Microbiome signatures of old- and young-onset colorectal cancer
Published in Cancer, Microbiology, and Genetics & Genomics
This’s my first research paper following my PhD training with Prof. Michael Inouye and Prof. Kathryn Holt in Melbourne and Cambridge.
My research journey with CRC microbiome traces back to 2012 when I collaborated with Prof. Jun Yu (@CUHK) and found bacterial taxa and functional genes associated with CRC1. Since then, the field has progressed fast, with well-established associations between CRC and gut microbiome2,3. However, few studies have delved into the gut microbiome concerning yCRC (aged <50), partly due to its lower incidence rate compared to old-onset CRC (oCRC, aged ≥50).
The gut microbiome is known to vary with age. With phenotypic differences in disease stage and tumor location between yCRC and oCRC4, one could hypothesize that the gut microbiome might also differ. Aiming to characterize the gut microbiome of yCRC, I collaborated with Prof. Pei-Rong Ding at Sun Yat-Sen University Cancer Center since 2021.
Negative findings
In our study, we counducted deep metagenomic sequencing on stool samples collected from a sizable cohort of CRC patients, including 167 yCRC. By comparing microbiome compositions between yCRC and oCRC, we observed only a few bacterial taxa that met the statistically significant threshold. Notably, the associations between age and microbial features across all studied samples were weak. Despite our rigorous efforts involving various analytical strategies, such as comparing individuals aged <40 to those aged >60, our investigations failed to yield promising results.
The Fudan Study
While we were under disappointment, a study by Yang et al from Fudan University working on the same topic published5. They reported distinct signatures of yCRC relative to oCRC. Our disappointment went deeper.
After thoroughly reviewing the Fudan paper multiple times, I found that their findings on oCRC were not corroborated well with other studies. One possible explanation is that their results primarily relied on 16S rRNA sequencing, although they were validated by shotgun metagenomic sequencing. Fortunately, the Fudan team also made the shotgun metagenomic sequencing (raw) data publicly available for 200 individuals. I downloaded and analyzed this data using our pipeline. Surprisingly, my findings diverged from those reported in the original paper. Notably, I found several well-known CRC-associated bacteria (Clostridium symbiosum, Hungatella hathewyi, Peptostreptococcus stomatis and Parvimonas micra) that were enriched in both oCRC and yCRC. Encouraged by these promising findings, we proceeded to integrate our data with the Fudan data, along with other published datasets, ultimately uncovering consistent signatures for both oCRC and yCRC.
Editor’s Support
Then, we found ourselves in a challenging position: eager to publish our results but lacking confidence in presenting contrasting findings to a previously published study. Despite our reservations, we decided to take a chance. I reached out to Dr. Javier Martinez-Vesga to discuss our intention to write a short paper detailing our contrasting results to a study published in Nature Communications. Interestingly, Javier suggested that we expand our scope and submit a full research article instead, highlighting the new data we had generated from numerous CRC patients.
Reviewers’ Support
During the peer-review process, we received positive feedback from all reviewers, who recommended conducting a more in-depth analysis on the bacterial genomes and exploring bacterial toxin and virulence factors. Following their suggestions, we conducted a more focused analysis, leading us to uncover insights into well-known CRC associated taxa with clear pathogenic pathways: Fusobacterium nucleatum, Bacteroides fragilis and Escherichia coli.
Story of F. nucleatum
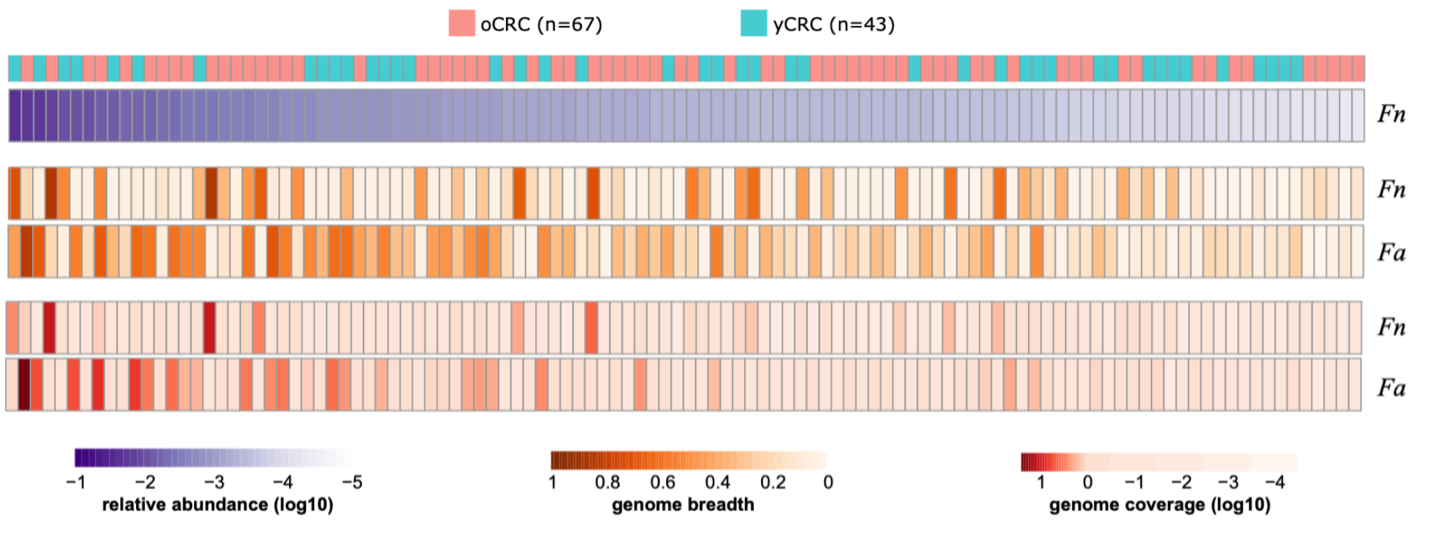
In our studied samples, we detected both F. nucleatum (Fn) and its close relative F. animalis (Fa; also known as Fn subspecies animalis, Fna). Strikingly, Fa had higher abundance and prevalence compared to Fn. This observation coincided with a very recent study published in Nature, which reported the enrichment of a distinct Fa clade in CRC niche6, consistent with an earlier study7. Moreover, in our Chinese cohort, we found no differentiation in the presence of the fadA gene (encoded adhesion protein A, mediates Fn attachment to and invasion of host epithelial cells) between samples enriched with either Fn or Fa.
