Early-life microbiota diversification facilitates enterocyte proliferation
Published in Microbiology
The adult gastrointestinal tract is colonized by trillions of bacteria that influence cellular homeostasis through microbial metabolism of dietary and hepatic compounds. The microbiota ecosystem and associated metabolic networks are gradually established in the first years of life (2-3 years) beginning at birth. In healthy term infants, lactic-acid producing bacteria typically colonize the gut shortly after or during delivery and gradually increase in abundance with limited diversity throughout infancy. Delivery at earlier gestational ages (pre-term) is associated with expedited microbiota diversification and altered metabolic profiles in the gut. Evidence from germ-free mouse models has shown that microbial acquisition in the gut facilitates tissue growth and organization of the crypt-villus architecture associated with a mature intestinal tract. However, it remains unclear if distinct patterns of term and pre-term microbiome maturation confer functional changes in the developing small intestine.
Two neonatal intensive care unit physicians, Dr. Oleksandr Kudin and Dr. Josef Neu had access to fecal samples from term and pre-term patients and shared our interest in elucidating host response to neonatal microbiota. Dr. Kudin joined our research group as part of his research fellowship, and together we optimized organoid cultures for in vitro screening of microbial metabolites from patient fecal samples. Simultaneously, other publications demonstrated that lactate (derived from lactic acid producing bacteria in the term gut or from ingested milk) induces enterocyte proliferation, while structural proteins from bacteria (lipopolysaccharide, flagellin) activate toll-like receptor signaling and inflammatory responses. Given this evidence, we expected pre-term microbial communities to induce an inflammatory response while term microbial metabolites would facilitate epithelial maturation.
We first characterized microbial and metabolic composition in term and pre-term cohorts. As expected, term samples harbored microbial communities characterized primarily by lactic-acid producing bacteria (LAB). In contrast, the pre-term microbiome was more diverse and lacking prominent LAB species. The pre-term cohort exhibited a significantly higher metabolic capacity, notably including significant increases in secondary bile acids. These biotransformed bile acids have been shown to increase proliferation and epithelial regeneration following chemical-induced injury. Mapping metabolomics profiles to microbial KEGG pathways showed a significant increase in amino acid and vitamin biosynthetic pathways with a concomitant increase in overall metabolite concentrations.
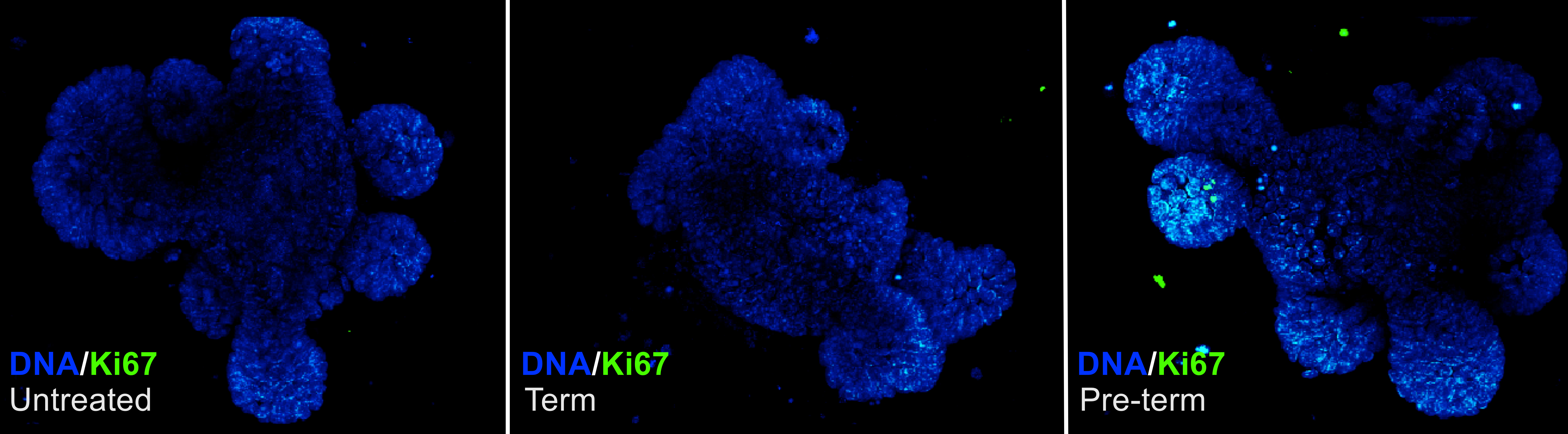
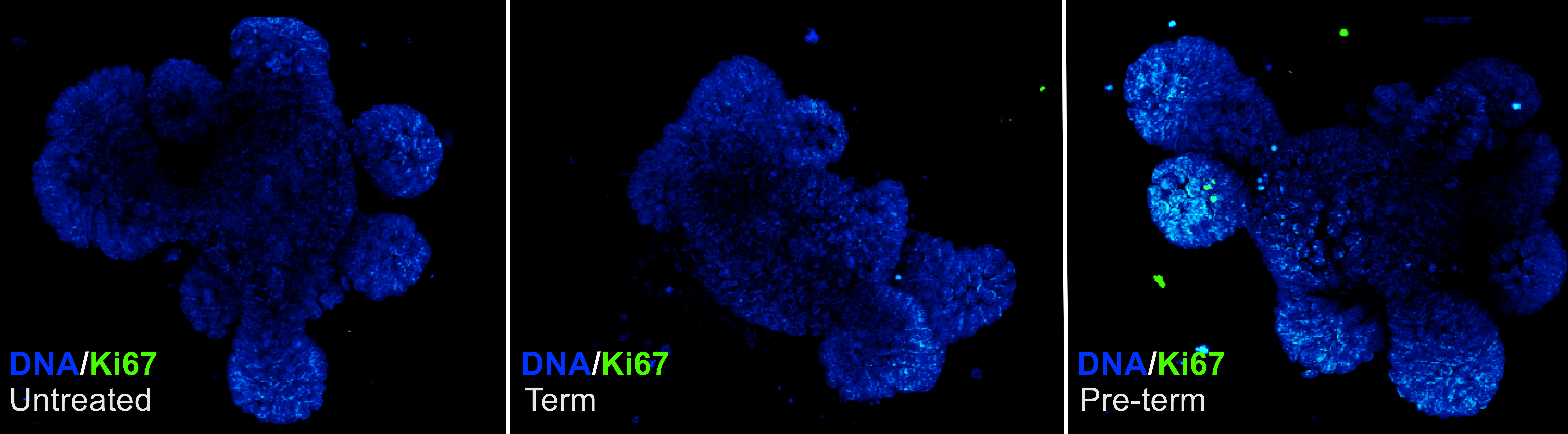
