Dancers in the Mud: Cable Break Spoils Microbial Party
Published in Microbiology
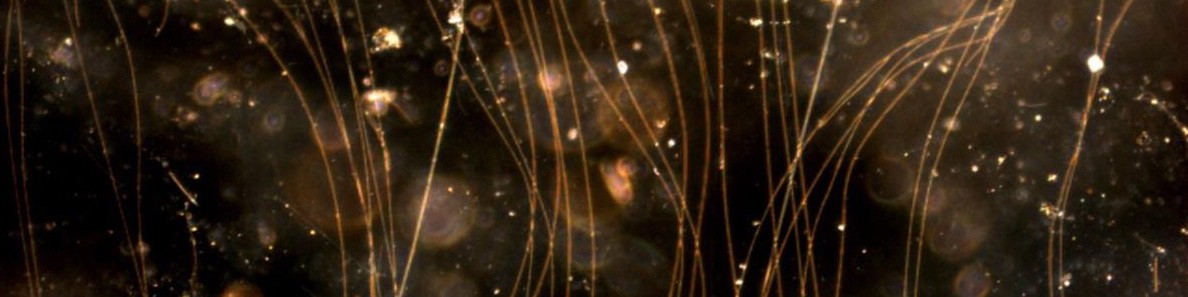
Several years ago, Jesper J. Bjerg, then a PhD student in the Microbiology group at Aarhus University, sat in the dark Raman microscope room of DOME, University of Vienna, measuring the cytochrome redox state of individual cable bacteria to follow their electron transport (see https://doi.org/10.1073/pnas.1800367115) – and nothing worked. While becoming increasingly impatient, suddenly something in the microscope caught his eyes: hundreds of tiny bacteria were swimming in a tight flock around the oxygen-free part of a cable bacterium; it looked as if they were dancing and having a party!
We had been suspecting earlier that other bacteria could “tap into the wire” by transferring electrons to cable bacteria and thus to indirectly “breathe” with oxygen in anoxic sediment. This idea was based on the 16S rRNA gene sequencing and FISH results of MSc student Signe Brokjær, who had seen certain bacteria associated with cable bacteria, soon to be substantiated by the great work of Diana Vasquez-Cardenas and colleagues (then at the Royal Netherlands Institute for Sea Research (NIOZ) in Yerseke.

But Jesper’s new observation was different, dynamic, and striking: it provided clear evidence that some motile bacteria (and there were many different kinds of them) were attracted by the cable bacterium. Was extracellular electron transfer (EET) the reason for this behavior? What followed was a year-long effort at the Center for Electromicrobiology (CEM) to unravel the nature of this phenomenon: Jesper (with scripting help by Paula Tataru) and Signe documented that the bacteria in the flock where metabolically more active when close to the cable bacterium; together with Anna Mueller, Markus Schmid, and Michi Wagner (the Raman microscopy experts at DOME Vienna), Jesper and Jamie Lustermans showed that c-type cytochromes of the flocking bacteria were more reduced when further away, and more oxidized when closer to the cable bacterium, implying EET from these bacteria to the cable bacterium. 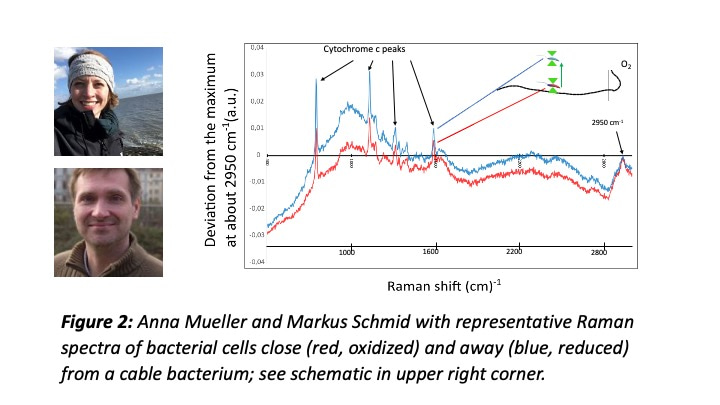 Metagenomic analysis by our bioinformatics wizards Casper Thorup and Ian Marshall (and genome analysis by Jamie) indicated a broad phylogenetic and metabolic diversity within the bacterial flocks, including sulfide oxidizers and chemoorganotrophs.