Story of B. fragilis
We identified two distinct clusters of B. fragilis in our data. In line with a previous study8, each individual was dominated by one strain. The pathogenic B. fragilis strains have the capability to produce enterotoxin, which plays a role in cell cycle regulation. While the specific identities of these two strain clusters unknown to us, we observed differential abundance and prevalence of the enterotoxin-encoding gene bft between the two clusters.
Story of E. coli
Only one strain cluster of E. coli was identified in our samples. As a commensal bacterium in the gut, most E. coli strains are non-pathogenic. However, pathogenic strains often harbor the pks genomic island, which encodes colibactin, a compound implicated in DNA damage. Subsequently, we examined the relationship between the abundances of E. coli and pks. Interestingly, we observed a positive association in the Guangzhou cohort, but no correlation was evident in the Fudan cohort. This population heterogeneity implies a complex microbiome background of CRC across different geographic regions.
Potential impact
The shared microbiome signatures between yCRC and oCRC could serve as a foundation for developing microbiome-based diagnostic methods for CRC screening. In our study, we showed that microbiome-based classifiers trained for oCRC had similar prediction performance for yCRC, and vice versa. The consistency in signatures across CRC cases of different ages suggests that the microbiota may play a pivotal role in CRC carcinogenesis.
Ongoing work
One of the most intriguing questions is whether these microbial signatures correlate with CRC clinical outcomes. To address this, we are actively collecting follow-up data for many of the patients included in our study and investigating the relationship between gut microbiome and CRC outcomes. Since many of the CRC-associated bacteria are commonly found in the oral cavity3, we have also generated matched metagenomes from saliva/tongue samples for most of our patients. It’s worth noting that our current study only focused on bacterial taxa. However, considering that fungal and viral signatures have also been reported to be associated with CRC3, conducting further analyses on fungi and viruses will offer new biological insights into CRC pathogenesis.
Simultaneously, BGI is embarking on a mission to sequence 10 million microbiome samples as part of a large-scale cancer screening project for CRC. This endeavor will yield an unprecedented volume of microbiome data for humans. Science thrives on collaboration, and I extend a warm invitation to researchers worldwide to join us in deciphering this vast dataset and harnessing its potential for the greater good.
References
1 Yu, J. et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66, 70, doi:10.1136/gutjnl-2015-309800 (2017).
2 Wong, S. H. & Yu, J. Gut microbiota in colorectal cancer: mechanisms of action and clinical applications. Nature Reviews Gastroenterology & Hepatology 16, 690-704, doi:10.1038/s41575-019-0209-8 (2019).
3 Wong, C. C. & Yu, J. Gut microbiota in colorectal cancer development and therapy. Nat Rev Clin Oncol 20, 429-452, doi:10.1038/s41571-023-00766-x (2023).
4 Keum, N. & Giovannucci, E. Global burden of colorectal cancer: emerging trends, risk factors and prevention strategies. Nature Reviews Gastroenterology & Hepatology 16, 713-732, doi:10.1038/s41575-019-0189-8 (2019).
5 Yang, Y. et al. Dysbiosis of human gut microbiome in young-onset colorectal cancer. Nature communications 12, 6757, doi:10.1038/s41467-021-27112-y (2021).
6 Zepeda-Rivera, M. et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature, doi:10.1038/s41586-024-07182-w (2024).
7 Younginger, B. S. et al. Enrichment of oral-derived bacteria in inflamed colorectal tumors and distinct associations of Fusobacterium in the mesenchymal subtype. Cell Rep Med 4, 100920, doi:10.1016/j.xcrm.2023.100920 (2023).
8 Verster, A. J. et al. The Landscape of Type VI Secretion across Human Gut Microbiomes Reveals Its Role in Community Composition. Cell host & microbe 22, 411-419.e414, doi:10.1016/j.chom.2017.08.010 (2017).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in