Efficient interhomolog gene conversion in both somatic and germline cells of Drosophila.
Published in Bioengineering & Biotechnology

Background context: In 2015, Valentino Gantz and I (Ethan Bier) published a paper in Science in which we demonstrated the first proof-of-principle for a CRISPR-mediated gene-drive system in metazoans that have specialized reproductive cell lineages (Gantz and Bier, 2015). The history behind that paper is Valentino wanted to develop a method to generate dominant alleles to aid the recovery of mutations in pioneer model organisms such as those of potential value for evolution and development (Evo/Devo) studies. His idea was simple: insert Cas9 and a guide RNA (gRNA) into the genome at the site where the Cas9/gRNA complex is programmed to make double strand breaks (DSB) the genome. Repair of such DSB using the homolog chromosome as a repair template should then copy the element to that chromosome and generate visible phenotypes in somatic cells of the body (e.g., wing venation patterning phenotypes). As we discussed this concept further, we also realized that such a system should behave similarly in the germline and that by measuring the frequency of its transmission we should to be able to obtain an estimate for the copying efficiency of such a process. We named this strategy the mutagenic chain reaction (MCR) in analogy to the polymerase chain reaction since both of these self-amplifying methods lead to a 2-fold increase in the desired template per cycle (PCR) or per generation (MCR).
The first prototype MCR gene-drive element was inserted into the yellow locus in Drosophila, and produced a full body mutant phenotype (lighter yellow body instead of the normal darker pigmentation) consistent the element copying itself between chromosomes as was evident from its efficient transmission through the germline. Although we imagined that the similar frequencies of copying were likely to be occurring in somatic cells in female flies inheriting the X-linked MCR element (yellow body phenotype of F1 females) as we observed through the germline (>90%), we recognized that we could not rule out the possibility that there could be different balances of repair outcomes in the germline (largely mediated by the homology directed repair - HDR - pathway) versus in somatic cells (e.g., either HDR or the mutagenic non-homologous end-joining - NHEJ - pathway). Indeed, we had identified NHEJ mutations being generated in this study and found by PCR analysis that F1 MCR-bearing females were mosaic for the element indicating that they had some mix of HDR versus NHEJ induced yellow- alleles in somatic cells, although the relative proportions of those events could not be resolved quantitatively.
As the gene-drive field matured and the prevalence of NHEJ induced alleles generated by maternal inheritance of Cas9/gRNA was raised as a potential obstacle to generating efficient drive systems via both sexes (a challenge that has since been overcome by several different approaches), a consensus emerged that the NHEJ repair pathway dominated over HDR in somatic cells and that HDR was only efficient in germline cells where meiotic cell lineages where recombination between homologs takes place. This view was consistent with the relatively low ratios of HDR/NHEJ that researchers observed in mammalian cells when attempting to perform Cas9 mediated editing using exogenously provided DNA repair templates. Also, the recognition that the normal biological function of HDR repair of DSB in somatic cells to preserve genome integrity following DNA replication, provided a rationale for why NHEJ dominated in somatic cells during non-replicative phases of the cell cycle, particularly given the fact that mutations that increase inter-homolog chromosome repair and thereby result in so-called loss-of heterozygosity can often to lead to pathological outcomes such as cancer (e.g., by rendering heterozygous mutations in tumor suppressor genes homozygous).
Current Study: As summarized above, the various arguments for or against potentially efficient inter-homolog HDR in somatic cells left the question unresolved. The goal of our new study was to develop genetic sensors to measure the frequency of inter-homolog repair in somatic cells, which I had actually included as an objective in an NIH grant submitted in early 2015 prior to Valentino's Science paper being published. As has often been the case, Valentino subsequently improved on the design I had proposed for such a detector element and Zhiqian Li further refined and developed prototypes of these newly-dubbed "CopyCatcher" elements.
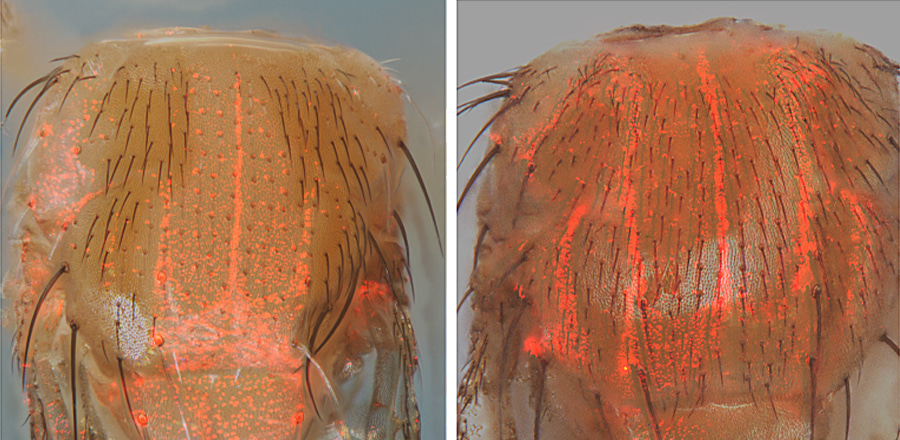

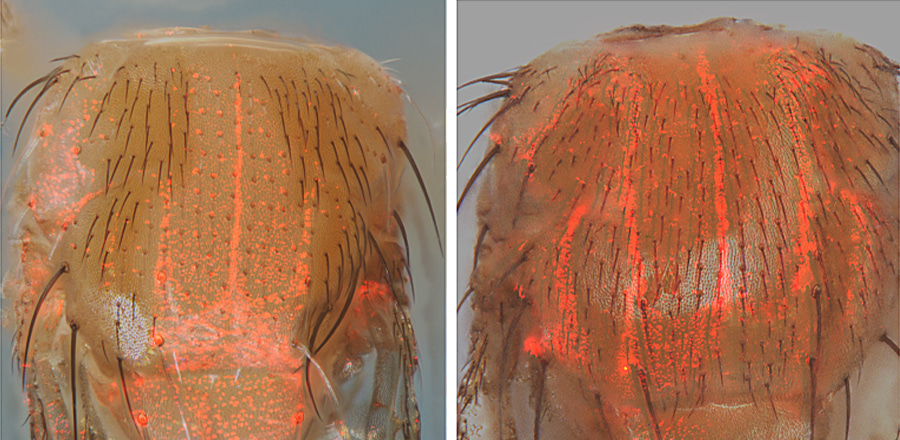
Caption: Red fluorescent detector proteins in fruit flies reveal detection from an actual copying experiment (left) and how perfect copying would appear (right). Fluorescent cells in the left panel also lack the function of a pigmentation gene called “pale” due to copying of the CopyCatcher element which eliminates function of both copies of the pale gene. Control flies in the right panel, which have only one copy of the pale gene disrupted by insertion of the CopyCatcher and one functional copy of the gene, have bright red cells and normal bristle pigmentation thought the body. Credit: Zhiqian Li
