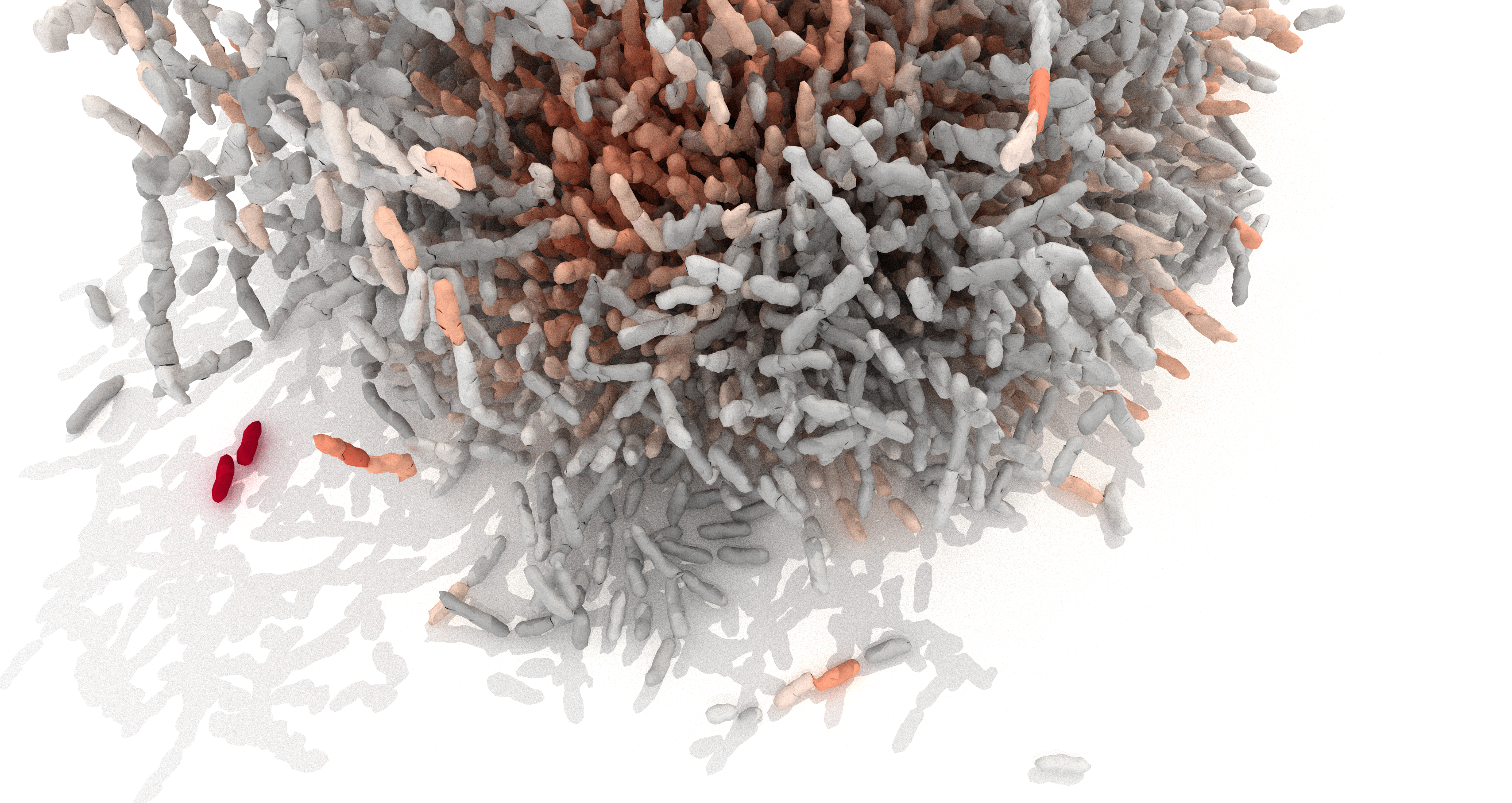
Which cell-cell interactions dominate during the development of bacterial biofilm architecture? One could imagine that such interactions are induced by a combination of cell metabolism, signaling, electric charge, thermal energy, and mechanical processes. In our study, we hypothesized that not all of these effects are equally important, and that perhaps mechanical cell-cell interactions could be dominant in shaping the architectural development of biofilms. However, the nature and magnitude of the forces between cells within biofilms were not known. Therefore, we aimed to shed light on the intercellular forces within Vibrio cholerae biofilms, and their importance in determining biofilm architecture, by combining experiments performed in the Drescher lab at the Max Planck Institute for Terrestrial Microbiology with theory developed in the Dunkel group at MIT.
Single-cell live imaging: observing biofilm growth at the cellular level
To begin, we required highly space- and time-resolved information on the dynamics of all single cells within biofilms. To this end, the Drescher lab developed a protocol for single-cell live imaging through automated confocal microscopy and novel image analysis. Using the developed protocol, we were able to observe complete 3D biofilm development at the single-cell level in many biofilms in parallel, with high enough temporal resolution to determine cell lineages and even measure individual cell growth rates (Figure 1). This rich dataset was the benchmark against which we could test all theories we could come up with for the relevant cell-cell interactions.
