Genetic function without charge
Published in Chemistry
How chemistry acquired a capacity for information storage and propagation is a question that has always fascinated me, because any answer to this question has immediate implications for our understanding of how life emerged on Earth (or elsewhere), a central unresolved question of science. But what are the chemical parameters required for an easily writable and readable molecular memory? Nature provides two molecular solutions, DNA and RNA and our enquiry must start from there.
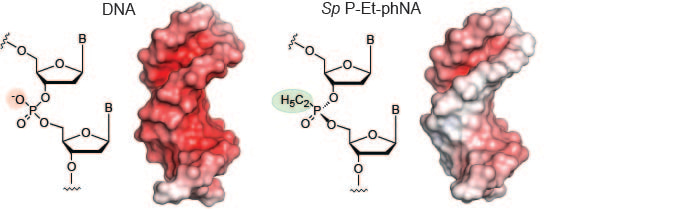
Starting with the insights of Westheimer that phosphate-esters occupy a special place of kinetic stability due to charge shielding of the phosphorus nucleus, the phosphodiester backbone of DNA and RNA is also special: it not just provides a kinetically stable scaffold but, in addition, its polyanionic nature is so dominant that it effectively decouples base sequence (i.e. information content) from the physiochemical bulk properties of the molecule. Recognising the fundamental importance of this for any form of molecular memory, Benner and Huttner proposed a “polyelectrolyte theory of the gene” stating that genetic molecules may be recognized by their highly charged nature (for example in the search for extraterrestrial life).
An experimental test of this conjecture interested us for both scientific and pragmatic reasons. Nucleic acids are not just molecules for information storage, but potentially also a source of powerful drugs. However, a polyanionic configuration poses many hurdles in drug development including poor delivery across cellular membranes and rapid renal clearance. This has spurred the development of nucleic acid analogues with uncharged backbones. In agreement with theory, many of these seem incompatible with information storage in that they preclude specific base-pairing with DNA (or among themselves). Notable exceptions are peptide nucleic acids (PNAs) developed by Nielsen and phosphorodiamidate morpholino oligomers (PMO) developed by Summerton, which form highly specific hybridization probes and anti-sense oligonucleotide (AOS) drugs like the FDA-approved PMO drug Eteplirsen. However, although specific hybridization may be a precondition for information transfer, it is in itself not sufficient but requires a means of replication.
In this paper, we chose to examine another uncharged analogue, alkyl-phosphonate nucleic acids (phNAs). phNAs are attractive analogues as they are nearly isosteric to DNA. They also had been explored in the 1980s as 1st generation AOS reagents, and found to be both biostable and non-toxic. Using a combination of in silico modelling and in vitro evolution we were able to discover an efficient DNA-templated phNA polymerase and show genetic information transfer from DNA into phNA and back into DNA as well as evolution of specific phNA aptamer ligands to a protein target (Streptavidin) directly from random phNA repertoires. This demonstrates that genetic information storage, propagation and evolution are compatible with an uncharged backbone chemistry (at least in combination with a DNA intermediate) indicating that the postulated stringent requirement for a polyelectrolyte backbone for all genetic materials may be an oversimplification. This also opens up the potentially rich properties of uncharged genetic polymers and their uncharted phenotypic space for exploration.
You can read more about the study in Nature Chemistry.
Find us at the MRC Laboratory of Molecular Biology Holliger lab and on twitter @PhilHolliger
Follow the Topic
-
Nature Chemistry
A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in