Evolutionary Insights into Cockle Transmissible Cancers
Published in Cancer

The story began by collecting almost 7,000 cockles along the Atlantic coasts of Europe and North Africa: from Morocco to Russia! We were on the hunt for cockle cancer because we knew that they suffer from transmissible or contagious cancers (Metzger et al., 2016). In particular, cockle transmissible cancers are leukemias, that is, blood tumors, in which the tumor cells can leave the animal they infect and travel through the oceans until they infect a new animal. No virus, bacteria, or parasite infects the new host, it is the cancer cell itself.
So we began to diagnose cockles to find out that only 390 cockles were leukemic. Surprisingly, all belonged to Southern European countries while the Northern European countries seem, in principle, to be free of this disease.
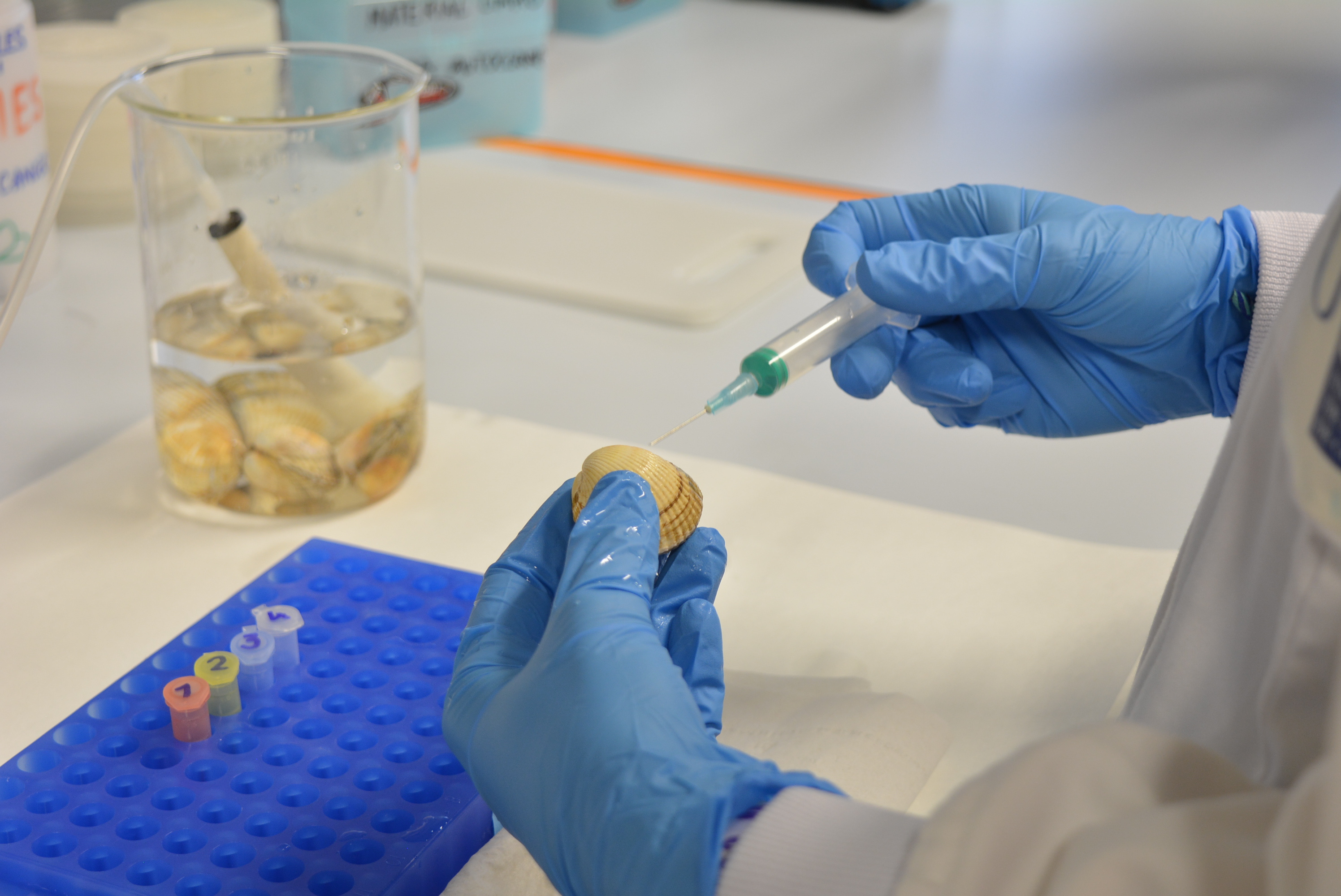

As we were interested in investigating the genetic causes of this contagious cancer, the first thing we had to obtain was the reference genome of the cockle, since it did not exist. For that, we applied a multiplatform DNA sequencing approach in a healthy adult male cockle carrying a standard karyotype with 19 chromosome pairs. Haploid genome size was estimated at 790 Mb –a third of the human genome– and we reconstructed a chromosome-level genome (N50=39.6 Mb). The transcriptome was constructed using seven tissues and the gene annotation resulted in a 42-Mb exome with 14,055 protein-coding genes.