To examine how these metabolites functionally affect the intestinal epithelium, we cultured small intestinal organoids with term and pre-term fecal metabolites. These organoids form “mini-guts” derived from intestinal stem cells which recapitulate epithelial morphology and cellular heterogeneity. Surprisingly, when cultured in the presence of pre-term fecal metabolites, we observed increased proliferation (fig. 1) and stem cell activation relative to term fecal metabolites without activation of inflammatory signaling pathways.
Organoid models are an increasingly common and powerful tool for defining host-microbe interactions intrinsically linked to the biology of the intestinal epithelium. Recently, human organoids have been used to identify the functional effects of individual microbial metabolites and characterize mutational profiles after exposure to bacterial genotoxins. Our study demonstrates the capability to induce functional changes associated with entire microbial ecosystems in vitro using host fecal material. This approach may help clarify complex metabolic interactions between host and microbes in intestinal diseases such as colon cancer.
For more details, please refer to our paper.
Follow the Topic
-
BMC Microbiology
This is an open access, peer-reviewed journal that considers articles on all microorganisms - bacteria, archaea, algae and fungi, viruses, unicellular parasites and helminths.
Related Collections
With Collections, you can get published faster and increase your visibility.
Mycobacteria: advances in physiology and mycobacterial diseases research
Mycobacteria are a large and diverse group of microorganisms, known for their adaptability and resistance to environmental stresses, particularly due to their unique thick, hydrophobic cell wall rich in mycolic acids. While most mycobacteria are not pathogenic, some of them (e.g. Mycobacterium tuberculosis or Mycobacterium leprae) have evolved to become very successful human pathogens, with complex pathogen-host interactions that are still poorly understood. Often because of their unique physiological properties that facilitate survival in different environments, persistence in a latent state within the host and evasion of host immune responses, mycobacteria pose significant challenges to global public health. A greater understanding of the physiology and mechanisms underlying mycobacterial infections is essential to develop effective treatments, diagnostics and preventive measures, especially considering the rise of drug-resistant mycobacterial strains, and the global burden of tuberculosis- and non-tuberculosis-related diseases. Recent advances in understanding mycobacterial basic biology and physiology, pathogenesis, and immune evasion are paving the way for novel therapeutic strategies, including the development of new vaccines, as well as new molecular diagnostics.
