Gene expression tuning made easy
Published in Bioengineering & Biotechnology
Gene expression tuning: how?
We have recently described a new toolkit for tuning endogenous gene expression using CRISPR: the CasTuner toolkit. CasTuner makes use of either a histone-deacetylase domain (hHDAC4) fused to a catalytically inactive Cas9 (dCas9) or a CasRx (Cas13d) system, which are fused to the FKBP12F36V degron and can thereby be tuned by titrating the quantity of the small molecule degrader dTAG-13. The hHDAC4 domain removes active marks (acetyl groups) from histones, leading to transcriptional repression, while CasRx targets and cuts RNAs, leading to their degradation, in a post-transcriptional fashion.
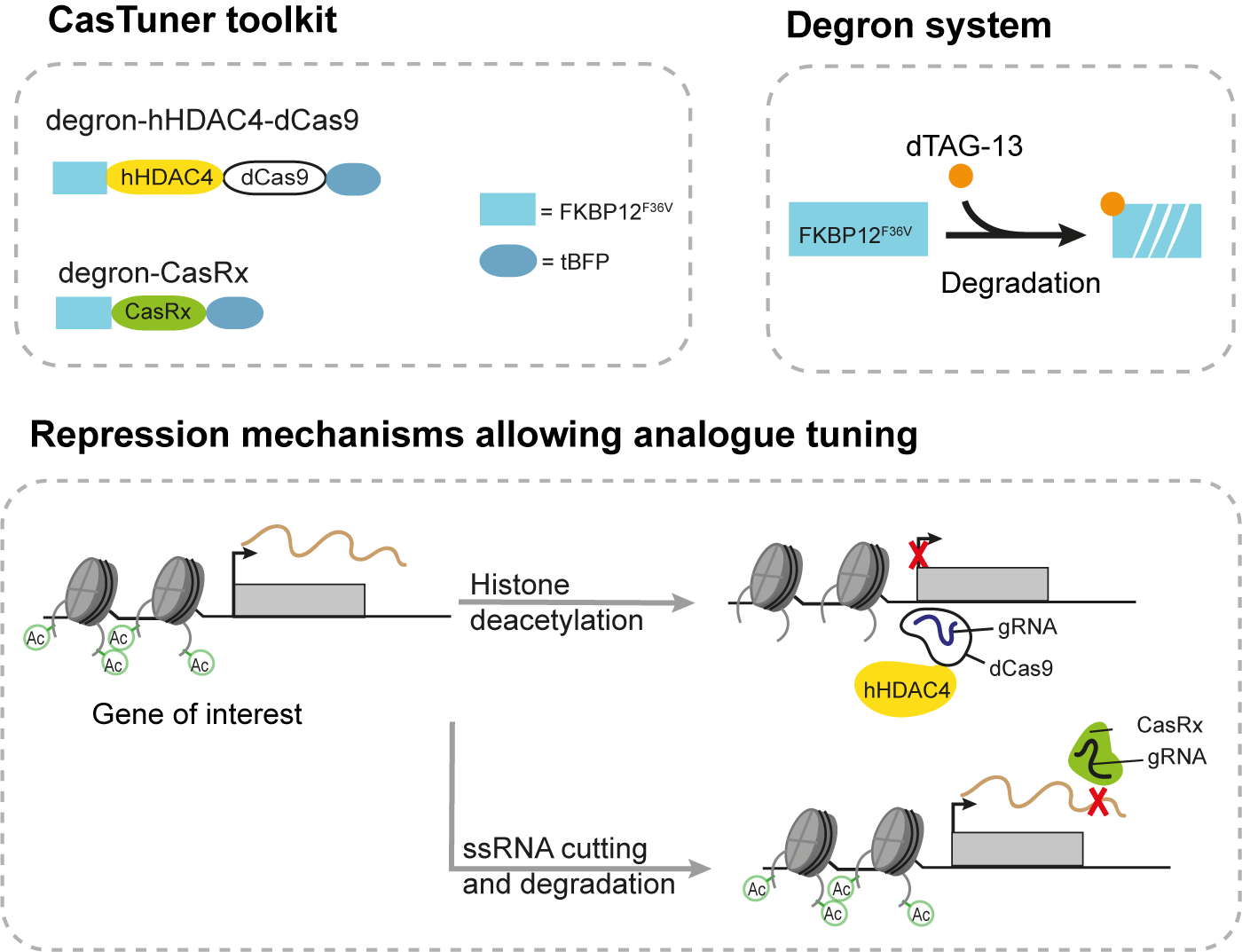
We have shown that these systems are able to induce homogenous intermediate expression levels of endogenous target genes, as opposed to the widely used KRAB-dCas9 system, which can only shut off gene expression. Using a metaphor, if we imagine an expressed gene as a light bulb, CasTuner would represent a dimmer switch, while KRAB-based systems are toggle switches. By extension, we refer to CasTuner as an analogue repressor and to KRAB-based systems as digital repressors.
