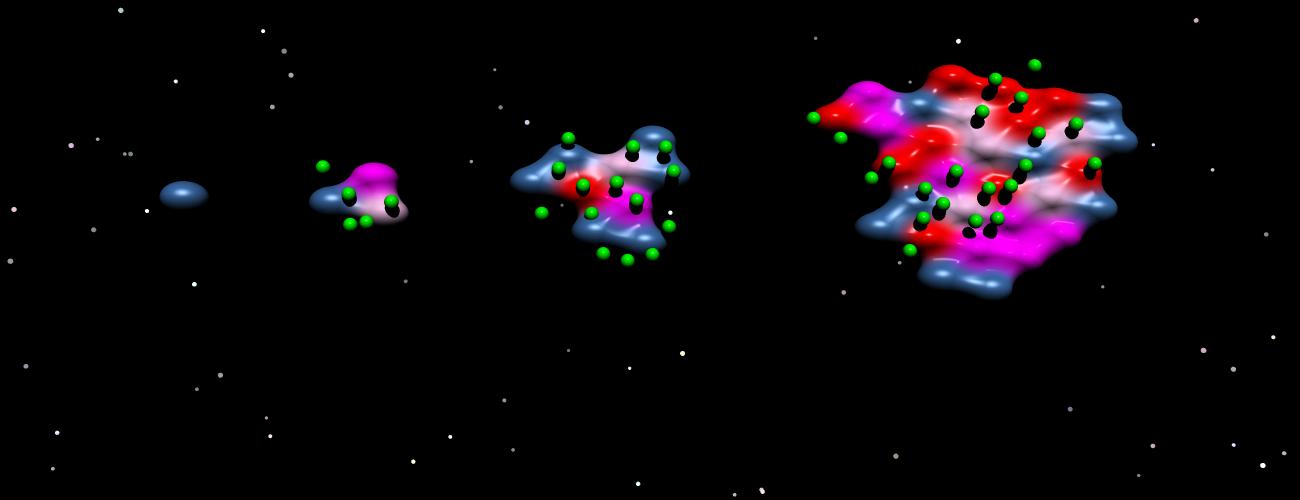

The classical understanding of cancer has been determined by observations made on tumor tissue obtained by surgical resection. However, this end-point perspective cannot account for one of the most fascinating aspects of cancer biology, the often long-year evolutionary process that is governed by principles of Darwinian evolution, eluding direct observation.
Cancer evolution recapitulates basic principles of biology in a nutshell, as surviving cell clones over time acquire mutations that allow them to thrive in an often hostile and changing environment. Adaptation processes leave their traces in the genome of manifest cancers, so studying cancer genomes can open a window into the past. In this sense, analyzing mutation signals in tumor genomes to unravel early steps of tumor formation shares similarities to cosmology research measuring cosmic microwave background radiation to shed light on the origin and history of the universe.
Importantly, studying cancer evolution is not only a fascinating exercise in basic science, it also has meaningful clinical implications. Individuals affected by Lynch syndrome, the most common hereditary cancer syndrome caused by germline variants in DNA mismatch repair (MMR) genes1, have a more than 50% life-time risk of developing MMR-deficient cancer. Therefore, there is a big medical need for novel approaches for effective tumor prevention in Lynch syndrome. For developing such prevention strategies, understanding the evolution of MMR-deficient cancers over time is essential.
Because MMR-deficient cells cannot recognize and correct small errors occurring during DNA replication, they accumulate an exceptionally high mutational load, particularly insertions or deletions (indels) of single nucleotides at repetitive sequences (microsatellite instability, MSI). The enormous amount of indels influences the phenotype of MSI tumors. After the onset of MMR deficiency, indels randomly affect microsatellites scattered over the genome. Whenever gene-encoding microsatellites are hit, the respective gene function can be abrogated. Importantly, the same events also lead to shifts of the translational reading frame, and frameshift peptides can be generated. Such frameshift peptides can be strong mutational neoantigens that make MSI cancers visible for the immune system and sensitive towards immune checkpoint blockade 2,3.
Using a genome-wide bioinformatics approach, our group has previously identified coding microsatellites as potentially relevant indel mutation hot spots 3. So far, genome-wide profiling of somatic mutations in MSI cancer has been hampered by the fact that indels at repetitive sequences are difficult to detect by next-generation sequencing approaches.
The present study4 was a collaboration between the Heidelberg University Hospital, the DKFZ Heidelberg, Heidelberg University, the Heidelberg Institute for Theoretical Studies (HITS) and international collaboration partners. We developed ReFrame, a new algorithm to quantitatively detect microsatellite indel mutations with high sensitivity. Using ReFrame, we identified mutations shared by most MSI colorectal and endometrial cancers.
