Looking at long non-coding RNAs from a 3D structural perspective
Published in Protocols & Methods

Written by Tina Uroda and Marco Marcia. Poster image by Jean-Luc Pellequer.
Human cells are constantly producing RNA molecules. While a small fraction of RNAs gets translated into proteins, the vast majority exert other functions and act as non-protein-coding molecules. The most complex of those molecules are long non-coding RNAs (lncRNAs). LncRNAs are involved in vital cellular processes, i.e. cellular differentiation and DNA replication, and in severe diseases, i.e. developmental disorders, neurological and cardiovascular dysfunctions, and cancer. Despite the important cellular roles of lncRNAs, their mechanisms of action remain largely undiscovered.
Looking at lncRNAs from a 3D structural perspective would help us better understand how they act. However, the large size of lncRNAs (usually between 1,000 and 10,000 nucleotides) makes them challenging targets for structural characterization with existing methods. Computational structure prediction methods are still unreliable. Biochemical and biophysical methods require the production of abundant homogeneous samples, which cannot be achieved with the same procedures used to produce short RNAs. As a result, to date there is no experimental high-resolution 3D structure of a full-length lncRNA, and the structures of small lncRNA fragments are insufficient to provide a comprehensive description of the molecular architecture of lncRNAs. Thus, there is a need for new approaches and this is where our recent publication in Nature Protocols provides an important contribution.
We were specifically interested in the human maternally expressed gene 3 (MEG3), an lncRNA that fine-tunes development and cell proliferation. Interestingly, MEG3 splicing isoforms display different levels of biological activity, despite sharing identical exons that cover up to 85% of its sequence. Such observation suggested us that short variable exons could alter the biological function of MEG3 by introducing discrete structural changes. Moreover, using a combination of biochemical and cell-based assays, we experimentally discovered that MEG3 possesses functionally crucial “pseudoknots”, a recurrent type of tertiary interactions typical of highly structured RNAs. This discovery made the characterization of the 3D topology and shape of MEG3 imperative.
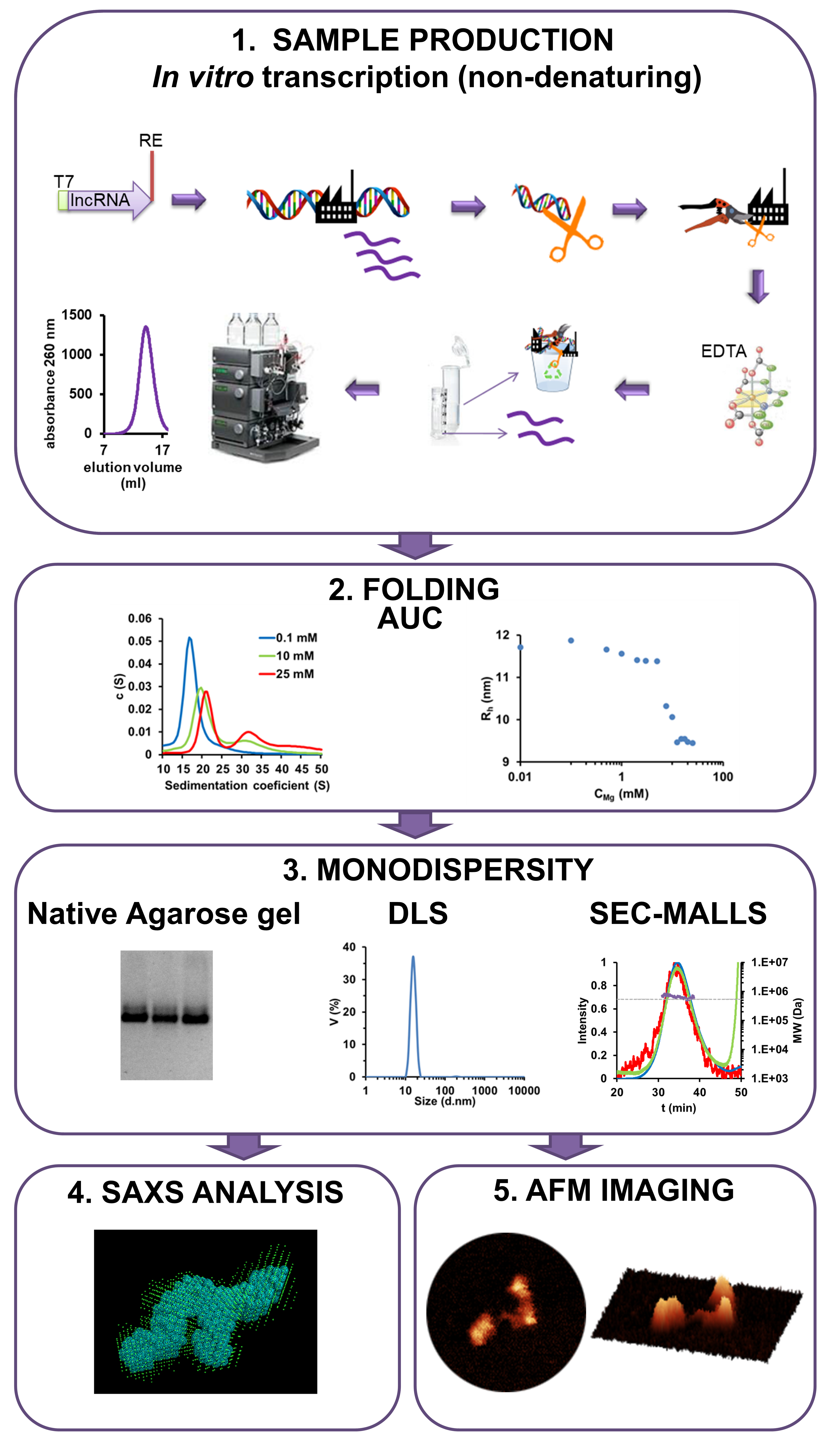
We thus designed this protocol to produce MEG3 by non-denaturing in vitro transcription, to determine appropriate folding conditions by analytical ultracentrifugation (AUC), to assess its monodispersity by electrophoresis and light scattering (MALLS, DLS), and finally to image it in 3D by small-angle X-ray scattering (SAXS) and atomic force microscopy (AFM). By applying this protocol, we were able to capture structural differences between active wild type MEG3 and inactive mutants with disrupted pseudoknots, providing for the first time a molecular visualization of functional and non-functional lncRNA particles. Our protocol thus constitutes a fundamental step forward onto the characterization of physiological 3D structural conformations of lncRNAs.
Our achievement could have impactful applications in many fields of life sciences since lncRNA are active research targets for epigeneticists, plant and developmental biologists, biochemists, and oncologists. Potentially, considering the expanding efforts to discover RNA-targeting drugs and the medical implications of many lncRNAs, our protocol could also be useful in the field of medicinal chemistry.
Follow the Topic
-
Nature Protocols
This journal publishes secondary research articles and covers new techniques and technologies, as well as established methods, used in all fields of the biological, chemical and clinical sciences.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in