Image Analysis Reveals Molecularly Distinct Patterns of TILs in NSCLC associated with Treatment Outcome
Published in Cancer
Non-small cell lung cancer (NSCLC) is the most common type of lung cancer and has a relatively low 5-year survival rate of less than 21%. Recently, a number of studies have found that tumor-infiltrating lymphocytes (TILs), cells that invade into the tumor tissue and kill tumor cells, are associated with clinical outcomes and response to chemotherapy and immunotherapy in NSCLC patients across different stages. A few studies [1, 2, 3, 4] have shown that computer-derived quantitative patterns related to TIL density and spatial arrangement observed on digitized tissue slides (H&E images) are associated with disease prognosis. However, it is unclear whether the spatial morphologic patterns of TILs are equally prognostic in LUAD and LUSC, and it is also unclear whether the TIL signatures that are prognostic in LUAD and LUSC have similar molecular composition in terms of the relative proportion of TIL subtypes. For example, Corredor et al. [1] previously found that the spatial arrangement of TILs related to the colonization TIL clusters and cancer nuclei clusters within the tumor area was associated with the likelihood of recurrence in early-stage NSCLC but did not consider potential differences between LUAD and LUSC.
Using over 1,000 LUAD and LUSC tumors across 6 different datasets, we evaluated the association of TIL density and spatial arrangement related features with outcome in LUAD and LUSC independently. We identified that computationally derived patterns of TILs on H&E images were different between LUAD and LUSC, with TIL density being prognostic of overall survival in LUAD and spatial arrangement being more prognostically relevant in LUSC. The overall workflow includes automated nuclei detection using watershed-based algorithm, which uses mathematical operations to segment out the nuclei boundaries. This was followed by a step to distinguish lymphocytes from cancer nuclei using features related to the shape, size, and texture of individual cells in conjunction with a machine learning classifier. Targeted tile selection was then performed to select the most representative tiles determined by a dimensionality reduction-based algorithm from each patient. From those selected tiles, TIL density (e.g. ratio between number of TILs and tissue area) and spatial arrangement features (e.g. areas of TIL clusters and intermixing of TIL and cancer nuclei clusters) were extracted. To investigate the association between TIL signatures and clinical outcome, we used Cox regression models which were trained and internally validated on 859 patients from the public dataset Cancer Genome Atlas of National Cancer Institute (TCGA) that consists of mostly early-stage LUAD and LUSC. We validated the models’ ability to prognosticate outcome externally on 100 advanced LUAD patients from University of Bern in Switzerland (UBern) [5]. A subset of those UBern patients were treated with more than 6 different regimens of neoadjuvant chemotherapy. We also used LUAD-specific signature to predict response to immunotherapy in 303 advanced NSCLC patients from a clinical trial study CA209-057 (CheckMate057) [6].
We found that in LUAD, the TIL count/density is much higher in model-predicted high-risk patients as compared to model-predicted low-risk patients. Further, the TIL density signatures were also prognostic of outcome in the UBern dataset. This indicates that the computationally derived signature is robust in capturing biological hallmarks of disease aggressiveness, independent of treatment and tissue sample type. In addition, the TIL density signature was also associated with objective response to immunotherapy in 303 advanced NSCLC patients from the completed CheckMate057 clinical trial [6]. This suggests that the TIL density model potentially captures signatures related to disease aggressiveness in prediction of both clinical outcome and treatment response. In LUSC, the spatial distance between TILs in tumor regions was larger in patients with worse survival outcomes. Interestingly, we found that LUSC patients actually had a significantly higher TIL density as compared to LUAD patients. However, even though the TIL density is high in LUSC patients, it is not as strongly associated with clinical outcome as compared to the spatial distribution of TILs. These results suggest the need for future TIL-based models to stratify survival risk and predict response to therapy independently in different histologic subtypes of NSCLC.