CopyCatchers are designed much like any other CRISPR-based drive elements in that they carry a gRNA that directs Cas9 cleavage at their site of genomic insertion. These elements are inserted into introns of target genes where the hope was that small insertions or deletions of sequence caused by NHEJ repair would not produce any detectable phenotype. CopyCatchers also carry a splice acceptor site followed by a T2A-DsRed transgene so that such elements express the fluorescent marker in the same pattern as that of the endogenous gene. CopyCatcher elements were then combined with upstream mutations located near the initiator ATG codon (ATG– mutations) that eliminated expression of both the endogenous gene and DsRed. These mutations are far enough (>1 kb) from the CopyCatcher that HDR-mediated somatic gene conversion (SGC) events would be very unlikely to co-copy the ATG– mutation with the element (typically local gene conversion tracts are <100-200 bp in length).
If a chromosome carrying the closely linked ATG– CopyCatcher alleles and placed in trans to a wild-type functional allele of the target gene, then in the presence of of an unlinked Cas9 source SGC events should result in the CopyCatcher being freed from the ATG– mutation, thereby restoring DsRed expression. Since CopyCatchers insertions also interrupt function of the target gene, they should also eliminate gene activity in DsRed+ cells (and their mitotic descendants) that have undergone copying events. This clonal copying events can be detected by coincident patterns of fluorescence and mutant adult phenotypes (e.g., loss of eye or body pigmentation).
We were initially surprised by Zhiqian's CopyCatcher tests and how efficiently they copied in cells giving rise to the adult eye and thorax, where ~30-50% of cells displayed the concordant DsRed+ and clonal mutant phenotypes. These substantial SGC frequencies could be further enhanced by optimizing delivery of the Cas9 enzyme (low expression in the early embryo and higher levels during later developmental stages) or by either RNAi knockdown or over-expression of a small set of modifier loci identified in a small-scale pilot screen of candidate interacting genes. I must admit to being fairly pleased to see that the original hypothesis that Valentino and I put forward was actually pretty close to the mark. Many of those yellow mutant cells in his F1 females were indeed likely to be the result of the MCR copying after all.
Zhiqian also went on to show that a GFP+ insertion element in human epithelial kidney cells could also be copied from one chromosome to its homolog, albeit at lower frequency 4-8% than observed in flies. In a pilot genetic screen, she found that was possible to increase SGC in flies by knock-down of specific genes and that similar manipulations in human cells also increased copying These findings suggested that the underlying genetic mechanisms for inter-homolog repair may be shared between flies and human cells.
The work continues: In future experiments Zhiqian and her colleagues will be trying to further improve SGC frequencies in human cells. Despite the concerns outlined above that HDR is not as efficient to repair DBS in somatic mammalian cells as NHEJ, there are reasons to be optimistic about such prospects. First, there is evidence when DSB are made that at the corresponding region of the homologous chromosome becomes paired with the broken chromosome during the repair process. Also, there are some cell lines in which chromosome arms are constitutively paired along their entire length as is the case for chromosomes in Drosophila where SGC is more efficient.
If higher frequencies of somatic gene conversion could be achieved in human cells, new gene therapy strategies to correct genetic disorders might be envisioned in which pathogenic lesions on one chromosome are repaired using wild-type sequences on the homolog chromosome. Such schemes could convert affected individuals carrying two different mutant alleles of a gene into asymptomatic carriers for the disease. This restoration of wild-type gene function from an endogenous copy of the locus would be a great improvement over current strategies in which a cDNA copy of the mutated gene is typically inserted into an ectopic chromosomal site under the control of synthetic regulatory elements that do not recapitulate either normal patterns or levels transgene expression.
The new study by Zhiqian and colleagues resolves an open question regarding the efficiency of inter-homolog templated of HDR in Drosophila and sets the stage for translating these encouraging results in flies to humans. This satisfying story thus weaves an interesting basic science result together with a potential for future impactful new therapies to benefit human health.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in