Fecal Immunochemical Test samples are suitable for virome studies; gut virome is associated with the individual’s lifestyle
Published in Microbiology and General & Internal Medicine

In the last decades, we have come to an understanding that microbial communities that live in and on our bodies, our microbiome, are not silent bystanders as previously believed, but rather play an important role in our health and well-being. This is especially true regarding gut microbes. Our gut microbiome is implicated in health issues far beyond the gut and has been linked to many immunological disorders. Much of the research on gut microbiome is focused on bacteria, but the evidence that gut viruses regulate these bacteria and thus also impact our health is persistently accumulating.
Colorectal cancer rates are increasing worldwide, and more and more early-onset cases are being reported. The gut microbiome is likely to play an important role in carcinogenesis, and several bacteria are often associated with the disease. To screen for colorectal cancer, the fecal immunochemical test (FIT) is widely used. The test is based on the detection of the hidden blood in the stool, and although it is extremely good in detecting cancer cases, it misses 6 out of 10 people with pre-cancerous lesions.
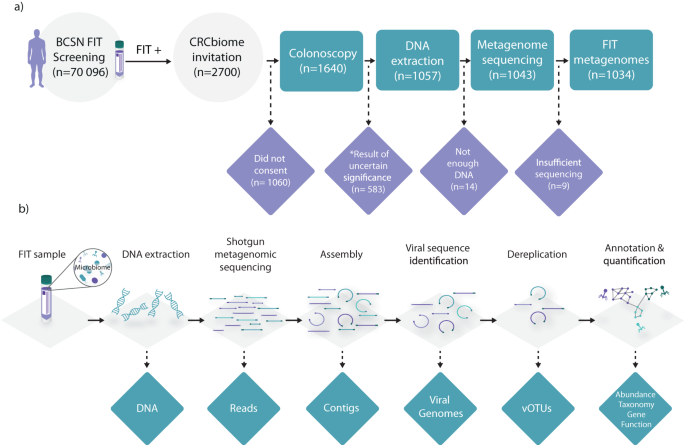
The CRCbiome project team led by Prof Rounge is set on extracting the information about the gut microbiome from FIT samples and adding it to the bleeding status when assessing gut health. This additional information could make the FIT test better at catching potential issues. We are convinced that although FIT sampling kits were not designed for microbial analyses, it is possible to analyze the microbiome using these samples. In this work, we used FIT samples collected from over 1000 people and demonstrated that we indeed could learn about the viral community in their guts. Showcasing the applicability of FIT samples to microbiome studies is extremely important, - it paves the way for large-scale studies in many countries where thousands of these tests are performed daily. We also discovered the vastly diverse and highly individual community of viruses, many of which were strongly indicative of lifestyle. For instance, viral communities varied greatly based on whether one was smoking or not, how much time per week they exercised, and how many fiber-rich products they included in their diet.
We believe that by using the information on microbes extracted from FIT samples, we will be able to better pinpoint who is at risk of colorectal cancer development and act preventively.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in