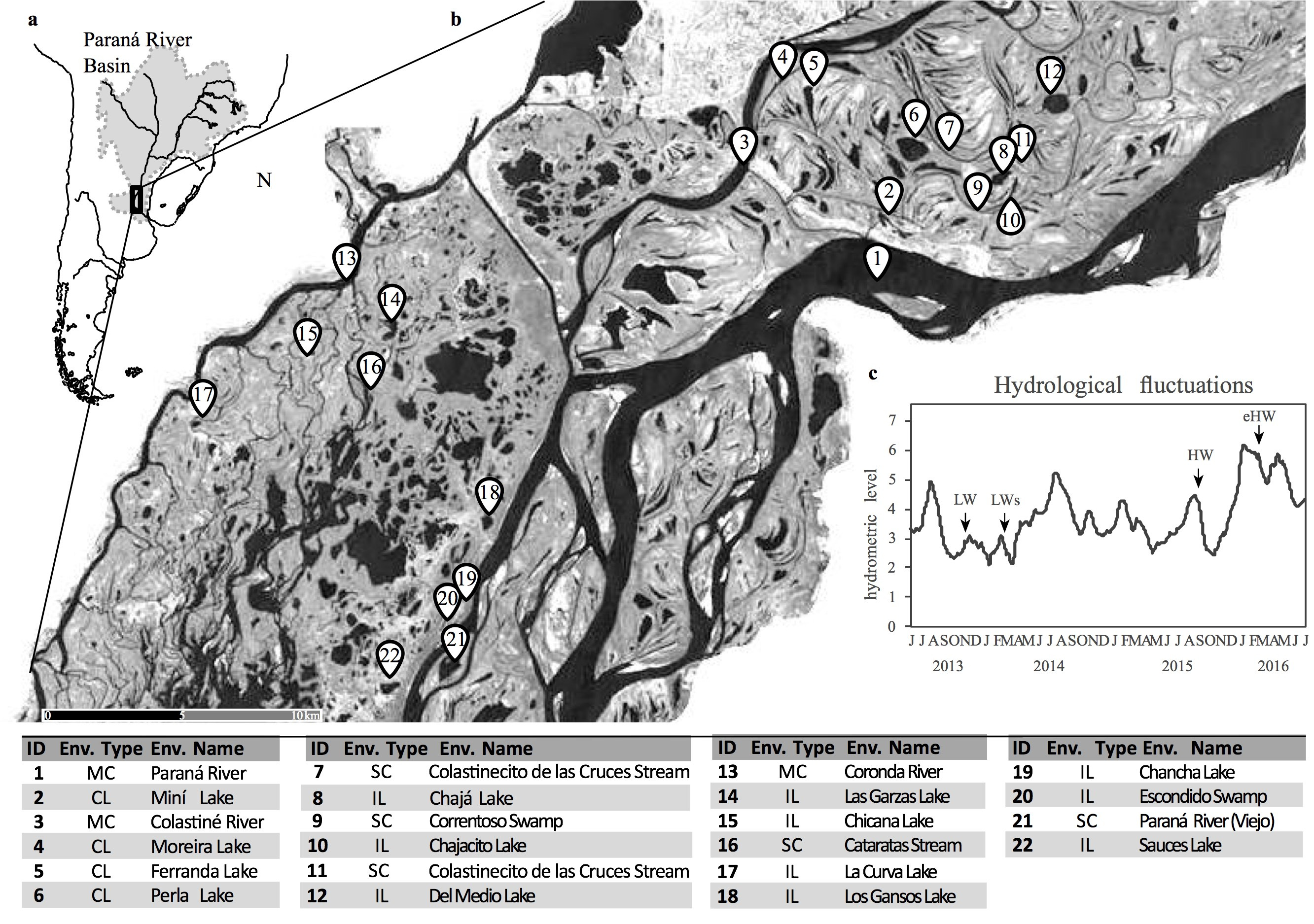
Figure: (a) The Paraná fluvial system location in South America and (b) the environments sampled (IL isolated lake, CL connected lake, SC secondary channel, MC main channel). (c) Daily water level in the Paraná River showing the sampled periods: LW low water, LWs low water with sedimentological pulse, HW high water, eHW extraordinary high water.
