Dense anatomical annotation of slit-lamp images improves the performance of deep learning for the diagnosis of ophthalmic disorders
Published in Bioengineering & Biotechnology
Previous medical datasets for machine learning were often collected for a single purpose, such as image-level classification of a specific disease, and therefore led to inadequate data for data mining and meaningful feature extractions, reflecting a major bottleneck of medical annotation for AI training. Moreover, data from most rare diseases is less readily available, undermining the representativeness of medical data, and hindering the development of algorithms. Therefore, we launched a Medical Artificial Intelligence ‘Lego’ Project, hoping to break through the data heterogeneity barriers of different disease disciplines by converting multidisciplinary medical data into "Lego" modules that can be combined together.
To address the challenges, the primary task was to establish a novel annotation approach for medical algorithm research. Moreover, the annotated dataset should be collected to be a general-purpose representation of clinical practice, without bias towards any particular task. Therefore, we took five years working on developing this annotation technique, which accurately extracts information from every part of a medical picture to mine the richness and potential of available data, hoping to improve the utilization of medical images and improve the performance of deep-learning algorithms.
The workflow of Visionome
Slit-lamp examination and photography is a common examination in our routine eye clinics. It helps to check and document multiple anterior segment structures, including eyelids, cornea, conjunctiva, iris, lens and sclera, facilitating the diagnosis of multiple diseases. Moreover, slit-lamp images have not been widely explored in the deep learning area. Therefore, it is suitable for exploring a novel annotation approach.
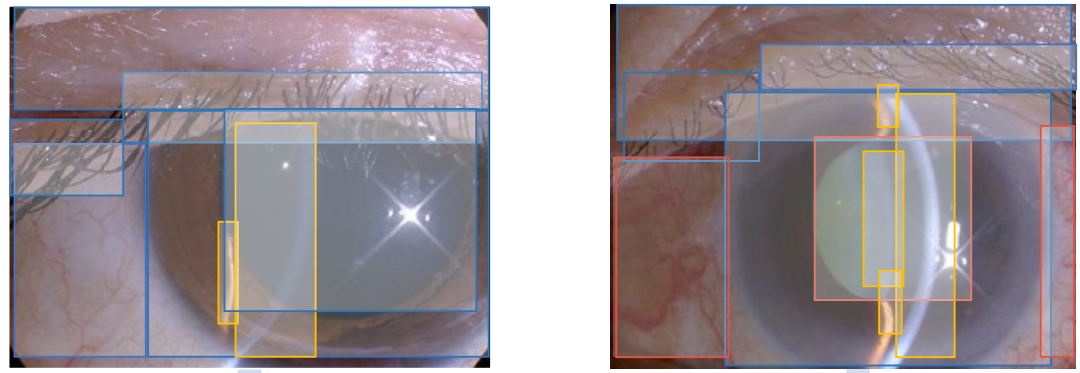
Inspired by genome sequencing, we combined genomics with computer vision, and developed a novel annotation technique named “Visionome”, to establish a densely annotated dataset, based on anatomical and pathological segmentations. In our study, a total of 1,772 slit lamp images of patients with infection-related diseases, age-related diseases, and environment-related diseases were processed by Visionome.
However, the implementation of ‘Visionome’ required a large amount of careful and precise annotation of medical image data. The annotators were also required to have a certain level of ophthalmology knowledge. To fill the talent gap, we established a Medical Artificial Intelligence Alliance to attract clinicians and trained a team of outstanding medical annotators through the recruitment-training-assessment system. We also established a high-standard review and error-correction mechanism to complete the enormous annotation work accurately. We finally generated 1,772 general classification labels, 13,404 segmented anatomical structures, and 8,329 pathological features, yielding 12 times more labels than the image-level classification for a single task.
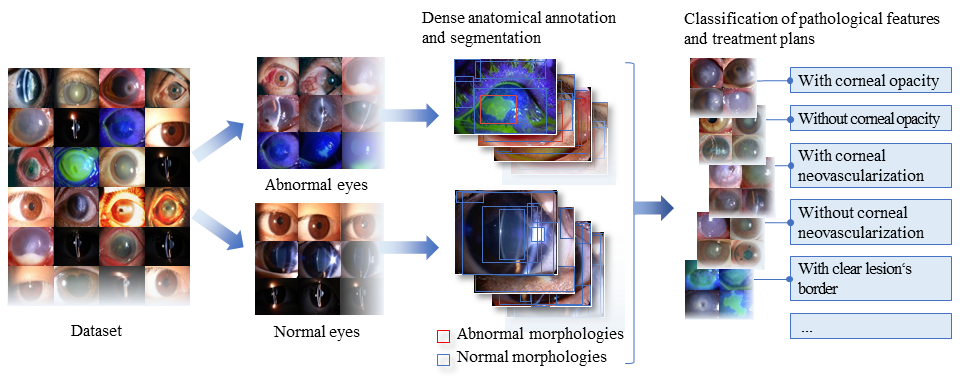
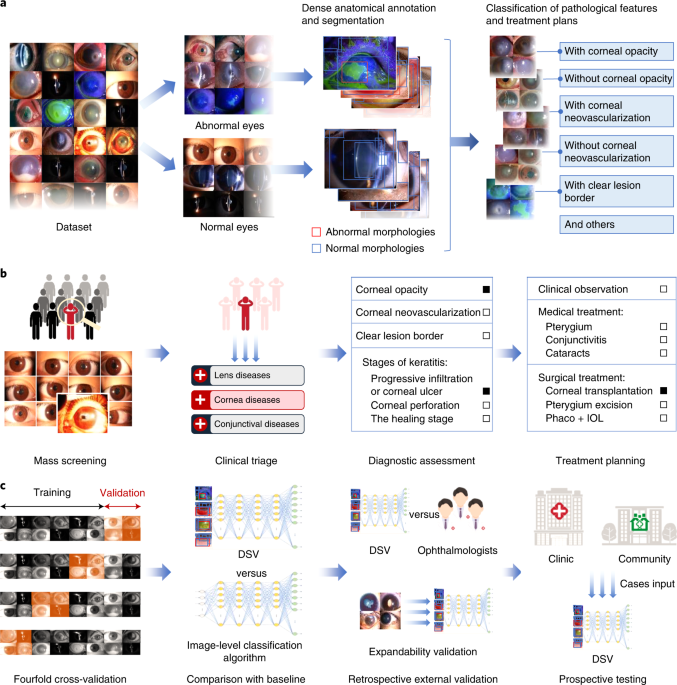
The workflow of Visionome
The performance of the diagnostic system using Visionome (DSV)
Using Visionome, we created an ophthalmic diagnostic system, the “DSV.” The DSV was designed to address four typical clinical scenarios: 1) mass screening to distinguish between normal and abnormal eyes, 2) comprehensive clinical triage to detect 14 lesion locations and ocular structures, 3) hyperfine diagnostic assessment of 22 (10 types) pathological features, and 4) multipath treatment planning based on 7 treatment options. The DSV showed better diagnostic performance than the algorithm that was trained with only the image-level classification labels.
