Canopy 4 inhibits the Hedgehog pathway by modulating membrane sterol lipids
Published in Cancer
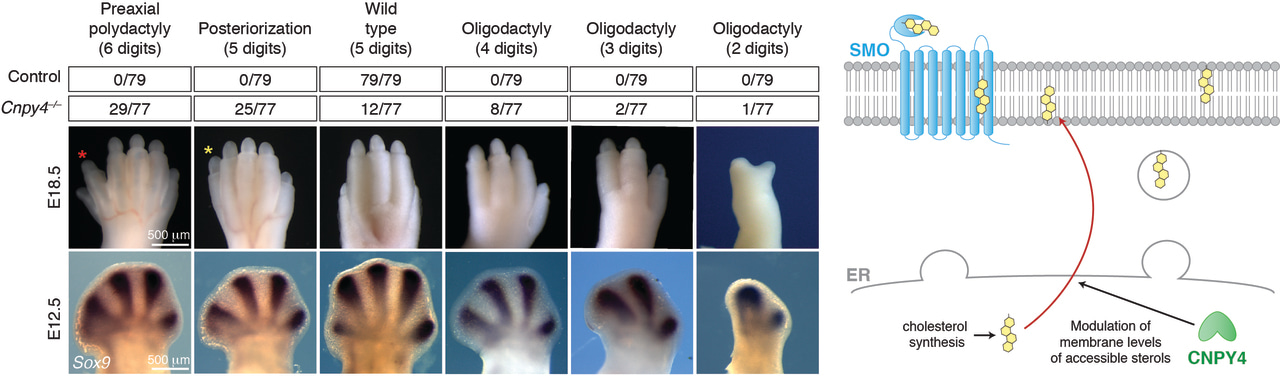
The Hedgehog pathway and cancer
Though perhaps best known for its critical role in development (1), the Hedgehog pathway (HH) is a major contributor to cancers, including medulloblastoma (2) and basal cell carcinoma (3). However, current therapeutic options against such HH-driven cancers, including the inhibitors vismodegib and sonidegib, have proven to be suboptimal with concerns over off-target effects, toxicity, low efficacy, and relapse (4, 5). As these drugs typically target the main activating receptor of the HH pathway, Smoothened (SMO), it has become increasingly clear that characterization of other nodes of the pathway, especially regulatory elements, is necessary for development of therapeutics with enhanced properties.
Sterol regulation of the HH signaling
One exciting new avenue of HH regulation that has only recently begun to be appreciated is the role of sterol lipids (6). While it has been known for some time that the HH pathway could be activated by treatment with a number of lipid molecules, including cholesterol and several of its precursors and metabolites (7), it was unclear how much of this effect was relevant in vivo. A recent study on the effect of a combination treatment of simvastatin, which broadly lowers cholesterol levels, and vismodegib on medulloblastoma in mice showed promising results in comparison to the effects seen in mice treated with vismodegib alone (8). Not only did the combination treatment inhibit the growth of tumors in mice more effectively than vismodegib alone, but it also did so without any apparent bone toxicity, a common side effect from vismodegib treatment. Thus, it is likely that cholesterol directly impacts HH signaling. However, how this mechanistically could be possible when cholesterol is ubiquitously expressed in cell membranes was only recently revealed in the publication by Kinnebrew, M. et al. The authors suggested that a particular subset of membrane cholesterol that is in a chemically accessible form is the endogenous ligand for SMO (9), a molecule whose identity has remained a mystery for decades despite continual effort on its elucidation.