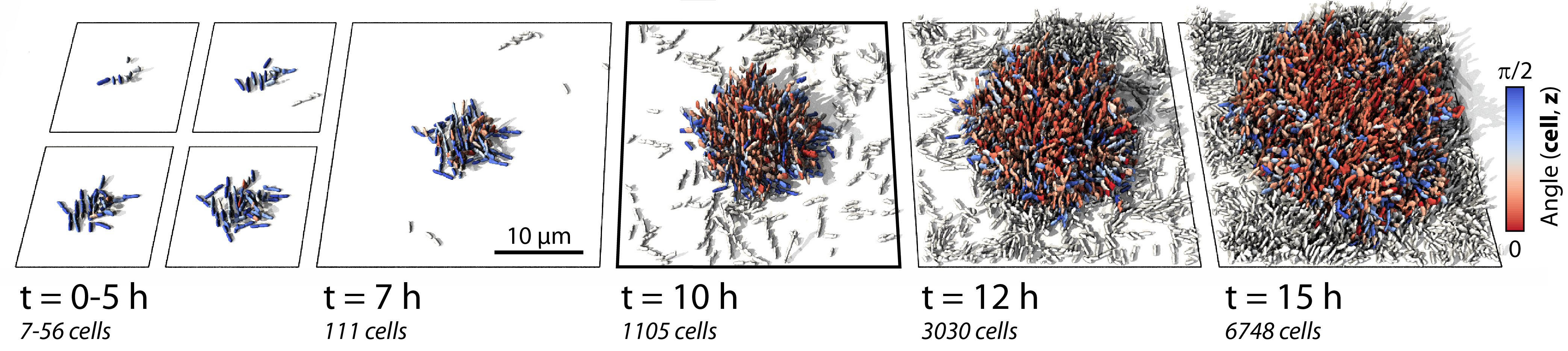
Figure 1. Time-resolved biofilm growth series. Each cell is colored by the cellular alignment with the z-axis. White cells are not direct descendants of the biofilm founder cell.
The golden ratio: attraction and repulsion
In parallel with the experiments, the Dunkel group developed a theoretical model containing the essential biophysical processes: cell growth, cell division, cell-surface interactions and cell-cell interactions. Crucially, we modeled cell-cell interactions using only two key mechanical ingredients: a repulsive force corresponding to steric interactions and osmotic pressure, and an attractive force corresponding to cell-cell linkage through the protein RbmA. We constrained these interaction forces in the simulations by quantitative comparison with experiments on genetically modified biofilms in which the production of the cell-cell-attraction-mediating protein RbmA was experimentally tuned. With this method, we leveraged the extensive experimental dataset to estimate the interaction forces between cells. Remarkably, even though we only compared global properties in constraining the forces, the optimized simulations reproduced local architectural properties from the experiments, including patches of highly aligned cells in biofilms with large amounts of attractive proteins (Figure 2).
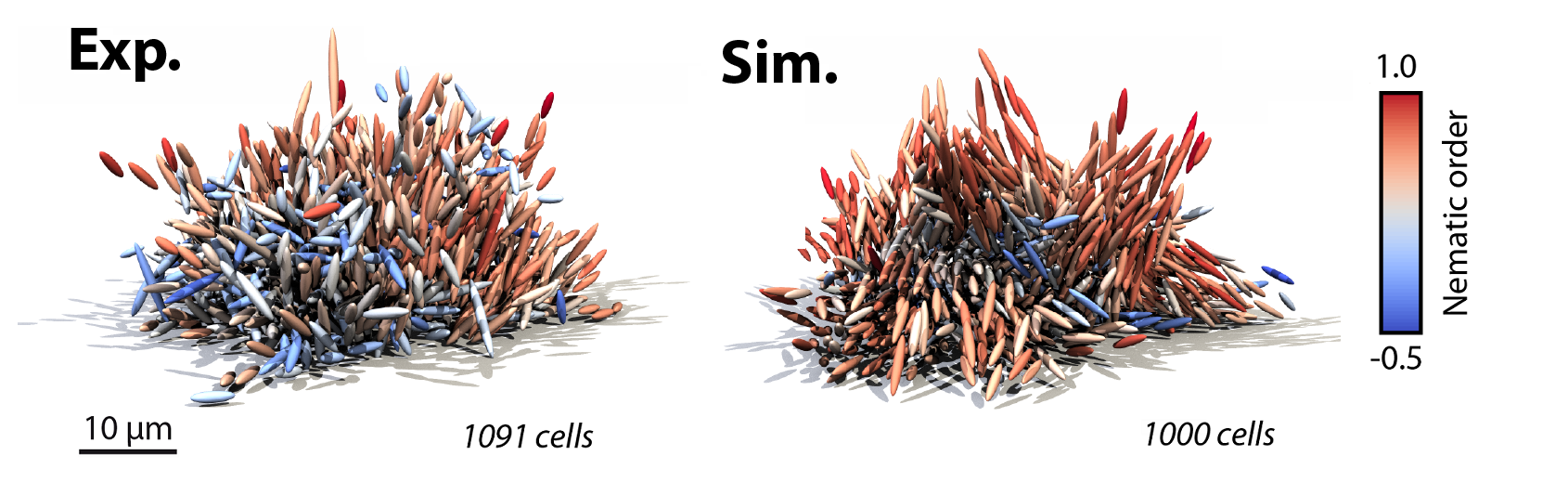

Figure 2. Simulation predictions (right) of biofilms show quantitative and qualitative agreement with experiments (left) with approximately 1000 cells. Each cell is colored by nematic order.
Towards a multi-scale understanding of biofilm growth
A fundamental quantitative understanding of the intercellular forces that sculpt bacterial biofilms requires a physical description in addition to knowledge of the molecular basis for cell interactions. Remarkably, our study showed that mechanical cell-cell interactions alone are sufficient for accounting for the development of V. cholerae biofilms up to a few thousand cells. For larger biofilms, our purely mechanical models of biofilm development differ from the experimentally observed biofilm architecture, leaving the verdict open as to which cell-cell interactions become dominant during the later stages of biofilm growth.
The paper in Nature Physics is here: https://www.nature.com/articles/s41567-018-0356-9
Follow the Topic
-
Nature Physics
This journal publishes papers of the highest quality and significance in all areas of physics, pure and applied.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in