Harnessing extreme underdominance to engineer a synthetic species
Published in Bioengineering & Biotechnology
“It is not always the magnitude of the differences observed between species that must determine specific distinctions, but the constant preservation of those differences in reproduction.” — Jean-Baptiste Lamarck
In nature, speciation occurs when a barrier to gene flow is introduced into a population, causing the two divided parts of the population to evolve separately, with each developing their own genetic differences and eventually becoming genetically independent species. Specifically, these genetic differences can create deviations in reproduction, with pre- and post-zygotic barriers eventually arising to completely prevent the production of viable offspring. Years ago, while Drs. Anna Buchman and Omar Akbari were working together in the lab of Dr. Bruce Hay at Caltech, they were fascinated by the idea of building an artificial species using extreme underdominance (a system where the heterozygous genotype is inferior to both the recessive and dominant homozygous genotypes). This seemed like the ultimate bioengineering feat - building a synthetic species, one that is completely reproductively isolated from its natural counterpart. But how do you engineer a synthetic speciation event?
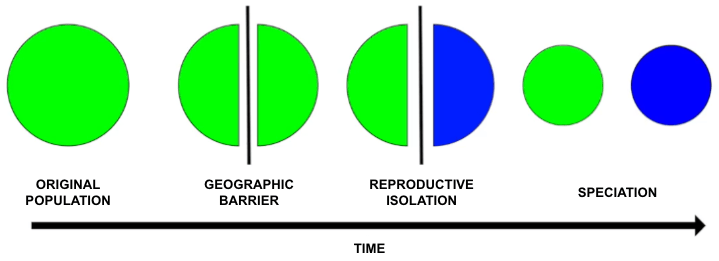
A few years later, while at UC San Diego, Anna and Omar set out to achieve this natural phenomenon by engineering synthetic reproductive isolation using CRISPR/Cas9. In practice, this synthetic speciation tool could be advantageous for preventing hybrid zygotes, for ecosystem engineering, or for developing - and controlling - gene drives.
Gene drives offer great promise to genetically control vectors capable of transmitting disease, or pests that can devastate crops, and thus reduce their impacts on human health and agriculture. Engineered gene drives work by biasing the inheritance of a heritable gene into a population using recombinant DNA techniques, effectively increasing the frequency of the gene in the population 1. These drives can be implemented as suppression or replacement drives. Suppression drives often target a gene of interest that induces a fitness or reproductive cost to progeny homozygous for the altered gene, which suppresses and decreases the entire population over time. In the case of vector control, this acts to decrease the number of disease-carrying vectors in an area, which effectively decreases the incidence of disease. Replacement drives operate in a similar manner, except that they aim to replace a wild-type population with an engineered population possessing a beneficial trait, such as increased sensitivity to insecticides. However, a significant challenge for gene drives is preventing them from spreading uncontrollably. There have been a number of promising methods to control gene drives via synthetic reproductive isolation 2–5, but these systems had limitations for acting as complete gene drives, as they were either not feasible in multicellular organisms 6–9, used complex expression systems 10, or did not achieve complete reproductive isolation 2,5,11. Anna and Omar began to brainstorm ways to overcome these hurdles and find tools to engineer synthetic speciation. They eventually determined that we could harness extreme underdominance using CRISPR-based genome editing and transcriptional transactivation (CRISPRa), which was previously used to build reproductive barriers in yeast 12, to build such a system in insects.
Shortly after this project began, I joined the Akbari Lab as a Research Assistant. I was fortunate to work closely with Anna, who explained the goal of engineering synthetic speciation and how it could ultimately be done. Maselko et al. 12 and Waters et al. 13 had demonstrated high-quality proof of concept synthetic speciation systems in yeast and flies respectively, but these systems were either limited to unicellular organisms or did not achieve complete reproductive isolation. So while the previous work provided a foundation for building synthetic speciation, we set out to develop a system in a complex organism that was 100% reproductively isolated.
