Chemical Communication Drive Microbial Community Structure
Published in Microbiology
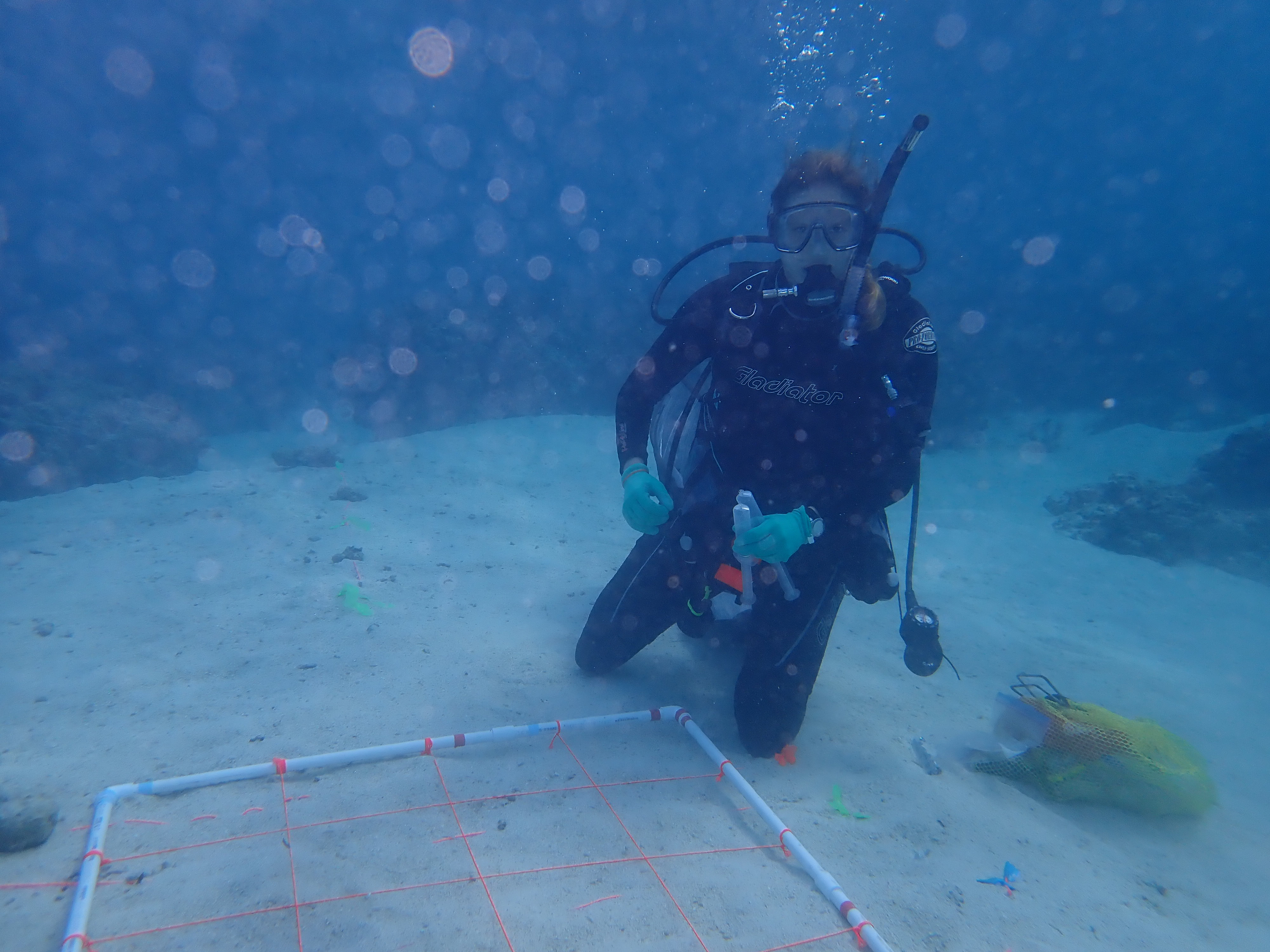

Microbial communities are assemblages organized by various ecological and evolutionary processes that contribute to the different types and number of species across environments. At broad biogeographic patterns, certain microbial taxa are largely associated with particular environments (e.g., Cyanobacteria in marine systems) due to conserved phylogenetic traits. Just as certain plant species live in tropical forests vs. temperate forests, microbial taxa disperse and differentially establish in environments afforded by their physiological traits, which are evolutionary constrained. While neutral processes like dispersal may contribute at these large spatial scales, microbial communities are largely composed of a subset of the regional species pool that have passed through environmental filtering due to adaptive differentiation of various abiotic factors. At smaller spatial scales at the local community, ecological theory predicts that biotic filters, such as predation, competition, and species interactions, further reduce these species pools resulting in the observed community. In our paper published in The ISME Journal (link), we examine the effects of these biotic interactions, as measured through the production of specialized metabolites, and their influence on microbial community structure in marine sediments (pictured above is co-author Dr. Alyssa Demko, currently a postdoc at the Smithsonian Institute, deploying our gridded plots for sample collection in Mo'orea, French Polynesia).
In these marine environments, chemistry is the language of choice for microbes. All organisms, including microbes, produce chemical compounds to interact with the environment. These may be for internal signaling, environmental responses, resource acquisition, or for antagonistic interactions like antibiotic production. In microbiomes, these metabolites perform a variety of ecological roles, with many mediating species interactions. Thus, they are classified as biotic filtering in classical ecological theory. However, we know little about the production of these metabolites in natural communities, their concentrations in natural systems, or even the roles most of these molecules perform in mediating interspecies relationships within these complex communities. This is due in part to the focus on these specialized metabolites for pharmaceutical exploitation, with most of the current drugs derived directly from these natural products.