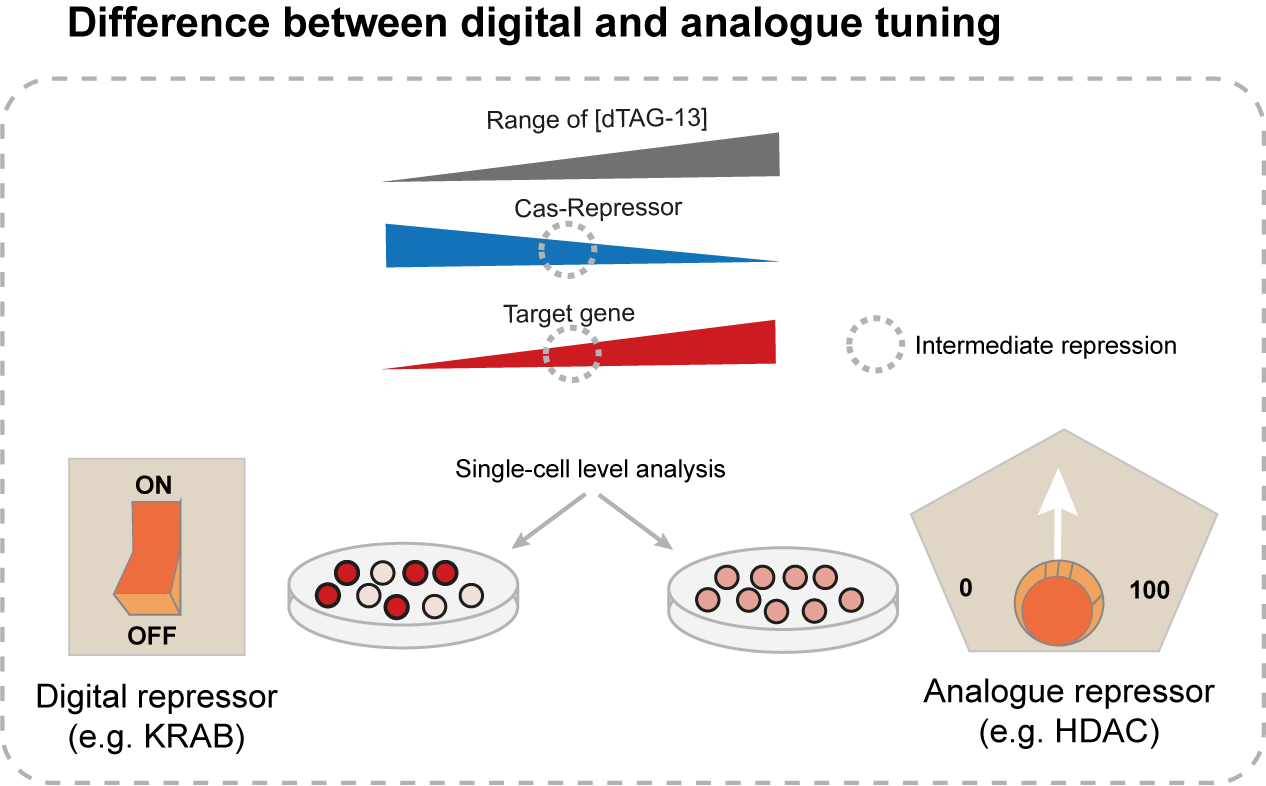
Alright, but how can I actually use CasTuner?
CasTuner systems can be stably expressed in cells using PiggyBac-transposition: the integrating part of the plasmids are flanked by PiggyBac recognition sites and, by co-transfection with a PiggyBac Transposase, will be stably integrated in the genome. An antibiotic selection marker facilitates the enrichment of successful integration events and the blue fluorescent protein (tBFP) directly fused to the repressor systems themselves allows to sort cells with the desired expression levels using flow cytometry.
