Developing modular and fast responsive systems for the release of pre-stored proteins from mammalian cells
Published in Bioengineering & Biotechnology
Synthetic biology aims to engineer biological systems for the desired controllable response to a selected combination of conditions. By and large the designed systems have been based on the transcriptional response, which is easier to construct and implement but is characterized by a slow response as the desired proteins have to be transcribed and translated1. Recently, protein posttranscriptional modification has been implemented, which considerably accelerated the response, achieving response within minutes2,3, however this type of systems was only able to address production of intracellular proteins, but not secreted or plasma membrane anchored proteins.
Protein secretion is a crucial mediator of an organism's biological activity. Translocation of a protein to the cell membrane or secretion of a soluble protein is one of the key mechanisms in multicellular organisms, either by downstream signaling of other cells through paracrine, endocrine or autocrine signaling, direct cell-cell interactions etc.
In eukaryotic organisms we can broadly distinguish two classes of secretion: the first type is constitutive secretion of proteins, where the protein passes straight to the plasma membrane, without being stored mid-way in a cellular compartment for future release. In this case, as long as the protein is expressed, it will be released from the cell4. The second type is regulated secretion where the secreted proteins are pre-synthesized and stored in specialized cellular compartments (e.g. synaptic vesicles, secretory granules etc.) and released from the cell upon the presence of a release signal. This type of secretion is able to respond faster upon the induction of an appropriate signal, as this type of release bypasses the need for the transcription and translation of the protein, and is therefore crucial in processes where a response needs to occur fast after the change in the environment5. The biosynthetic pathways that generate the specialized storage compartments are currently not fully understood and would require the coordinated action of tens of genes6,7. Additionally, the diverse storage compartments differ in their packing, storage and release mechanisms, making them less suitable for the designed modular synthetic protein release systems.
Compared to storage compartments, a relatively simpler natural system for stalling proteins on the way to the cell exterior occurs for proteins resident in the endoplasmic reticulum (ER), which need to elicit their function inside of the ER and therefore need to be prevented from passing through the ER and post-ER compartments to the cell exterior. This is achieved by the presence of so-called retention signals - short peptide sequences recognized by the cellular trafficking machinery, which returns the proteins back to the ER through the retrotransport system8. Targeting a protein of interest to the ER by the addition of a signal sequence, and retaining that protein inside the ER by the addition of a retention signal would thus position a synthesized protein inside the ER, where it would accumulate. To ensure the release of the protein from the cell, all that one would need to do is remove the retention signal.
The idea how to address this type of engineered systems has been first conceived and proof of the concept tested within the Slovenian iGEM research project in 2016 (http://2016.igem.org/Team:Slovenia/Demonstrate), along with protein signaling logic, which has already been published as a SPOC logic2; nevertheless it took us some time to fine tune the regulated secretion system, perform all required controls explore and demonstrate its potentials.
The team developed two systems where we could control the removal of the retention signal, and thus control protein secretion by highly specific orthogonal viral proteases. These proteases from the Potyviridae plant virus family have been previously shown to be able to provide modular and highly specific tools for controlling signal processing inside of mammalian cells, without interfering with naturally occurring pathways2,3. We opted for the use of these proteases, as they can be split into inactive fragments and rapidly reconstituted with the help of chemically inducible dimerization (CID) domains. The two systems, which we termed membER (for membrane bound ER proteins that feature a KKXX retention signal) and lumER (for luminal ER proteins that feature a KDEL retention signal) were both able to elicit secretion upon CID with an appropriate inducer molecule at a much faster rate than comparable systems based on the induction of gene transcription. Out of the two, the lumER system performed faster, where the protein was detected in the media as early as 30 minutes after the induction. Both systems are orthogonal to each other and could be stimulated multiple times, with different viral proteases.