Most of our knowledge of microbial natural products comes from culture-based studies under artificial laboratory conditions that provide limited inferences for the ecological roles of these chemical compounds under natural conditions. Even in current assessments of environmental metabolomes, we are further limited in reference databases for chemical characterization, inasmuch that our results highlight the deficits in relying on poorly annotated reference spectra and other unsupervised computational and chemoinformatic approaches. While most metabolites from these communities cannot be characterized, we compared the composition of various molecular features to correlate directly to community and functional composition, including the biosynthetic enzymes that produce these molecules. Our paper provides a multi-omic approach to connect taxonomic composition and biosynthetic potential to community metabolomes.
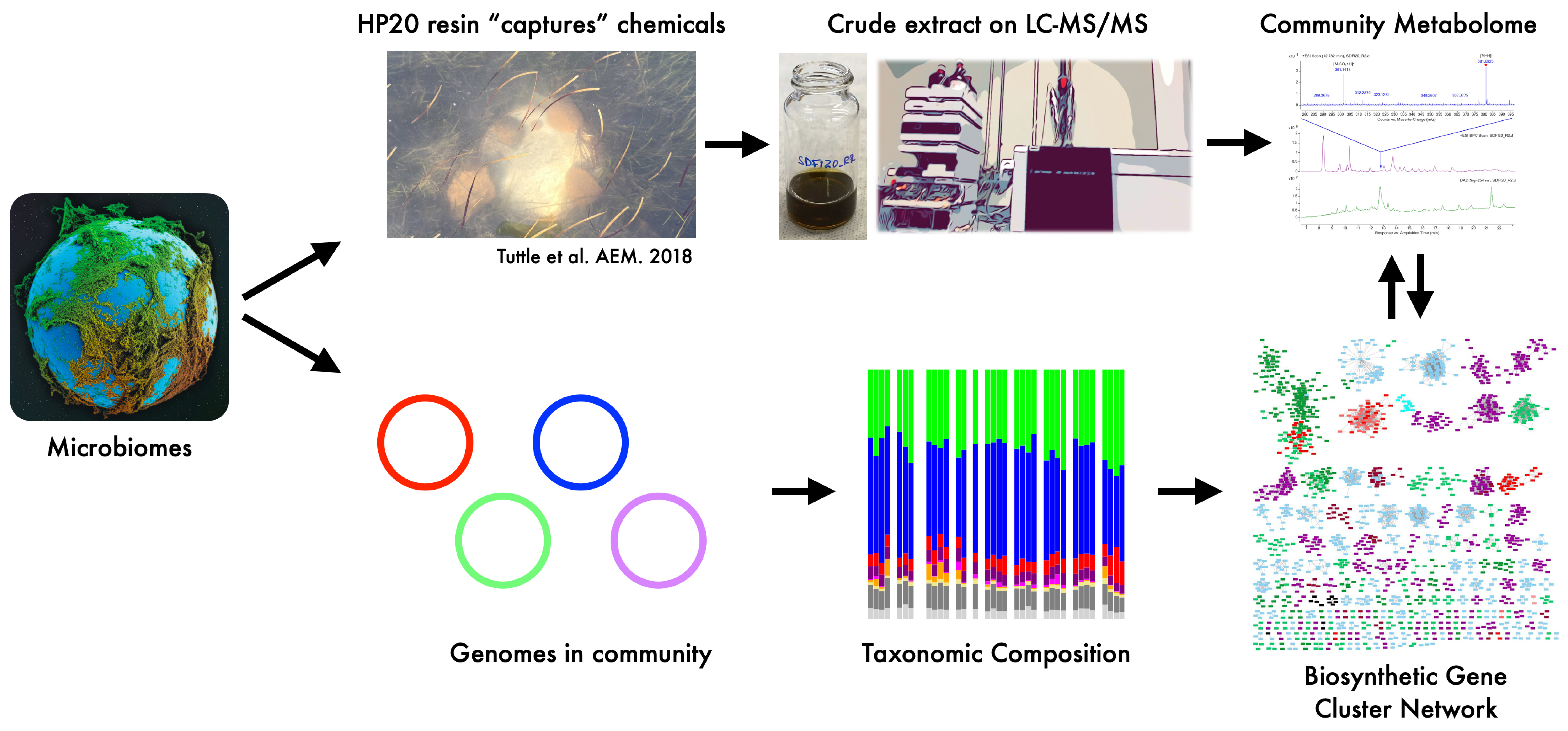
By reducing environmental variation among sites, we find that environmental selection was minimized across reef-associated sediment communities at varying spatial scales. Rather, at fine spatial scales, communities were differentiated by differences among closely-related taxa that varied extensively in their biosynthetic enzymes and composition of biosynthetic gene clusters (BGCs), suggesting that 16S rRNA resolved taxa are insufficient to link to metabolite production. These differences were further accentuated at the metabolome level, where each community harbored its unique assemblages of the metabolites mediating ecological interactions.
Based on our findings, we propose that future studies on natural products should prioritize ecological relevance and the characterization of their roles in mediating biotic interactions in natural communities. Here, we identified novel natural products (although more rigorous NMR techniques are needed to resolve absolute structures - something we are optimizing with "resin bags" in future work) underlying a vast array of uncharacterized chemical space. In particular, we found that understudied taxa, such as Myxococcus and Acidobacteria, harbor extensive, yet unexplored, biosynthetic potential that dwarfs our most known and prolific natural product producing bacteria, such as Streptomyces. Given the vast and largely unexplored diversity of natural products, there is tremendous potential for incorporating ecological and evolutionary perspectives for discovering novel compounds that may have biotechnological and biomedical applications.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in