We were then ready to start our journey into the genomics of cockle transmissible cancers. To do so, we sequenced the whole genomes of 61 cancer, and 462 non-cancer animals.
Two stories from the same book
A first outlook into the nuclear genomes of these singular marine leukaemias confirmed the two independent cancer lineages consistently match two histological phenotypes. However, the phylogenetic analysis of mitochondrial genomes did not reveal those two cancer lineages. Instead, nine mtDNA lineages were inferred. The presence of multiple mtDNA lineages within each phenotypic-nuclear cancer lineage indicates that mitochondria from transient hosts have repeatedly been acquired. Cancer lineages evolved from one ancestral cell, and they propagate by asexual reproduction resulting in the accumulation of deleterious mutations that lead to a fitness decline. However, capturing host mtDNA seems to be the mechanism by which cockle contagious cancers have avoided their extinction.
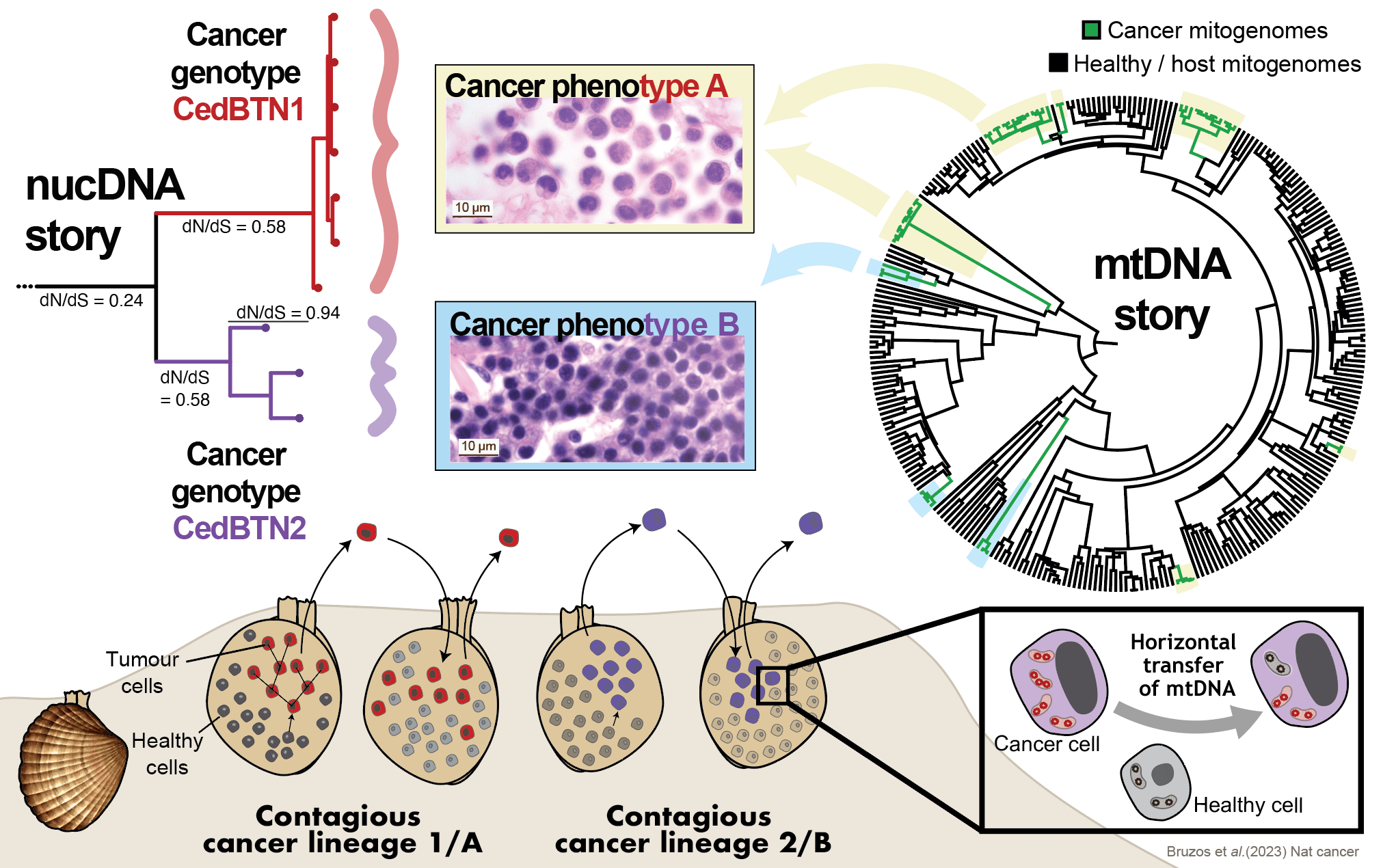
Genomic Archaeology: from past to present
Since cancer is a genetic disease, we were interested in finding the genetic mutations that caused a cell (from a cockle that lived thousands of years ago) to transform into a tumor that acquired the ability to spread among the population.
First of all, to shed light onto the origins of these two cockle cancer lineages, we inquired about the tissue of origin of these cancer cells. The normal tissue stem cell of this cancer is an ongoing debate in the scientific community