We began testing sgRNAs that targeted the promoters of genes essential for development or viability. The goal was to overexpress these genes in the post-zygotic stage, inducing lethality before a viable offspring could form. We ultimately used four essential genes - eve, hid, hh, and wg - and began testing them using engineered flies that expressed two different dCas9 activator domain fusions (dCas9-VP64 and dCas9-VPR) under the control of either a ubiquitous (Ubiquitin-63E) or early embryonic (bottleneck, bnk) promoter. We found that dCas9-VP64 didn’t overexpress target genes enough, while ubiquitous dCas9-VPR expression proved to be too toxic. So we ultimately used bnk-driven dCas9-VPR, which gave 100% lethality when paired with our sgRNAs with no noticeable toxicity. In order to “rescue” individuals from this lethality, we used maternally-deposited spCas9 to generate site-directed protective indels in the sgRNA target site, which prevents dCas9/sgRNA from binding and inducing lethal overexpression. Using spCas9, dCas9-VPR, and sgRNAs targeting genes essential for development, we engineered eight SPECIES (Synthetic Postzygotic barriers Exploiting CRISPR-based Incompatibilities for Engineering Species), which are reproductively isolated from each other and wild-type, due to lethal overexpression induced by dCas9-VPR. We initially attempted to target single genes; however, we found that despite harboring the required CRISPR machinery and protective indels, outcrosses to wild-type resulted in viable offspring, as one protected target site was sufficient to rescue offspring individuals from lethal overexpression, which was similar to the results demonstrated in Waters et al. 13. So we instead tested multiplexed overexpression, targeting two or more genes per SPECIES, which ultimately proved to successfully induce lethal overexpression in offspring individuals. After creating a homozygous stock of our rescued SPECIES individuals and demonstrating that each line was reproductively isolated from wild-type, we concluded that we had accomplished the ultimate bioengineering feat - building a synthetic species!
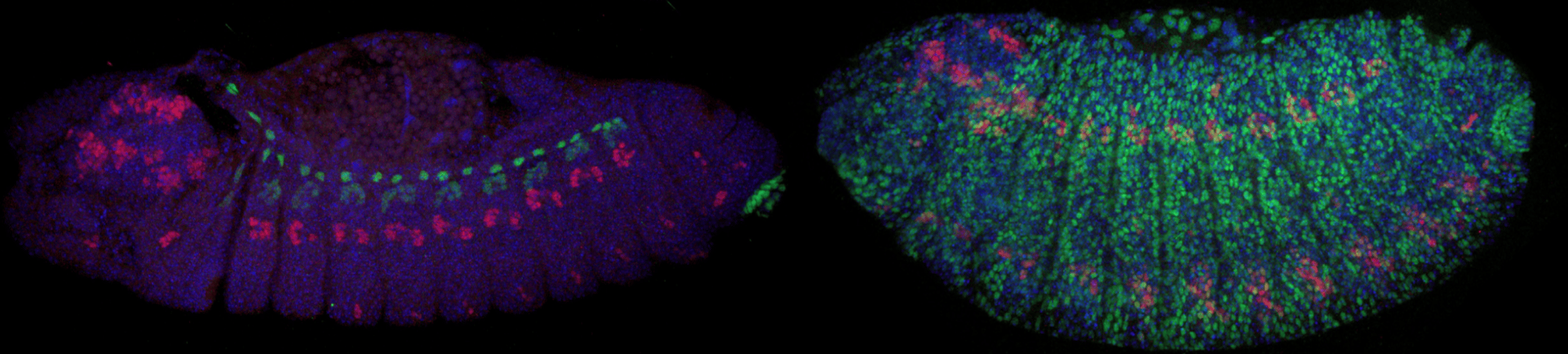
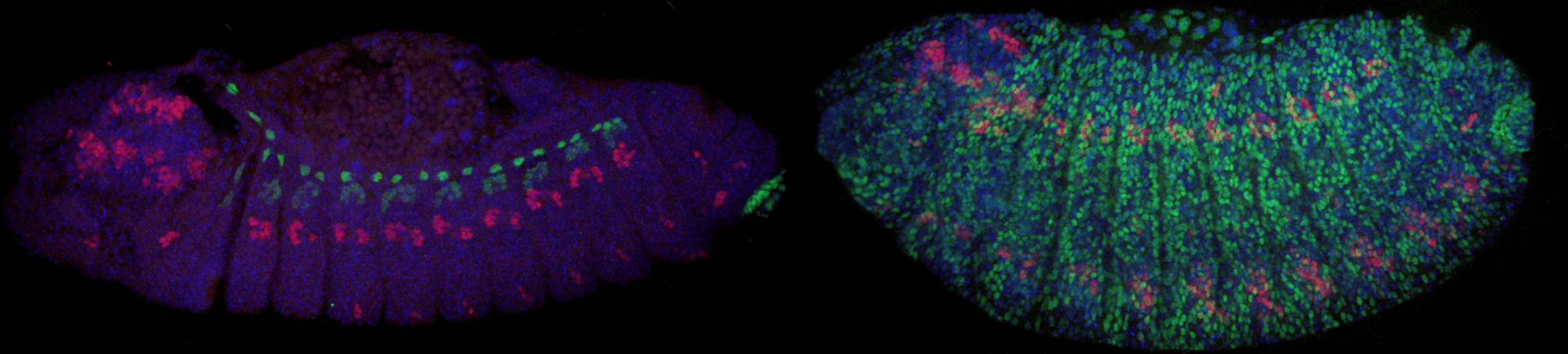
Antibody stains by co-author Michael Perry. Left is a SPECIES selfcross with normal eve expression. Right is a SPECIES outcross with lethal eve overexpression.