We observed for the first time that selection is the most important process structuring the bacterial metacommunity at both extremes of environmental heterogeneity and homogeneity, changing the general view that selection is weakened by dispersal homogenization. At high environmental heterogeneity, community structure is mainly determined by heterogeneous selection increasing the divergence in local communities, whereas, in extremely homogeneous environmental conditions, the low diversity of niche habitats leads to an increase of homogeneous selection, and local communities are strongly filtered by common environmental factors. Besides, we advance in showing how the ecological processes also determined the bacterial network associations, and that spatiotemporal heterogeneity is an important factor determining the most highly connected taxa in the system.
Integrating all the empirical evidence, we devised a conceptual model that synthesizes how environmental heterogeneity determines the action of the ecological processes assembling the bacterial metacommunity.
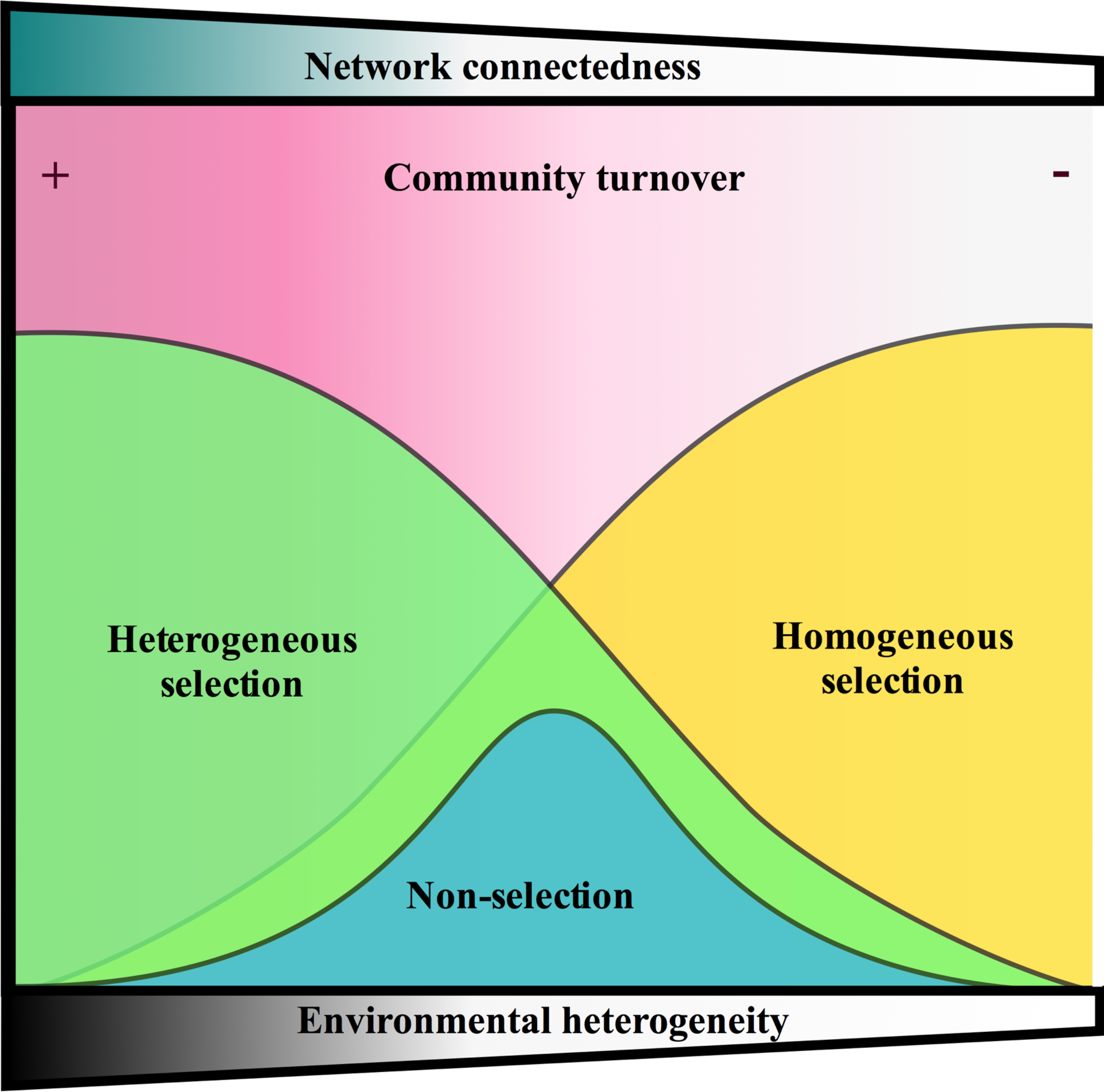
Figure: Conceptual model that synthetizes how environmental heterogeneity determines the action of the ecological processes on bacterial metacommunity assemblage and influences in the network associations.