In support of United Nations’ SDG 3 (Good health and well-being), BMC Microbiology presents the Collection Mycobacteria: advances in physiology and mycobacterial diseases research. This Collection invites research on mycobacteria, focusing on advances in understanding their physiology, pathogenesis, and the diseases they cause. We welcome submissions that explore host-pathogen interactions, immune system responses, drug-resistance mechanisms, and innovative approaches for the prevention, diagnosis and treatment of mycobacterial infections. Research without a clear focus on microorganisms or pathogen-host interactions will not be considered. We will consider research articles that investigate, but are not limited to, the following topics:
- Physiology, metabolism, stress response mechanisms, adaptation to environmental stresses, sources of transmission and environmental persistence of mycobacteria
- Mycobacterial pathogenesis and immune evasion
- Innate and adaptive cellular immune responses to M. tuberculosis and non-tuberculous mycobacteria (NTM) infections
- Drug-resistance mechanisms, persistence and dormancy in mycobacteria
- Mycobacterial reactivation and transmission
- Host-pathogen interactions in mycobacterial diseases
- Advances for combating mycobacterial co-infections
- Conserved and specialized functions of Type VII secretion systems (T7SS) (ESX or ESAT-6 systems) of mycobacteria
- Advances in molecular diagnostics and biomarkers for the detection of mycobacterial infections, disease progression and cure
- Vaccine development for mycobacterial infections
- Novel drug targets and therapeutic strategies against mycobacterial infections
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Jul 17, 2026
Microbes in built environments
The microbial communities (e.g. bacteria, fungi, viruses) that inhabit human-built environments are diverse, dynamic and complex, and they play a significant role in human health and wellbeing. From indoor air quality to the surfaces and building materials we touch daily, we interact with a range of microbes that come from a variety of sources such as other humans, pets, plants, pests, and other outdoor sources. Investigating and understanding these interactions require interdisciplinary approaches that integrate microbiology, ecology, and engineering principles, among other disciplines. As our understanding of microbial dynamics within built environments advances, it becomes increasingly clear that built-environment microbes and microbiomes can strongly affect human health, either positively or negatively.
Recent research has highlighted the potential importance of indoor microbial diversity in influencing human health (e.g. respiratory health, allergies, and the spread of multidrug-resistant organisms, particularly in healthcare settings). Innovations in monitoring and modelling microbial communities are opening new avenues for designing healthier indoor environments and mitigating risks from unwanted microbial exposures. Ongoing research in the field has the potential to yield novel insights into how these microbes respond metabolically to certain stressors and how we can leverage that knowledge. Additionally, a better understanding of microbe-host interactions could aid in developing novel approaches in building design and public health initiatives. Adaptive antimicrobial methods based on real-world conditions (such as occupancy, ventilation, and microbial indicators) offer a promising pathway to control microbial risks while minimizing unintended impacts.
In support of the United Nations’ Sustainable Development Goal 3 (SDG 3, Good Health and Well-Being), BMC Microbiology launches the Collection Microbes in built environments. This Collection invites research articles that contribute to the understanding of microbes and microbiomes in human-built environments, encompassing microbial ecology, diversity, metabolism, and their implications for human health. Submissions are welcome on topics ranging from indoor microbial communities to the impact of design on microbe-host interactions in the built environment. Research without a clear focus on microorganisms, microbial communities, or microbiomes in built environments will not be considered. We invite researchers and experts in the field to submit research articles covering a broad range of topics, including, but not limited to:
- Interactions between microorganisms/viruses and hosts in built environments
- Microbial physiology, metabolism, adaptation and diversity of the built environment
- Impact of microbes in the built environment on human health and diseases
- Antimicrobial resistance of the built environment
- Holistic pathogen defence in the built environment
- Microbial ecology of the built environment
- Microbial adaptation and evolution in response to anthropogenic disturbances in built environments
- Viruses in the built environment
- Food surfaces and disinfection in the built environment
- Role of indoor air quality, ventilation and HVAC systems in microbial community dynamics
- Impact of surface and building materials on microbial colonisation
- Bioremediation and microbial management in built environments
- Interventions to promote/discourage microbial exposures in built environments
- Environmental monitoring methods of microbes in built environments
- Adaptive antimicrobial strategies in built environments
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Oct 22, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in