Identifying signaling molecules from Gram-positive bacteria
Published in Chemistry

The expression of virulence factors in Gram-positive bacteria, including staphylococci, is regulated through a process called quorum sensing (QS). This cell-to-cell communication is mediated by the secretion and detection of molecules termed auto-inducing peptides (AIPs). AIPs from non-aureus staphylococci have received attention as potential anti-virulence agents to inhibit QS and virulence gene expression in the human pathogen, Staphylococcus aureus. In an effort to investigate inter-species QS modulation, our colleagues at the Department of Veterinary and Animal Sciences observed strong inhibition of S. aureus QS by the supernatant from a culture of the dog pathogen S. schleiferi. Based on the genome sequence of the bacterium, an AIP was predicted and we (Olsen Lab) were asked if we could synthesize this molecule to substantiate the claim that an AIP from the dog pathogen was responsible for the observed QS inhibition. The synthesized thiolactone-containing octapeptide indeed exhibited strong inhibition of QS in S. aureus. However, it is not possible to predict the full structure of an AIP based on genetic information alone, because the cleavage site of the so-called leader peptide during processing of the natural AIP in the bacterium varies between species and specificity groups.
We were therefore keen to come up with a method for identifying the AIP directly from spent medium, but ideally without performing lengthy activity-guided fractionations and HPLC purifications as in traditional natural products chemistry. After failing at detection of the AIP by performing simple extraction procedures, it was envisioned that some sort of enrichment process by binding of the molecule to a solid support might do the trick. This was when the graduate student working on the project, Bengt Gless, thought of native chemical ligation (NCL), which is a highly chemo-selective reaction between an N-terminal cysteine and a thioester (in this case the thiolactone). A simple polymeric resin, functionalized with cysteine, proved highly efficient for trapping synthetic AIP molecules from medium in preliminary experiments, and we were excited to find that expressed AIPs from cultured bacteria could also be enriched and identified by standard LC-MS analysis after cleavage from the resin.
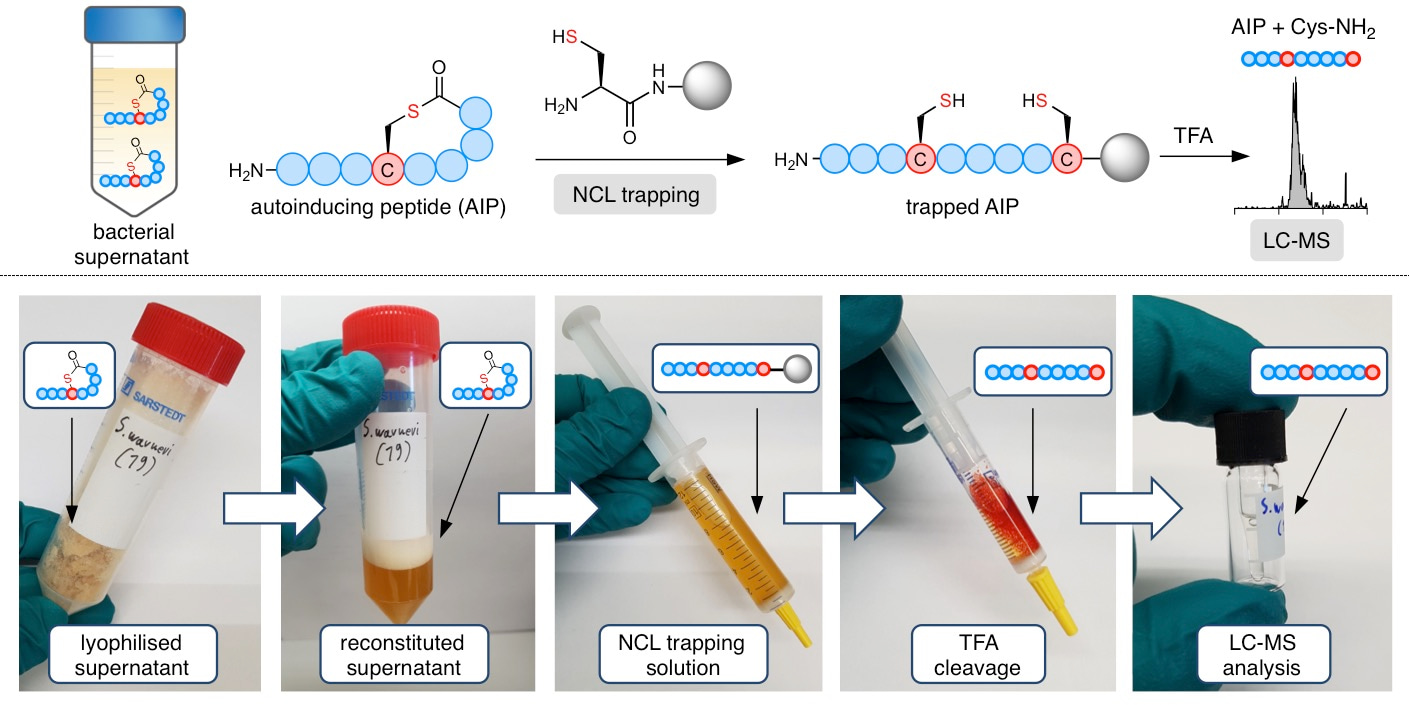
Ultimately, the S. schleiferi AIP proved to be a nonamer rather than the initially predicted octamer. Further application of our enrichment protocol, followed by standard LC-MS analysis guided by genome sequencing data, allowed us to confirm the identity of 5 known AIPs and to identify the AIPs of 11 species that were previously unknown. This effort doubles the number of known staphylococcal AIPs and we expect that the method should be extendable to AIP-expressing Gram-positive bacteria beyond the staphylococcal genus.
We hope that access to new AIPs will pave the way to detailed studies of QS and its impact on important pathogen properties such as biofilm formation, colonization, virulence, and interactions with commensal staphylococci.
You can read more about the study in Nature Chemistry.
This post was co-authored by Bengt H. Gless and Christian A. Olsen.
Find us on twitter @BengtGless and @ChristianAOlsen
Follow the Topic
-
Nature Chemistry
A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in