with various hypotheses proposed such as the hemolymph, gills, or gonads. Comparing the gene expression of the two cancer lineages and several healthy cockle tissues, we found that both types of cancer lineages shared a similar expression profile and that both are similar to that of hemolymph. This leads us to deduce that both lines of cancer originated in the hemolymph. The recurrent origin in this tissue may reflect the cancer's ability to exploit the transmission opportunity offered by the circulatory system.
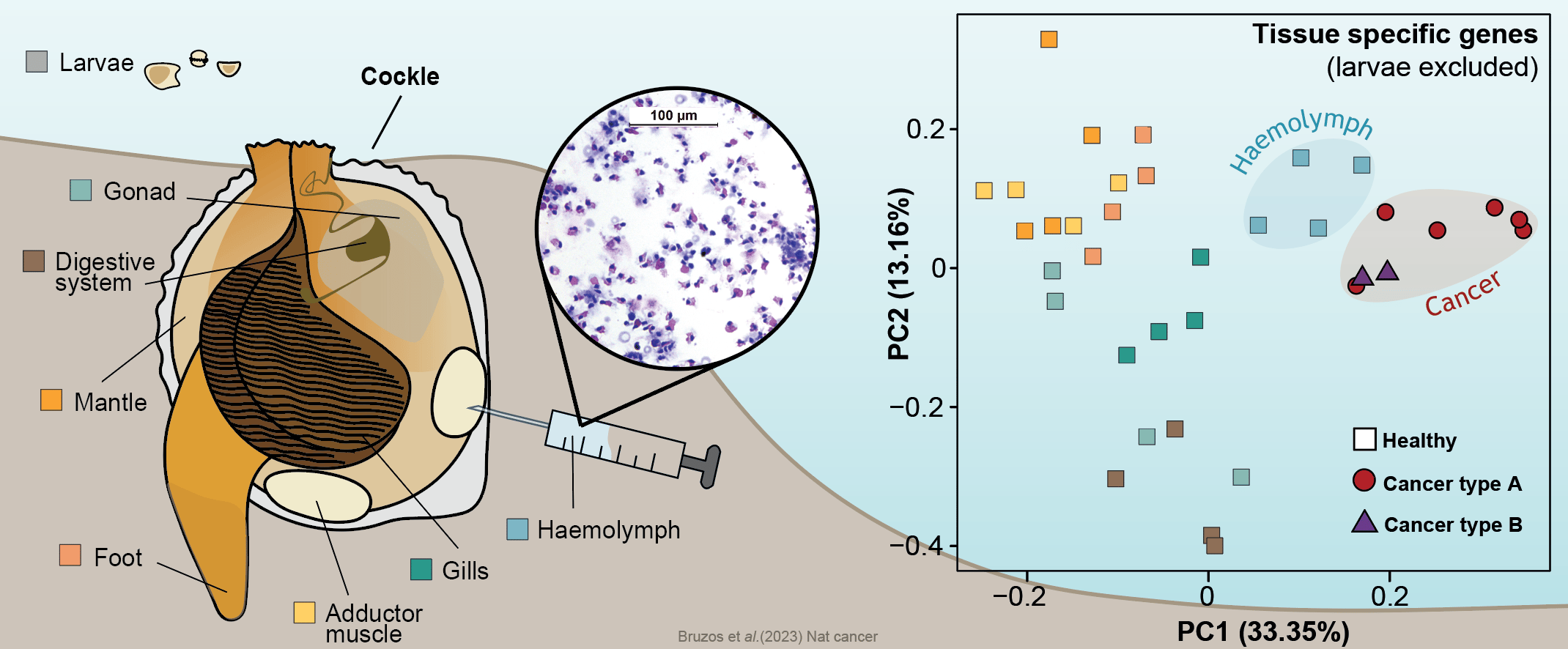
The transcriptome of different cockle tissues/organs were analyzed to inquire about the normal tissue that originated cockle contagious cancer lineages. Data supports a hemocytic origin of both lineages which may reflect the cancer's ability to exploit the transmission opportunity offered by the circulatory system
Focusing on the genetic mutations that shaped the evolution of a cell to become a contagious cancer cell, the main cause that we have found in both cancer lineages is a duplication of the complete genome. When genetic material is duplicated, the contribution of genes to cellular functioning is decompensated, which deregulates other genes and can trigger the origin of a tumor. This is common in many human cancers and it indicates that invertebrate cancers can also arise in the same way.
