Forward-predictive SERS-based chemical taxonomy for untargeted structural elucidation of epimeric cerebrosides
Published in Chemistry, Ecology & Evolution, and Materials
This story started in early 2023 when I worked alongside a computer science student specializing in machine learning. We were having a debate on whether models can predict unknown data that are outside the training dataset. At first, it seems impossible as machine learning models are known to identify what it has been trained on, not beyond. A model train to identify cats and dogs no doubt can identify cat and dogs with high accuracies, but not birds and rabbits. The "unknown" in question is not the same as extrapolation outside of a continuum, I want to predict "new" molecules and their labels beyond the boundaries of the training dataset. Naturally, I thought of biologists who have depended on taxonomy to systematically classify "new" species based on their relationship to existing species. A characteristic that differentiates the species is determined in the different "clades" of the taxonomy. This is how the hierarchical 5-level machine learning framework model is established. Using this methodology, we can now achieve untargeted structural elucidation and identification of new and unknown chemicals/molecules using just their SERS spectra (notably even at LOD=10−10 M). This signifies an important advancement for SERS-based sensing where the identity and concentration of chemicals are often unknown.
To begin, we look at our target molecules: epimeric glucocerebrosides (GlcCerX:Y) and galactocerebrosides (GalCerX:Y) differ in the spatial orientation of their C4 OH-groups (C4 site of isomerism) in their glycosyl/ galactosyl moiety and consist of ceramides moieties with varied carbon chain length (X) and saturation degrees (Y). Due to their structural diversity, they possess different bioactivities and play distinct functional and constitutional roles in cellular signaling and metabolism. Each has its own set of five structural characteristics, allowing us to identify them i.e., (1) the presence or absence of epimers, (2) monosaccharide vs. cerebroside, (3) saturated vs. unsaturated ceramide, (4) glucosyl vs. galactosyl moieties, and (5) GlcCer or GalCer’s carbon chain lengths. Using these 5 characteristics, we establish hierarchical levels within the SERS-based chemical taxonomy machine learning framework. Each level is linked to a molecular structural characteristic, such as the types and numbers of functional groups. Leveraging the taxonomic ML model, we can predict individual structural attributes in a stepwise manner. This progressive process can be done by analyzing and pairwise profiling similarities and differences in structure and SERS spectra. Crucially, this approach facilitates unprecedented forward prediction, allowing for the deduction of “unidentified molecules” situated beyond the boundaries of the ML model. Specifically, our proposed process systematically excludes alternative structural possibilities when the SERS spectra traverse the hierarchical levels of the chemical taxonomy, culminating in the precise identification of the exact molecular structure. In contrast, such forward prediction remains elusive through a single classification ML model, which inaccurately classifies the “unidentified molecules” as one of the pre-existing labeled classes in that model.
This unprecedented forward prediction ability is a notable departure from current ML-detection work as illustrated by the figure below:
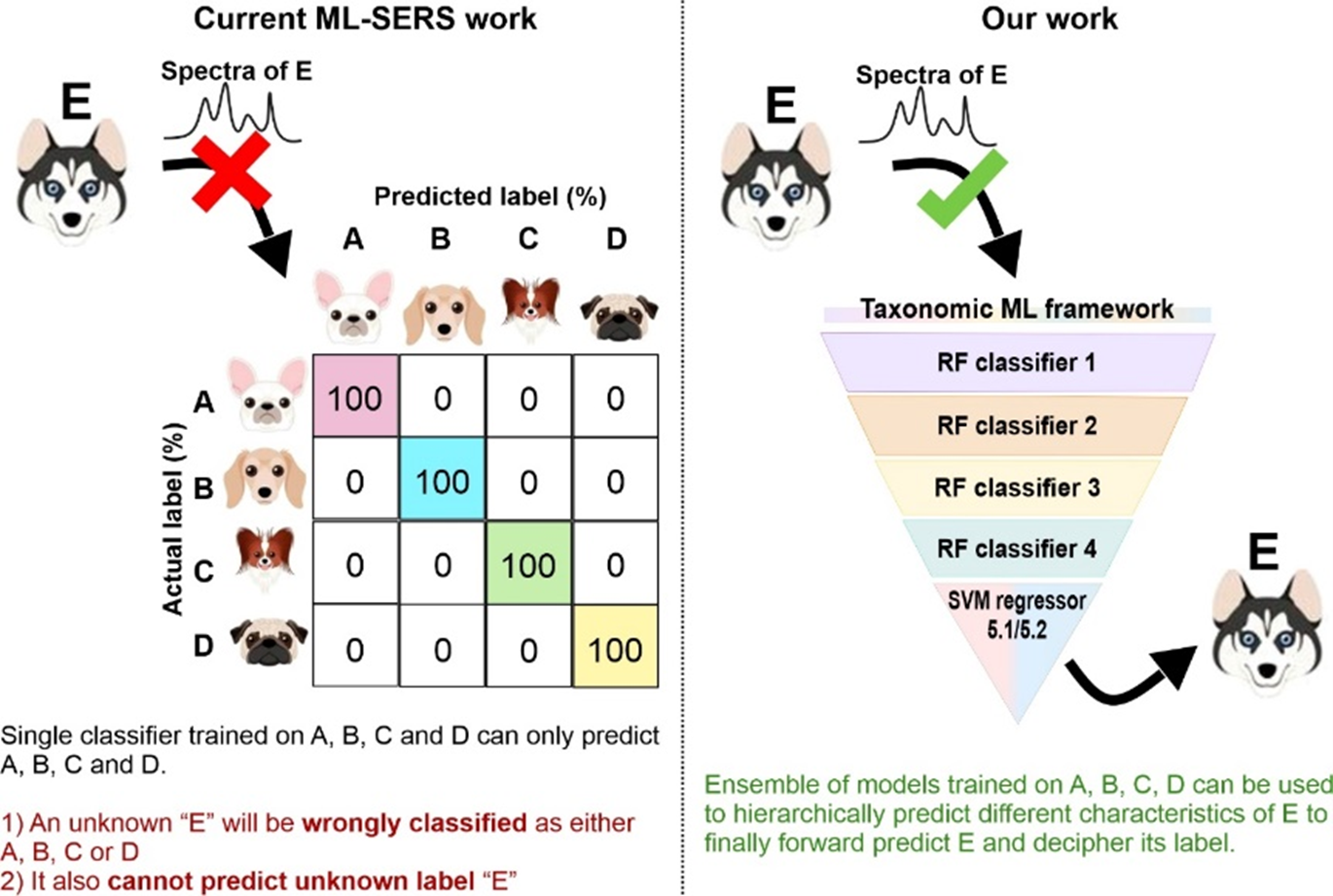
But what if the molecules are present in much lower concentrations? What if we do not know its concentration, can we still identify it? We then performed a series of tests to prove that although the model is established using spectra of cerebrosides at 10−4 M, it is still effective in predicting “unidentified cerebrosides” at concentrations 1–6 orders of magnitude lower than those in the trained model (i.e., at 10−5–10−10 M) with accuracies ranging from 87 to 100% with <1 carbon chain length discrepancy. This demonstrates the robustness and applicability of our chemical taxonomy framework for practical SERS sensing applications, where the concentration of analytes is frequently unknown.
Further integrating with our series of pre-trained quantification models will allow swift quantification following their identification for both qualitative and quantitative analysis of these epimeric biomolecules within a few seconds of inputting their spectra. These models are designed to accurately quantify the concentrations of all 11 pure cerebrosides from 10−4 to 10−10 M. Our models show near-ideal linearity spanning seven orders of magnitude with R2 of 0.95–1.00 and low RMSEprediction of 0.09-0.44 for each epimer, confirming the ultra-trace sensitivity of our SERS platform with a detection limit of 10−10 M. Beyond this study, we posit the creation of a global SERS molecular space using high-throughput platforms to test various probe-analyte combinations. This innovation can synergize effectively with miniaturized SERS spectrometers and microfluidic chips to realize the point-of-need lab-on-a-chip concept by streamlining sample separation and pretreatment to improve SERS detection in complex and heterogeneous mediums.
We hope to continue to borrow new tools from emerging fields like artificial intelligence and machine learning but invigorate them with our ideas about how to utilize them. May we learn but not get bounded by our own knowledge, think outside the box and dream about new ways of applying established methods to achieve things that were thought to be “impossible” before.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in