A particular strength of this model is that it was conceived relying on different metacommunity features (e.g., taxonomic and phylogenetic turnover 14, network associations15) but taking into account the spatial and temporal variability scales. We challenged future studies to test it in other communities.
References:
1. Leibold MA, Holyoak M, Mouquet N, Amarasekare P, Chase JM, Hoopes MF, et al. The metacommunity concept: a framework for multi-scale community ecology. Ecol Lett. 2004; 7:601–13.
2. Wilson DS. Complex interactions in metacommunities, with implications for biodiversity and higher levels of selection. Ecology. 1992;73:1984–2000.
3. Vellend M. The theory of ecological communities. 1st ed. Woodstock, UK: Princeton University Press; 2016.
4. Lindström ES, Langenheder S. Local and regional factors influencing bacterial community assembly. Environ Microbiol Rep. 2012;4:1–9.
5. Nemergut DR, Schmidt SK, Fukami T, O’Neill SP, Bilinski TM, Stanish LF, et al. Patterns and processes of microbial community assembly. Microbiol Mol Biol Rev. 2013;77:342–56
6. Comte J, Lovejoy C, Crevecoeur S, Vincent WF. Co-occurrence patterns in aquatic bacterial communities across changing permafrost landscapes. Biogeosciences. 2016;13:175–90.
7. Heino J, Melo AS, Siqueira T, Soininen J, Valanko S, Bini LM. Metacommunity organisation, spatial extent and dispersal in aquatic systems: Patterns, processes and prospects. Freshw Biol 2015;60:845–69.
8. Logares R, Deutschmann IM, Giner CR, Krabberød AK. Different processes shape prokaryotic and picoeukaryotic assemblages in the sunlit ocean microbiome. Environ Microbiol. 2018;20:37–49.
9. Wang J, Shen J, Wu Y, Tu C, Soininen J, Stegen JC, et al. Phylogenetic beta diversity in bacterial assemblages across ecosystems: deterministic versus stochastic processes. ISME J. 2013;7:1310–21
10. Dini-Andreote F, Stegen JC, Van Elsas JD, Salles JF. Disentangling mechanisms that mediate the balance between stochastic and deterministic processes in microbial succession. Proc Natl. Acad Sci USA. 2015;112:E1326–E1332.
11. Declerck SAJ, Winter C, Shurin JB, Suttle CA, Matthews B. Effects of patch connectivity and heterogeneity on metacommunity structure of planktonic bacteria and viruses. ISME J. 2013;7:533–42.
12. Fodelianakis S, Lorz A, Valenzuela-Cuevas A, Barozzi A, Booth JM, Daffonchio D. Dispersal homogenizes communities via immigration even at low rates in a simplified synthetic bacterial metacommunity. Nat Commun. 2019;10:1. https://doi.org/10. 1038/s41467-019-09306-7.
13. Mayora G, Scarabotti P, Schneider B, Alvarenga P, Marchese M. Multiscale environmental heterogeneity in a large riverfloodplain system. J South Am Earth Sci. 2020;100:102546.
14. Stegen JC, Lin X, Konopka AE, Fredrickson JK. Stochastic and deterministic assembly processes in subsurface microbial communities. ISME J. 2012;6:1653–64.
15. Faust K, Raes J. Microbial interactions: from networks to models. Nat Rev Microbiol. 2012;10:538–50.
This study was supported by Agencia Nacional de Promoción Científica y Tecnológica (PICT 2016-0465 PI, MD), Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET).
Photograph by Sofía Jiménez.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in