Metagenomic analysis by our bioinformatics wizards Casper Thorup and Ian Marshall (and genome analysis by Jamie) indicated a broad phylogenetic and metabolic diversity within the bacterial flocks, including sulfide oxidizers and chemoorganotrophs. 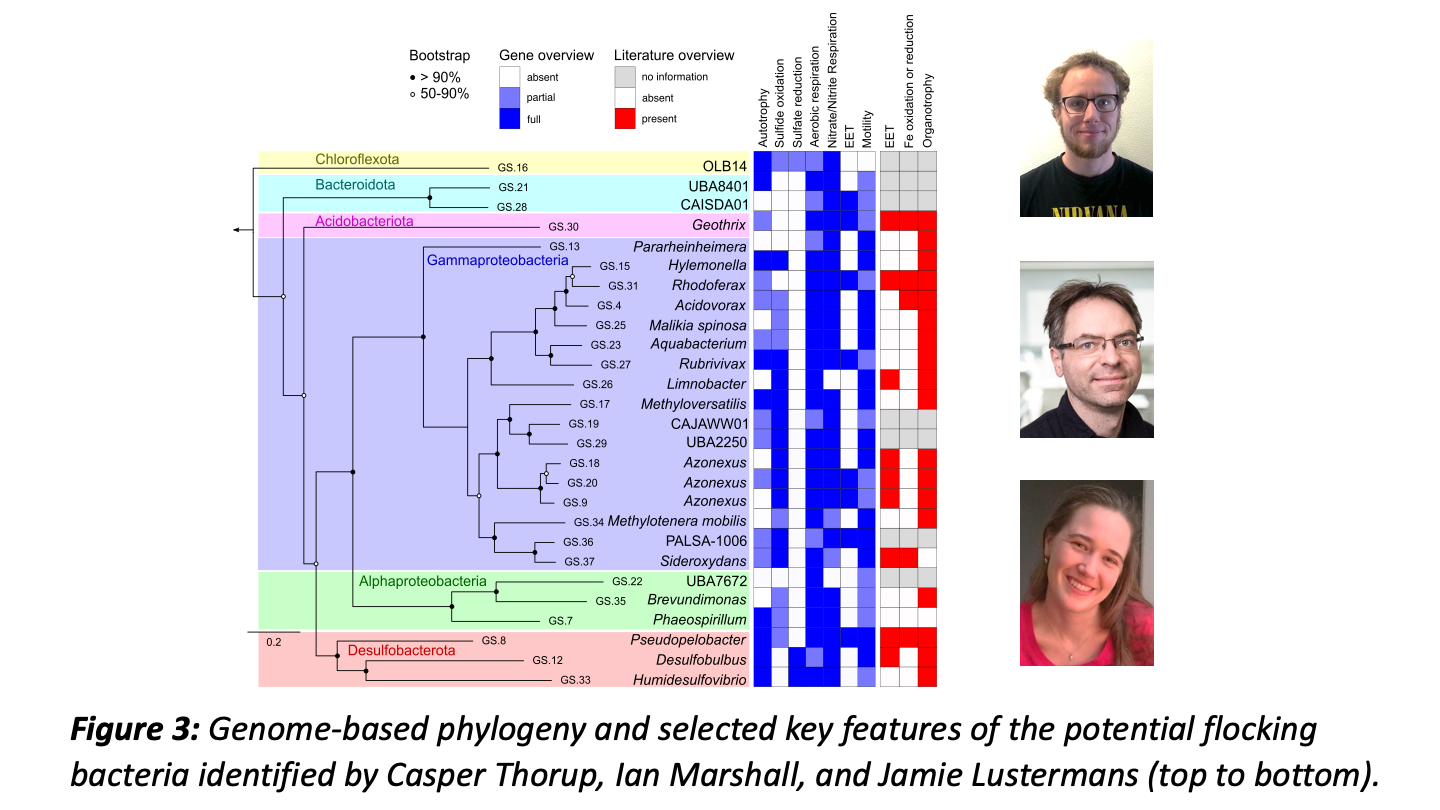
The most intriguing visualization of this close interaction, however, was a cut-experiment by Jesper, using a Laser Microdissection Microscope (LMD): when he cut the cable bacterium with the laser at a position with “dancing microbes”, the dancing stopped abruptly around those cable bacterium cells that had missed the electrical connection to oxygen.

Movie 2: Dispersing flock of bacteria after a cable bacterium is laser-cut (at 0.04 s) at the vertical line. The left side of the filament is disconnected from oxygen, so the flock disperses; the right side of the filament stays connected to oxygen, and consequently retains its flocking bacteria.
This cut experiment corroborated that the flocking bacteria use the cable bacterium as an electrical lifeline to oxygen, facilitating a quasi-aerobic metabolism in anoxic sediment. The most likely mechanisms for EET are (yet unidentified) electron shuttles, of which (according to calculations by Lars Peter Nielsen, the Head of CEM) nanomolar concentrations are sufficient to explain the flocking phenomenon and the rapid dispersal of the flocks after cutting the wire. Whether the cable bacterium benefits or suffers from the interaction, or can control it somehow, is yet to be determined. Read the complete story at https://www.nature.com/articles/s41467-023-37272-8.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in