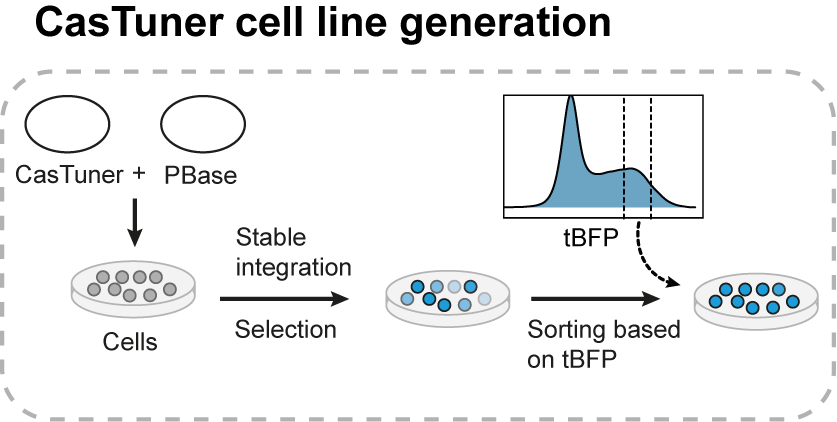
Once a cell line has been obtained, the next step is to choose a gene of interest and design single guide RNAs targeting its promoter (for hHDAC4-dCas9) or the transcribed RNA (for CasRx). Typically we include up to four guides in a lentiviral plasmid that we transduce in the cells. By treating cells with 500nM of dTAG-13 prior to transduction, we make sure to degrade all the repressor and avoid leaky repression prior to the start of the experiment of interest. Our systems are inducible, which can be important both for perturbations of essential genes and for those experiments that require a specific timing: at the onset of the experiment, the ligand can be removed from medium to allow repressor stabilization and consequently target gene repression. In addition to that, perturbations can be reversed by adding the ligand back into the medium. In mouse embryonic stem cells, we also measured the kinetic parameters of repression and derepression for a target gene, for the main systems we tested.
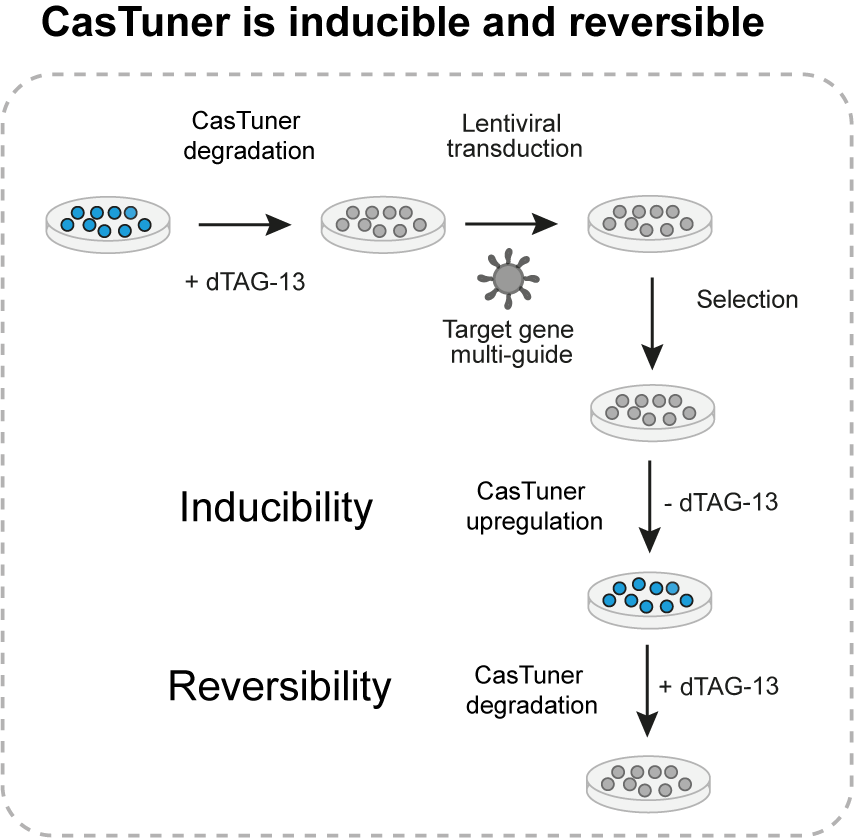
Clearly the main purpose and advantage of CasTuner is that it allows to tune endogenous gene expression. By using sub-saturating concentrations of ligand in the medium, we can obtain intermediate CasTuner levels which in turn repress a target gene only partially. In a typical experiment, we would take cells with the CasTuner system and target guide RNAs stably integrated and seed them in multiple wells, with each well having a different concentration of dTAG-13 within the medium. This way, in each well we can have a different level of target gene expression, ranging from its wild-type level (500nM dTAG-13) to the minimal level achievable (0nM dTAG-13).