To access whether the morphologic differences in TIL density and arrangement were also reflected in differences in molecular composition of TILs, we further investigated the molecular composition of the TIL signatures. We used 83 NSCLC patients from Yale Pathology [7] who had multiplexing TIL immunofluorescence stained images which capture the major subtypes of TIL, namely CD4+ T, CD8+ T, and CD20+ B cells. We found that in LUAD, the prognostic TIL signature was primarily comprised of CD4+ T and CD8+ T cells, whereas in LUSC, the immune patterns were comprised of CD4+ T, CD8+ T, and CD20+ B cells. Specifically, density measures of CD4+ T cells and of CD8+ T cells were both independently found to be prognostic of outcome in LUAD patients. The spatial interaction between CD4+ T and CD8+ T cells and the interaction between CD4+ T and CD20+ B cells were associated with outcome in LUSC patients. Spatial interaction was defined as increasing overlapping area between two TIL subtypes, and it was associated with better survival. These results suggest that not only the density of TIL subtypes, but also their spatial co-localization might provide valuable insight for cancer prognosis. Moreover, in both subtypes, prognostic TIL features were associated with 1) transcriptomics-derived immune scores, and 2) biological pathways implicated in immune recognition, response, and evasion. The Immune scores were generated using immune and normal cell related gene expression data obtained by RNA-sequencing from formalin fixed paraffin-embedded tumor samples, and these scores were used to quantify the presence of immune cells in tumor tissues. By using routinely acquired diagnostic H&E slides instead of immune scores, the approach that involves computationally-derived TIL features could offer a lower cost approach for outcome prediction compared to more expensive gene expression-based assays [8], which are usually tissue-destructive.