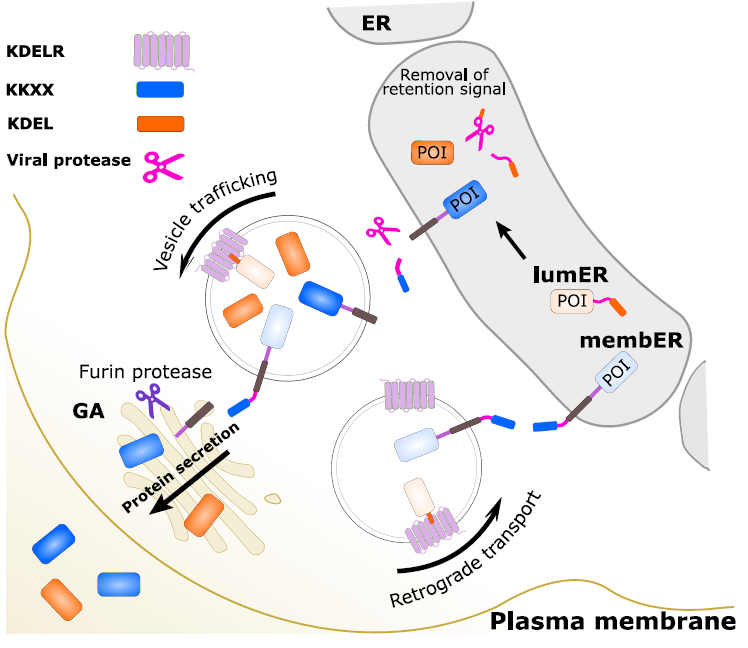
Figure 1: The design and principle of the membER and lumER systems. Proteins of interest (POI) are retained in the ER by the addition of a C-terminal KDEL (for soluble luminal proteins) or KKXX (for membrane bound proteins) peptide sequence. Activation of a viral protease via CID removes the retention signal, releasing cargo protein from the retrograde transport and allows its secretion. Additionally, in the membER system, the POI needs to be released from the membrane by the GA-resident furin protease.
Moreover, we wanted to advance the system further. Since both membER and lumER systems deploy modular orthogonal elements, we wanted to devise the system so that the secretion could be controlled by more than a single input signal. In this approach, the elaborate signal processing elements can be incorporated so that the functional output, in our case the POI secretion, is achieved only when the right combination of input signals is present. Using the membER system, we were able to combine it with our teams’ previously described SPOC system2, where a combination of split viral proteases and complementary coiled-coil peptides were used to regulate the output according to the selected Boolean logic functions. The output of the SPOC system results in the reconstitution of a split viral protease, which in turn drives the removal of the retention signal on our membER constructs.
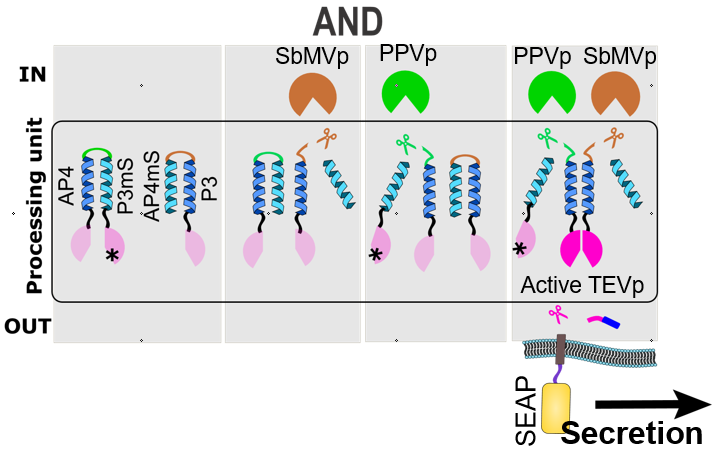
Figure 2: A demonstration of a two-input signal processing according to an AND Boolean logic function using SPOC system, which controls the release of a SEAP reporter protein with the membER system. The highly specific split TEV protease (magenta) is reconstituted only upon the activation of two input proteases, in this case SbMVp (brown) and PPVp (green), which in turn could be regulated by chemical signals. The reconstituted TEVp removes the retention signal (dark blue), leading to the release of the desired protein from the ER and its secretion.