Following this initial duplication event, many other structural mutations have caused the cancer to evolve. We see that the number and structure of chromosomes in cancer cells is very chaotic. While the normal number of chromosomes in a cockle is 19 pairs, cancer cells have a highly variable number of chromosomes, which can reach several hundred. Their structure is greatly altered, with highly fragmented chromosomes and regions that have multiplied several times.
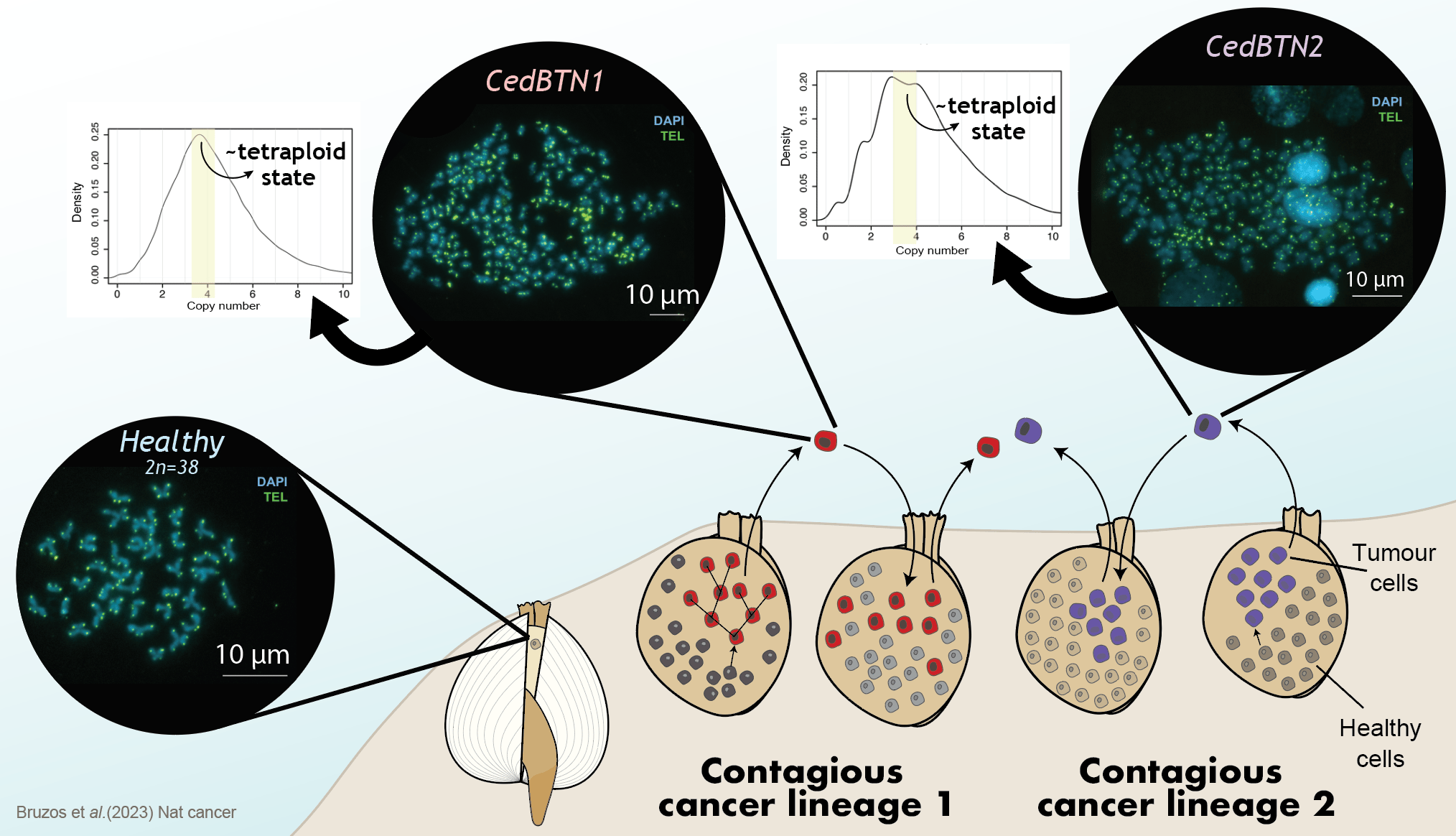
We also observed other mutational events such as a signature of unknown etiology described in human myeloid and brain tumors. Or a very common type of mutation that is a consequence of the loss of a gene called MGMT, which is responsible for repairing a modification in DNA called base alkylation which is a process necessary for the correct functioning of the genomes. Among others, these findings represent one of the most important points of our work: how is it possible that a cancer with such a damaged genome could survive for thousands of years? This question has no answer yet and will be the focus of future research.
To sum up, studying these mutations and the mechanisms by which cancer cells overcome the effects of such instability promises to broaden our understanding of the conditions required for tumors to survive and adapt over the long term.
This is just a start, want to know about the cancer age? Want to know about co-infections? Take a look at our paper where you can read more details and findings:
- Bruzos AL et al. (2023). Somatic evolution of marine transmissible leukaemias in the common cockle, Cerastoderma edule. Nature Cancer. DOI: 10.1038/s43018-023-00641-9
Want to know more? Check this independent study by researchers in the United States, who examined a similar transmissible cancer affecting the soft-shell clam, another species of shellfish found on the East coast of North America.
- Hart SF et al. (2023). Centuries of genome instability and evolution in soft-shell clam, Mya arenaria, bivalve transmissible neoplasia. Nature Cancer. DOI: 10.1038/s43018-023-00643-7
They also found that, like in cockles, clam transmissible cancer originated from a hemocyte and its genome displays widespread duplication, reorganization, and mutation. They did not observe mtDNA capture, but they did observe amplification of a portion of the mtDNA genome in the clam cancers, and they found an error-prone polymerase mutational signature that is different than the cancer-specific signatures seen in cockles. Taken together, these studies reveal divergent and convergent mechanisms that may have allowed each cancer to arise and survive long-term as a contagious cancer in their respective host populations.
Main funding: European Research Council (ERC) Starting Grant SCUBA CANCERS (No.716290)
Follow the Topic
-
Nature Cancer
This journal aims to provide a unique forum through which the cancer community will learn about the latest, most significant cancer-related advances across the life, physical, applied and social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Cancer Neuroscience: from mechanisms to therapy
Publishing Model: Hybrid
Deadline: Jan 30, 2027



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in