In summary, we identified unique spatial morphologic TIL signatures that were independently prognostic in LUAD and LUSC based on H&E images, with TIL density measures being prognostic in LUAD and spatial arrangement of TILs being prognostic in LUSC. Further, the LUAD specific TIL signature was associated with outcome in an external validation set of 100 NSCLC patients treated with more than six different neoadjuvant chemotherapy regimens, and predictive of response to immunotherapy in the clinical trial CA209-057.We further showed that the immune composition of the morphologically distinct TIL signatures in LUAD and LUSC was different. Specifically, CD4+ T and CD8+ T cells dominate the prognostic signals that capture TIL density measures in LUAD and CD4+ T, CD8+ T, and CD20+ B cells dominate the prognostic signals related to spatial distribution of TILs in LUSC. In both LUAD and LUSC, we discovered associations between prognostic TIL features and transcriptomics-derived ISs. Biological pathways implicated in immune recognition, response, and evasion were significantly differentially expressed with respect to the prognostic features in both LUAD and LUSC.
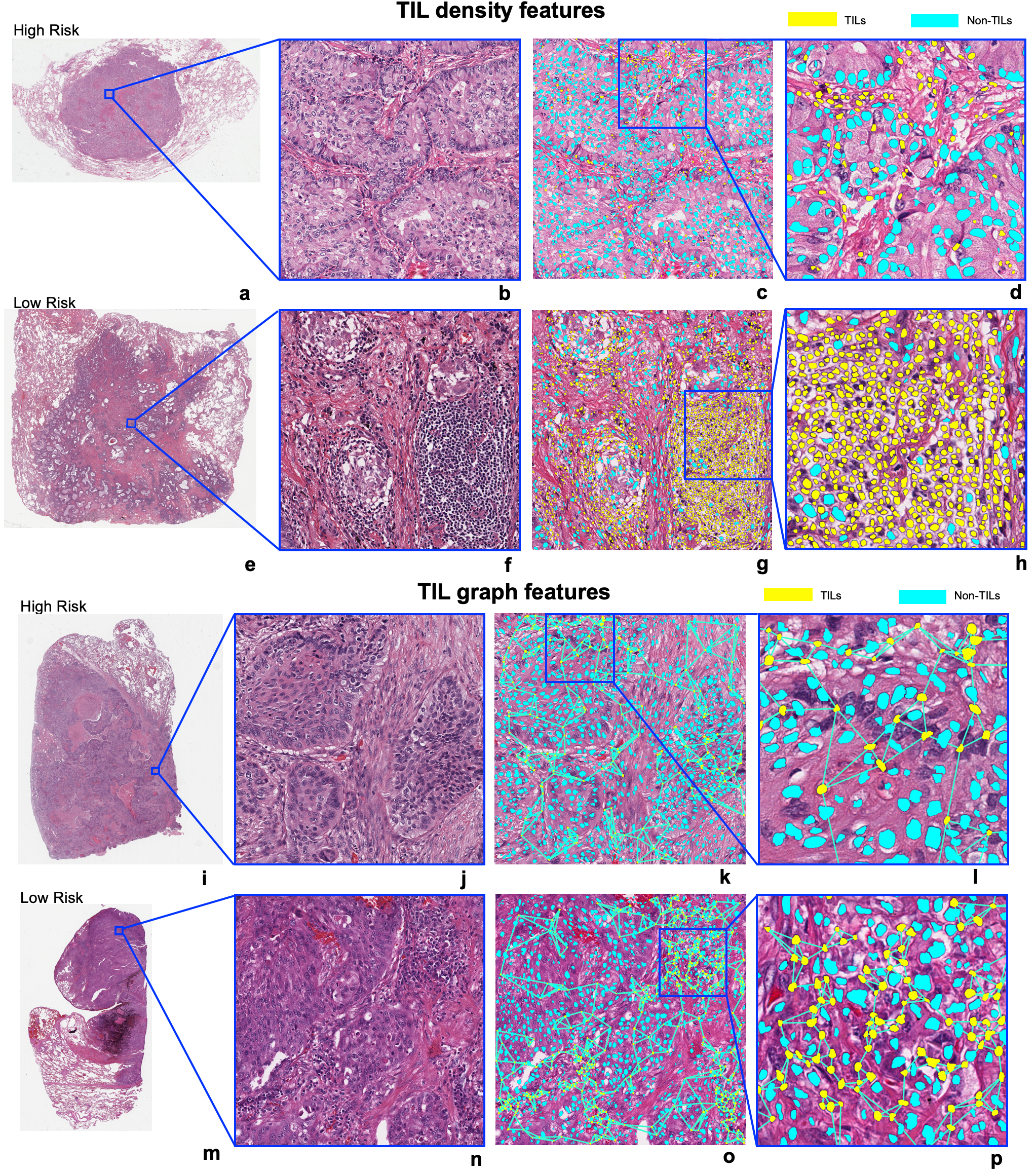
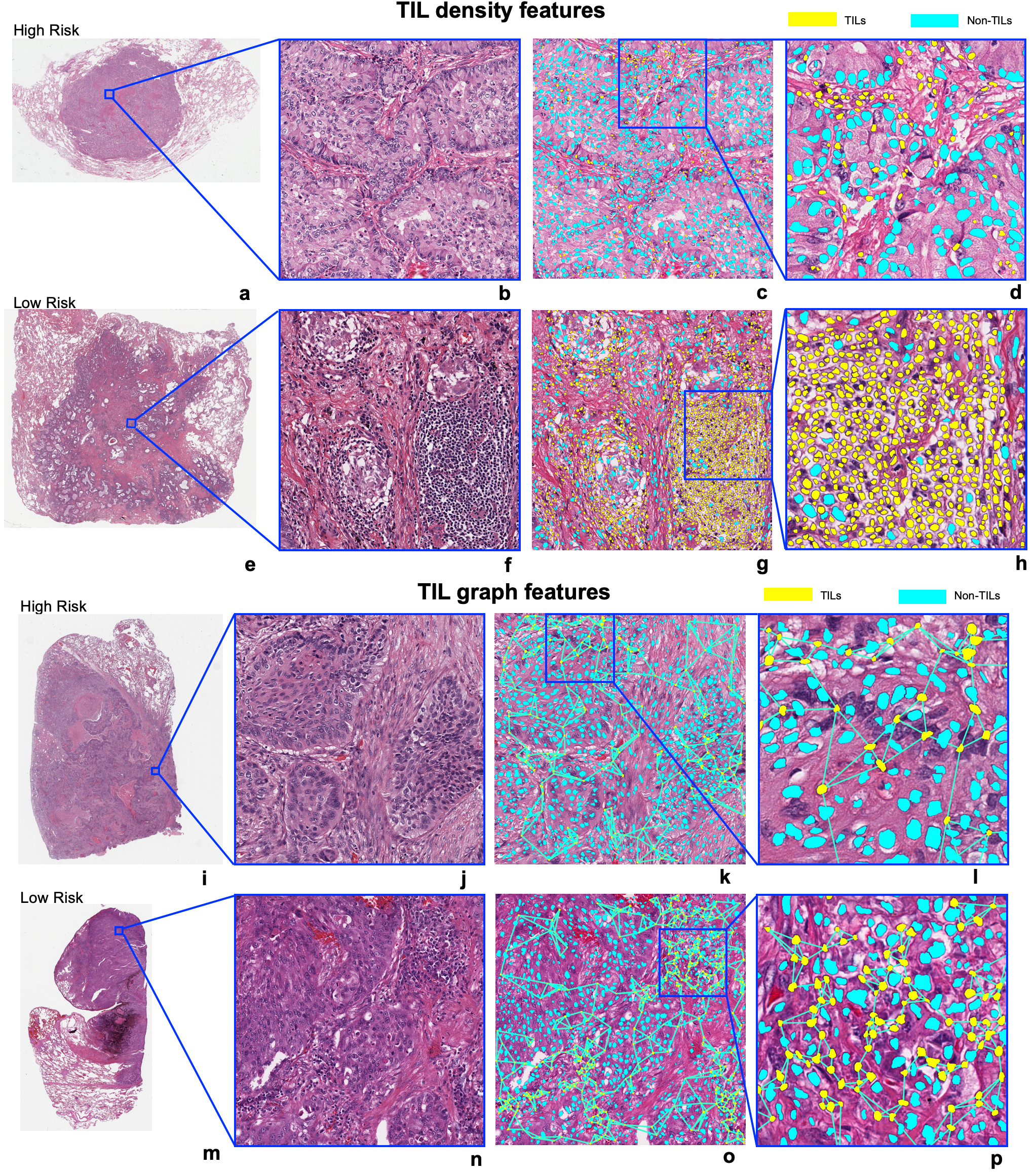
Figure: Visualization of prognostic TIL features in LUAD and LUSC. (a, b, c, d, e, f, g, h) represent the TIL density measures in LUAD. TIL density in the low-risk patient is prominently higher than in the high-risk patient. (i, j, k, l, m, n, o, p) represent the visualization of Euclidean distance to three nearest neighbors of each TIL in LUSC and the distance was measured from the centroid of each TIL. The TIL distribution is sparser in a high-risk patient as compared to a low-risk patient.
References
[1] Corredor, G. et al. Spatial architecture and arrangement of tumor-infiltrating lymphocytes for predicting likelihood of recurrence in early-stage non–small cell lung cancer. Clin. Cancer Res. 25, 1526–1534 (2019).