Identification of Canopy4 as a regulator of the HH pathway
In our present study, we demonstrate that changes to such accessible cholesterol levels in the cell membrane directly impacts HH-mediated development in mice lacking the Canopy 4 (CNPY4) protein. Though these effects were most prominent in the limbs, presenting as abnormalities in digit number and development of the long bones, Cnpy4 null mice additionally had perturbations to their overall skeletal morphology and brain development. While the CNPY family of proteins has been predominantly studied in the context of fibroblast growth factor signaling (10), our examinations of CNPY4’s effect on the major developmental signaling pathways suggested that CNPY4 operated primarily on the HH pathway instead. Cells lacking CNPY4 had significantly elevated basal HH signaling, and this effect was dramatically increased in the presence of any one of the varied SMO agonists, including recombinant HH ligand, a chemical SMO agonist SAG, and multiple cholesterol metabolites.
As the CNPY family of proteins belongs to a larger superfamily of proteins collectively termed Saposin-like proteins, which have been largely linked to lipid biology (11), we examined the effect CNPY4 may have on the newly postulated endogenous ligand of the HH pathway, i.e., membrane levels of accessible cholesterol. We found that in both cells directly derived from Cnpy4 null limbs and immortalized cells that had Cnpy4 transiently knocked down, the membrane levels of accessible cholesterol were significantly increased. Furthermore, the stimulatory effect CNPY4 had on the HH pathway was completely eliminated in the absence of cholesterol. Taken together, we concluded that the severe developmental phenotypes we observed in Cnpy4 null mice were likely a direct result of alterations to the levels of accessible cholesterol and thus hyperactive HH signaling.
In summary, our study demonstrates that CNPY4 is a newly reported negative regulator of the HH pathway. Deletion of Cnpy4 in mice manifests in severe developmental abnormalities, and knockdown of Cnpy4 in cells results in significant hyperactivation of the HH transcriptional pathway. We traced the possible source of this Cnpy4 null-dependent alteration to changes in the membrane levels of accessible cholesterol, which is believed to be the endogenous ligand for SMO. Thus, our study has uncovered a new node by which the HH pathway can be indirectly modulated and further corroborated the idea that drugs which alter the levels of cholesterol may have a pharmacologically significant impact on the treatment of HH-derived cancers.
The entirety of our findings have been published here: https:/rdcu.be/cMArf
References:
- Nieuwenhuis, E. & Hui, C. c. Hedgehog signaling and congenital malformations. Clin. Genet. 67, 193–208 (2004).
- Raleigh, D. R. & Reiter, J. F. Misactivation of Hedgehog signaling causes inherited and sporadic cancers. J. Clin. Investig. 129, 465–475 (2019).
- Bakshi, A. et al. Basal cell carcinoma pathogenesis and therapy involving hedgehog signaling and beyond. Mol Carcinog. 56, 2543–2557 (2017).
- Yauch, R.L. et al. Smoothened mutation confers resistance to a Hedgehog pathway inhibitor in medulloblastoma. Science 326, 572–574 (2009).
- Robinson, G.W., et al. Irreversible growth plate fusions in children with medulloblastoma treated with a targeted hedgehog pathway inhibitor. Oncotarget. 8, 69295–69302 (2017).
- Cooper, M. K. et al. A defective response to Hedgehog signaling in disorders of cholesterol biosynthesis. Nat. Genet. 33, 508–513 (2003).
- Gofflot, F. Molecular mechanisms underlying limb anomalies associated with cholesterol deficiency during gestation: implications of Hedgehog signaling. Hum. Mol. Genet. 12, 1187–1198 (2003).
- Fan, Q. et al. Statins repress hedgehog signaling in medulloblastoma with no bone toxicities. Oncogene 40, 2258–2272 (2021).
- Kinnebrew, M. et al. Cholesterol accessibility at the ciliary membrane controls hedgehog signaling. eLIFE 8, e50051 (2019).
- Hirate, Y. & Okamoto, H. Canopy1, a Novel Regulator of FGF Signaling around the Midbrain-Hindbrain Boundary in Zebrafish. Curr. Biol. 16, 421–427 (2006).
- Bruhn, H. A short guided tour through functional and structural features of saposin-like proteins. Biochem. J. 389, 249–257 (2005).
Follow the Topic
-
Nature Communications

An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Mar 24, 2026
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in