Next came the bulk of the experiments - proving that these individuals were 100% reproductively isolated from as many wild type flies as possible. Countless grape-agar plates were made, hundreds of crosses were set-up, and thousands of embryos were counted and observed, and to our excitement, no viable adults were found! We also wanted to demonstrate that the extreme underdominance system in our SPECIES could function as a gene drive in wild populations, which could prove useful in vector and pest control. In theory, SPECIES could act to replace a WT population with a desirable one (i.e. one resistant to disease or sensitive to insecticides). So we set up population studies, crossing our SPECIES individuals to WT at various thresholds and observing them over generations. After many replicates and months of counting, we determined that our SPECIES could effectively drive into a population at various release percentages, eventually resulting in WT population replacement. We also determined that our drive was threshold dependent, meaning that at certain release percentages, we had the control to either replace the WT population with SPECIES, or effectively reverse the drive and replace it with a WT population again. This was further confirmed by mathematical models created by our co-author John Marshall at UC Berkeley, which also demonstrated that SPECIES are more confinable than other gene drives due to their inherent threshold dependence and ability to modify localized populations (see Figs. 4-6 in our main text).
These results are exciting, as they demonstrate that synthetic species that are reproductively isolated can be engineered using extreme underdominance thanks to CRISPR. Our SPECIES system is threshold dependent and could, in theory, be paired with a useful cargo gene, such as disease resistance, and drive into populations to achieve a desired result (i.e. reducing disease). There have been other recent advances in this branch of bioengineering as well, as last year Maselko et al. built on their previous work and demonstrated synthetic speciation in flies, a system they describe as Engineered Genetic Incompatibility (EGI) 14. Their results, as well as ours, reveal the possibility of applying these systems to animals in which it will be particularly useful, such as mosquitoes or crop pests, which could have profound impacts on the fields of public health and agriculture.
Our full publication can be accessed via the link here: https://www.nature.com/articles/s41467-021-23531-z
Link to video created by co-author Junru Liu summarizing the engineering of SPECIES: https://www.youtube.com/watch?v=380IWQabI1U
References:
-
Alphey, L. S., Crisanti, A., Randazzo, F. F. & Akbari, O. S. Opinion: Standardizing the definition of gene drive. Proc. Natl. Acad. Sci. U. S. A. 117, 30864–30867 (2020).
-
Akbari, O. S. et al. A synthetic gene drive system for local, reversible modification and suppression of insect populations. Curr. Biol. 23, 671–677 (2013).
-
Altrock, P. M., Traulsen, A., Reeves, R. G. & Reed, F. A. Using underdominance to bi-stably transform local populations. J. Theor. Biol. 267, 62–75 (2010).
-
Edgington, M. P. & Alphey, L. S. Conditions for success of engineered underdominance gene drive systems. J. Theor. Biol. 430, 128–140 (2017).
-
Buchman, A. B., Ivy, T., Marshall, J. M., Akbari, O. S. & Hay, B. A. Engineered Reciprocal Chromosome Translocations Drive High Threshold, Reversible Population Replacement in Drosophila. ACS Synth. Biol. 7, 1359–1370 (2018).
-
Lajoie, M. J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
-
Rovner, A. J. et al. Recoded organisms engineered to depend on synthetic amino acids. Nature 518, 89–93 (2015).
-
Boeke, J. D. et al. GENOME ENGINEERING. The Genome Project-Write. Science 353, 126–127 (2016).
-
Foo, J. L. & Chang, M. W. Synthetic yeast genome reveals its versatility. Nature vol. 557 647–648 (2018).
-
Moreno, E. Design and Construction of ‘Synthetic Species’. PLoS ONE vol. 7 e39054 (2012).
-
Reeves, R. G., Bryk, J., Altrock, P. M., Denton, J. A. & Reed, F. A. First steps towards underdominant genetic transformation of insect populations. PLoS One 9, e97557 (2014).
-
Maselko, M., Heinsch, S. C., Chacón, J. M., Harcombe, W. R. & Smanski, M. J. Engineering species-like barriers to sexual reproduction. Nat. Commun. 8, 883 (2017).
-
Waters, A. J., Capriotti, P., Gaboriau, D. C. A., Papathanos, P. A. & Windbichler, N. Rationally-engineered reproductive barriers using CRISPR & CRISPRa: an evaluation of the synthetic species concept in Drosophila melanogaster. Sci. Rep. 8, 13125 (2018).
-
Maselko, M. et al. Engineering multiple species-like genetic incompatibilities in insects. Nat. Commun. 11, 4468 (2020).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in