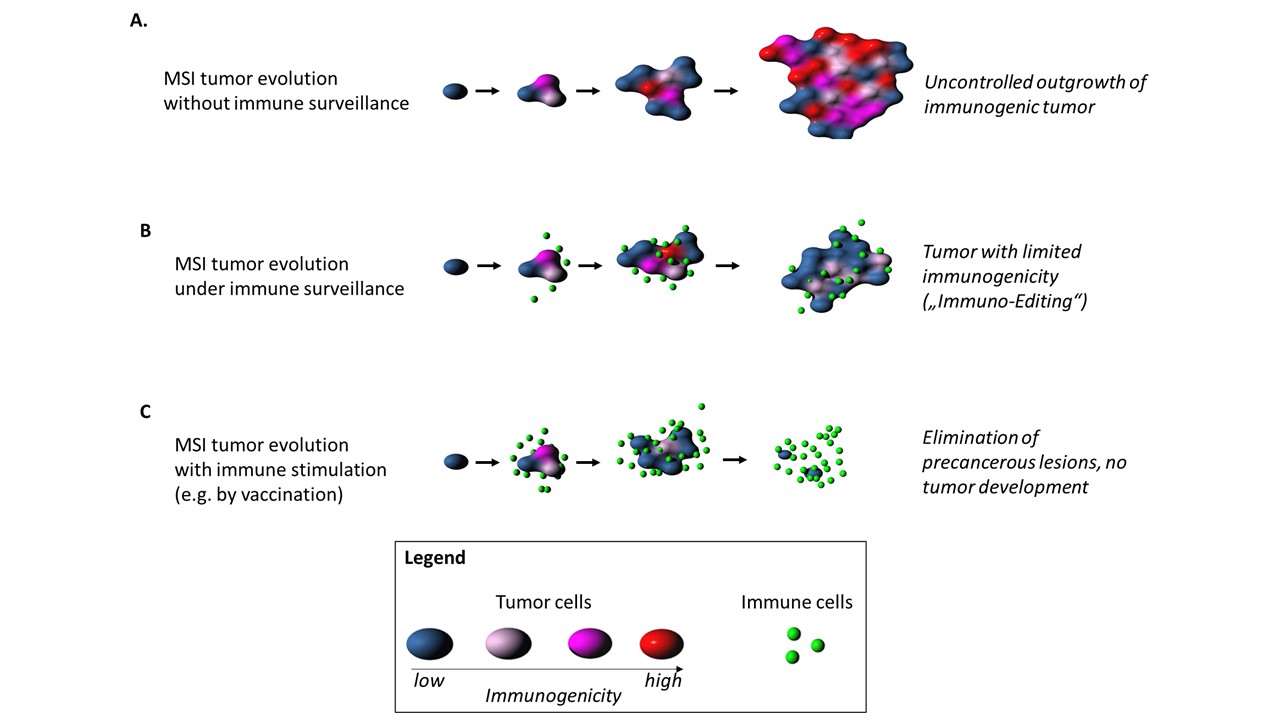
We further discovered a negative correlation between the prevalence of a defined indel mutation in MMR-deficient colorectal or endometrial cancers and the predicted immunogenicity of the resulting frameshift peptide. Our study strongly supports the concept of continuous immunoediting in human cancers and provides new evidence for the hypothesis that immunogenic cancers and pre-cancer cell clones can be attacked and potentially eradicated by the host’s immune system.
This strongly encourages the concept of neoantigen-based cancer-preventive vaccines that may in the future help to reduce tumor risk in Lynch syndrome and potentially beyond. We have recently demonstrated the safety and immunological efficacy of a prototype frameshift peptide vaccine in a phase I/IIa clinical trial5.
https://www.nature.com/articles/s41467-020-18514-5
The source code of the ReFrame algorithm can be accessed on GitHub (https://github.com/atb-data/neoantigen-landscape-msi).
References
1 Lynch, H. T., Snyder, C. L., Shaw, T. G., Heinen, C. D. & Hitchins, M. P. Milestones of Lynch syndrome: 1895-2015. Nature Reviews Cancer 15, 181-194, doi:10.1038/nrc3878 (2015).
2 Le, D. T. et al. Mismatch-repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409-413, doi:10.1126/science.aan6733 (2017).
3 Kloor, M., von Knebel Doeberitz, M. The immune biology of microsatellite-unstable cancer. Trends in Cancer 2, 121-131 (2016).
4 Ballhausen, A., Przybilla, M., Jendrusch, M. et al. The shared frameshift mutation landscape of microsatellite-unstable cancers suggests immunoediting during tumor evolution. Nature Communications 11, 4740, doi:10.1038/s41467-020-18514-5 (2020).
5 Kloor, M. et al. A Frameshift Peptide Neoantigen-Based Vaccine for Mismatch Repair-Deficient Cancers: A Phase I/IIa Clinical Trial. Clinical cancer research : an official journal of the American Association for Cancer Research 26, 4503-4510, doi:10.1158/1078-0432.CCR-19-3517 (2020).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in