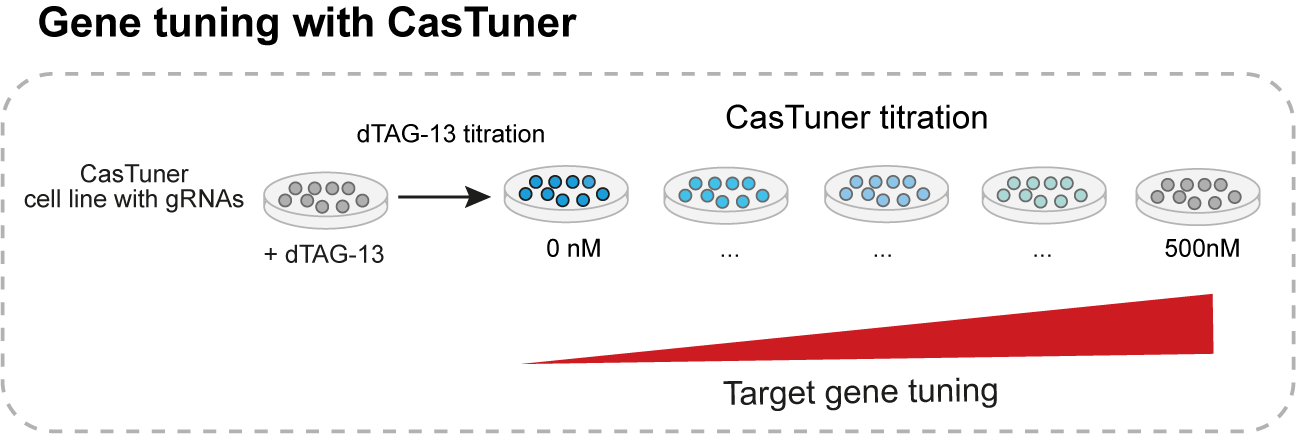
Why do we think it is important to be able to perform quantitative perturbations within physiologically relevant ranges?
Historically, gene functions have been elucidated by analyzing the effects of a genetic alteration. In other words, genes have been broken in order to study them – think about deletions or inactivating mutations. For various reasons, oftentimes it is preferable to leave the genetic sequence of interest untouched and, rather, to alter the level of expression of its product. CRISPRa and CRISPRi systems, for example, are able to up- or down-regulate a gene, respectively. There are a large number of biological processes, however, that rely on specific quantities of proteins or RNAs, which we refer to as “dose-dependent processes”. Such processes include body patterns specification, differentiation of embryonic stem cells into different lineages, random X chromosome inactivation, to name a few. Broadly speaking, we can say that cells decode information on their state in a quantitative fashion. Genes are not simply turned ON or OFF at a given moment, but their level of expression, their quantities, are finely modulated in order to confer specific identities.
Sometimes, the same protein might have different roles depending on its dose. For example, when the pluripotency factor OCT4 drops below a certain threshold, mouse embryonic stem cells were observed to dedifferentiate into the trophectodermal lineage, while an OCT4 over-dose drives the primitive endoderm lineage specification (Ref. 1). Considering that both trophectoderm and primitive endoderm are extra-embryonic lineages, these observations should make the point of how crucial the quantitative modulation of gene expression is and how necessary it is for researchers to understand it. Ironically, because tuning gene expression has been rather non-trivial in the past, it is hard to estimate which processes are dose-dependent and what are the molecular mechanisms underlining this fascinating property. In this sense, I am confident that our new CasTuner toolkit, which we are glad to share with the scientific community, will enable faster discovery and characterization of dose-dependent biological processes.
After geneticist have been breaking genes for decades, it is now time for us to tune them.