The employment of SPOC is made possible since in the case of membER, the retention signal is facing the cytosolic side of the ER membrane, and can therefore be accessible by cytosolic proteins. The situation is different with the lumER system, where the retention signal needs to be present on the luminal side, making it impossible to target with cytosolic proteins. Therefore, several of the logic processing constructs needed to be optimized for use inside the ER, by modifications to prevent nondesired glycosylation, endowing constructs with additional retention signals, and co-transfecting the cells with additional proteins involved in the retrotransport to increase the ER-retention capacity. This ensured that the secretion with both of our systems could be regulated by two input signals according to the desired logic operations, thereby providing the much needed regulation to control protein secretion. This could be especially useful for the implementation of a negative feedback, to prevent overactivation of a biological process, or when a more stringent control would be required.
This new type of regulation could be used in diverse types of applications, such as for the regulation of therapeutic proteins, both soluble and membrane bound. For soluble proteins, we opted for the release of human insulin9 and anti-inflammatory cytokine interleukin 1010, both valuable therapeutic proteins, where secretion needs to occur ASAP after the specific (patho)physiological condition is achieved. For therapeutically relevant membrane proteins, membER system was demonstrated to control positioning of the anticancer chimeric antigen receptor (CAR) to the plasma membrane, upon the chemical activation.
In summary, we have developed two systems for the release of both membrane-bound and soluble proteins from the ER, where both systems outperformed systems based on the initiation of protein expression in terms of the secretion kinetics. The systems also do not rely on protein aggregation or competitive binding, as do previous systems based pre-storing proteins in the ER11,12, making them applicable to a broader area of biological applications, as well as allowing for the combination of our systems with intricate signal processing modules.
References
- Verbič, A., Praznik, A. & Jerala, R. A guide to the design of synthetic gene networks in mammalian cells. FEBS J. 288, 5265–5288 (2021).
- Fink, T. et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells. Nat. Chem. Biol. 15, 115–122 (2019).
- Gao, X. J., Chong, L. S., Kim, M. S. & Elowitz, M. B. Programmable protein circuits in living cells. Science (80-. ). 361, 1252–1258 (2018).
- Baldwin, G. S. Post-translational Processing of Gastrointestinal Peptides. Physiology of the Gastrointestinal Tract vol. 1 43–63 (2012).
- Scott, M. & Sandkvist, M. Toxin secretion systems. in The Comprehensive Sourcebook of Bacterial Protein Toxins 83–105 (Elsevier, 2006). doi:10.1016/B978-012088445-2/50010-X.
- Guest, P. C. Biogenesis of the insulin secretory granule in health and disease. in Advances in Experimental Medicine and Biology vol. 1134 17–32 (2019).
- Omar-Hmeadi, M. & Idevall-Hagren, O. Insulin granule biogenesis and exocytosis. Cell. Mol. Life Sci. 78, 1957–1970 (2021).
- Stornaiuolo, M. et al. KDEL and KKXX retrieval signals appended to the same reporter protein determine different trafficking between endoplasmic reticulum, intermediate compartment, and golgi complex. Molecular Biology of the Cell vol. 14 889–902 (2003).
- Hay, C. W. & Docherty, K. Enhanced expression of a furin-cleavable proinsulin. J. Mol. Endocrinol. 31, 597–607 (2003).
- Saraiva, M., Vieira, P. & O’Garra, A. Biology and therapeutic potential of interleukin-10. J. Exp. Med. 217, (2020).
- Rivera, V. M. et al. Regulation of protein secretion through controlled aggregation in the endoplasmic reticulum. Science (80-. ). 287, 826–830 (2000).
- Boncompain, G. et al. Synchronization of secretory protein traffic in populations of cells. Nat. Methods 2012 95 9, 493–498 (2012).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in