[2] Brambilla, E. et al. Prognostic effect of tumor lymphocytic infiltration in resectable non-small-cell lung cancer. J. Clin. Oncol. 34, 1223–1230 (2016).
[3] AbdulJabbar, K. et al. Geospatial immune variability illuminates differential evolution of lung adenocarcinoma. Nat. Med. 26, 1054–1062 (2020).
[4] Saltz, J. et al. Spatial Organization and Molecular Correlation of Tumor-Infiltrating Lymphocytes Using Deep Learning on Pathology Images. Cell Rep. 23, 181-193.e7 (2018).
[5] Keller, M. D. et al. Adverse prognostic value of PD-L1 expression in primary resected pulmonary squamous cell carcinomas and paired mediastinal lymph node metastases. Mod. Pathol. 31, 101–110 (2018).
[6] Borghaei, H. et al. Nivolumab versus Docetaxel in Advanced Nonsquamous Non–Small-Cell Lung Cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
[7] Schalper, K. A. et al. Objective measurement and clinical significance of TILs in non-small cell lung cancer. J. Natl. Cancer Inst. 107, 435 (2015).
[8] Wong, Kit Man, et al. "A Cost-Effectiveness Analysis of Using the JBR. 10-Based 15-Gene Expression Signature to Guide Adjuvant Chemotherapy in Early Stage Non–Small-Cell Lung Cancer." Clinical Lung Cancer 18.1 (2017): e41-e47.
Follow the Topic
-
npj Precision Oncology
An international, peer-reviewed journal committed to publishing cutting-edge scientific research in all aspects of precision oncology from basic science to translational applications to clinical medicine.
Related Collections
With Collections, you can get published faster and increase your visibility.
Minimal Residual Disease and Circulating Tumor DNA Dynamics in Personalized Cancer Treatment
Publishing Model: Open Access
Deadline: Mar 12, 2027
Genomic Instability
Publishing Model: Open Access
Deadline: Jun 24, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in