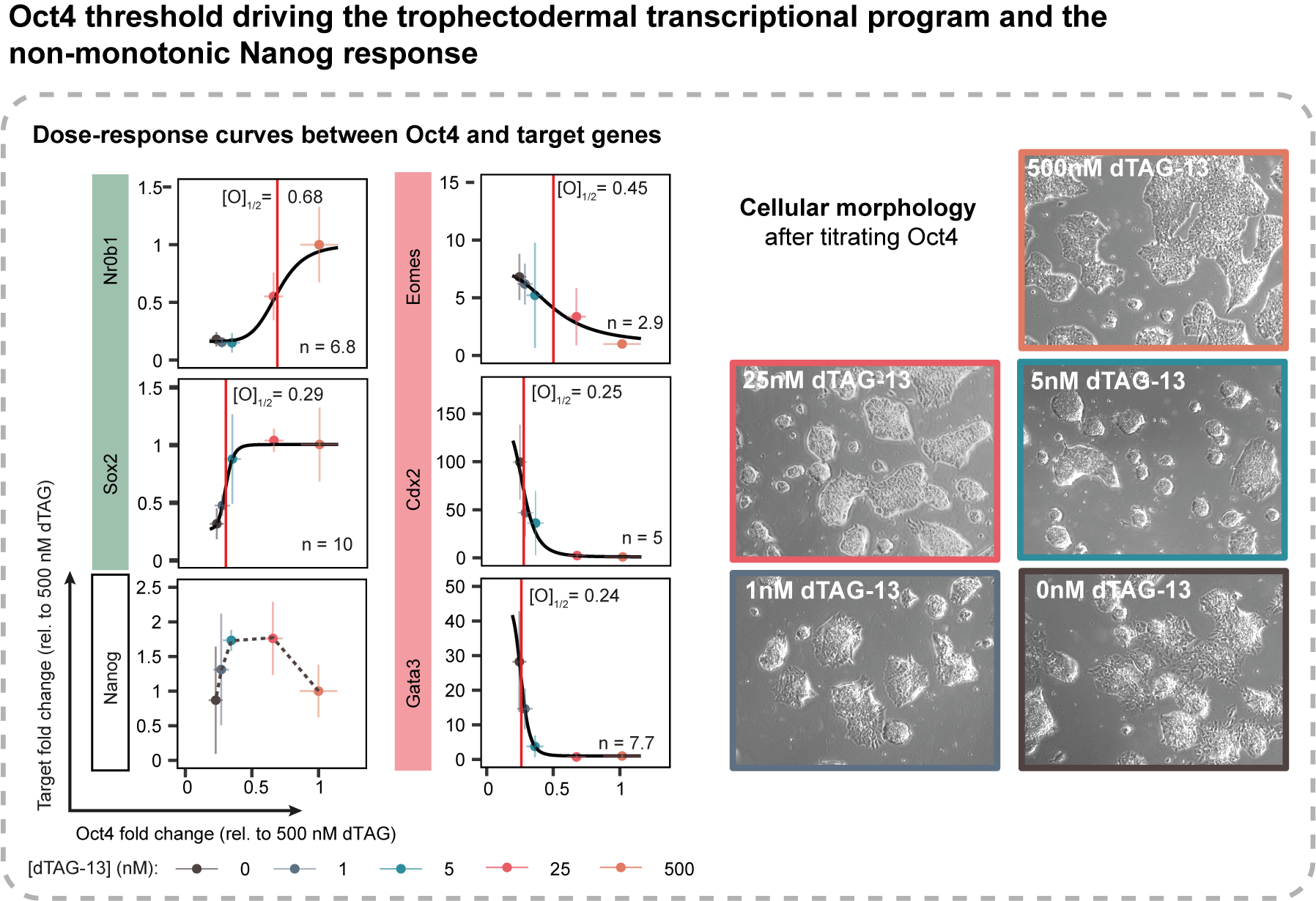
Clearly, there are a few other ways to achieve gene expression tuning. For instance, in a recent work, Naqvi and colleagues have altered the dosage of the transcription factor SOX9 via direct degron fusion (Ref. 2). Altering protein stability in such a way has the advantage of a more rapid control over protein turn-over, as it acts directly at the post-translational level, rather than at the transcriptional or post-transcriptional level as it is the case for CasTuner. However, not only the generation of knock-in cell lines is time-consuming, it can also present other major draw-backs, depending on the specific application. Degron domains can for example cause an alteration of protein stability even in the uninduced condition (degradation leakiness), or not fully destabilize the protein of interest when induced (low degradation efficiency). Such aspects can however be tested prior to generating the knock-in line. The major problem that would still remain concerns the fusion of a domain to protein in itself. If the object of study is a protein forming multiple interactions, the fusion domain could interfere with them. Since in recent years protein concentration-dependent phenomena (see liquid-liquid phase separation/condensate formation) have been described playing a pervasive role in biological processes (Ref. 3), probing their physical-chemical properties in vivo, in physiologic conditions and without additions to the protein sequence can help to clarify the controversial aspects in the field.
References
-
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
- Naqvi, S., Kim, S., Hoskens, H. et al. Precise modulation of transcription factor levels identifies features underlying dosage sensitivity. Nat. Genet. 55, 841–851 (2023).
- Boeynaems S, Alberti S, Fawzi NL, et al. Protein Phase Separation: A New Phase in Cell Biology. Trends Cell Biol. 28(6), 420-435 (2018).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in