Although the system performed well during internal validation, it was not convincing enough for clinical application. Therefore, we further designed retrospective external validation tests and a prospective test to compare the performance of the DSV with ophthalmologists and to assess the capability of the DSV in real-world clinics. We also established a website to perform these tests and provide patients with remote real-time monitoring of disease conditions. Our system had performance similar to ophthalmologists across four clinically relevant retrospective scenarios, and correctly diagnosed most of the consensus outcomes of 615 clinical reports in prospective datasets for the same four scenarios. When facing diseases that the machine had not learned, including the top 10 eye emergencies, and other common complex diseases, the diagnostic ACC of the mass screening scenario was 84%, demonstrating the adaptability of Visionome.
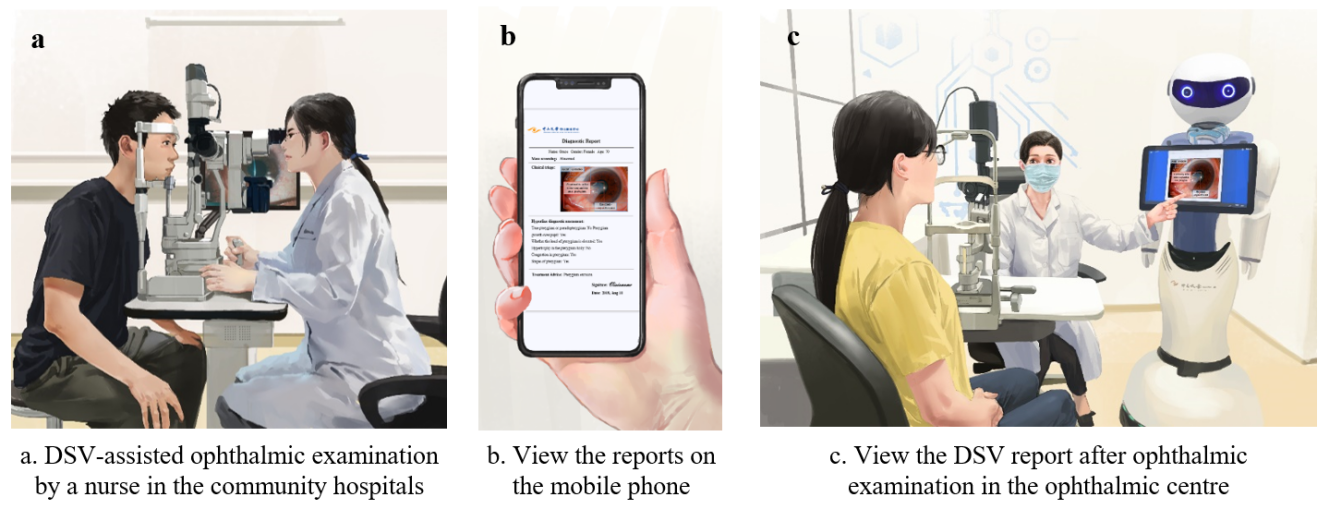
DSV clinical application
The future of Visionome
Visionome has infinite possibilities to become an excellent ‘doctor.’ Using Visionome, a user can obtain a comprehensive multi-region diagnostic report by uploading an image of the anterior segment to the DSV within seconds, promoting active healthcare and a shift in the mindset of clinicians and patients who entrust clinical care to machines. However, the application of Visionome is still limited by the requirements of enormous manual work. To solve the problem, we are now working to create a more convenient annotation technique, which may reduce the workload of medical workers and generate well-curated labels on the premise of accurate annotation. Maybe in the future, healthcare workers without coding expertise can train and upgrade annotation models automatically by a one-stop platform to annotate multi-discipline medical data.
Written by Wangting Li, Yahan Yang, Dongyuan Yun and Haotian Lin.
Follow the Topic
-
Nature Biomedical Engineering
This journal aspires to become the most prominent publishing venue in biomedical engineering by bringing together the most important advances in the discipline, enhancing their visibility, and providing overviews of the state of the art in each field.
Related Collections
With Collections, you can get published faster and increase your visibility.
Implantable wireless communication technologies
Publishing Model: Hybrid
Deadline: Nov 28, 2026
Medical Ultrasound: Emerging Techniques and Applications
Publishing Model: Hybrid
Deadline: Jan 29, 2027


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in