The bioactive form of vitamin B6, pyridoxal phosphate (PLP), acts as a cofactor in more than 230 PLP dependent enzymes (PLP-DE), representing ~4% of all known catalytic activities. PLP-DEs are not only involved in many chemical transformations, but also play an important role in the regulation of immune function and the metabolism of neurotransmitters. Due to the involvement of PLP in basic metabolic processes, some PLP-DEs are crucial in diseases and recognized as drug targets.
Because of the diversity and evolution of PLP-DEs, resulting in five different fold types, their classification by sequence homology with known enzymes is difficult. In order to profile known and so far unassigned PLP-DEs, we developed a novel chemo-proteomic strategy for the global characterization of the PLPome of living cells. This strategy was inspired by activity-based protein profiling (ABPP) and involved the use of PL-derived probes.
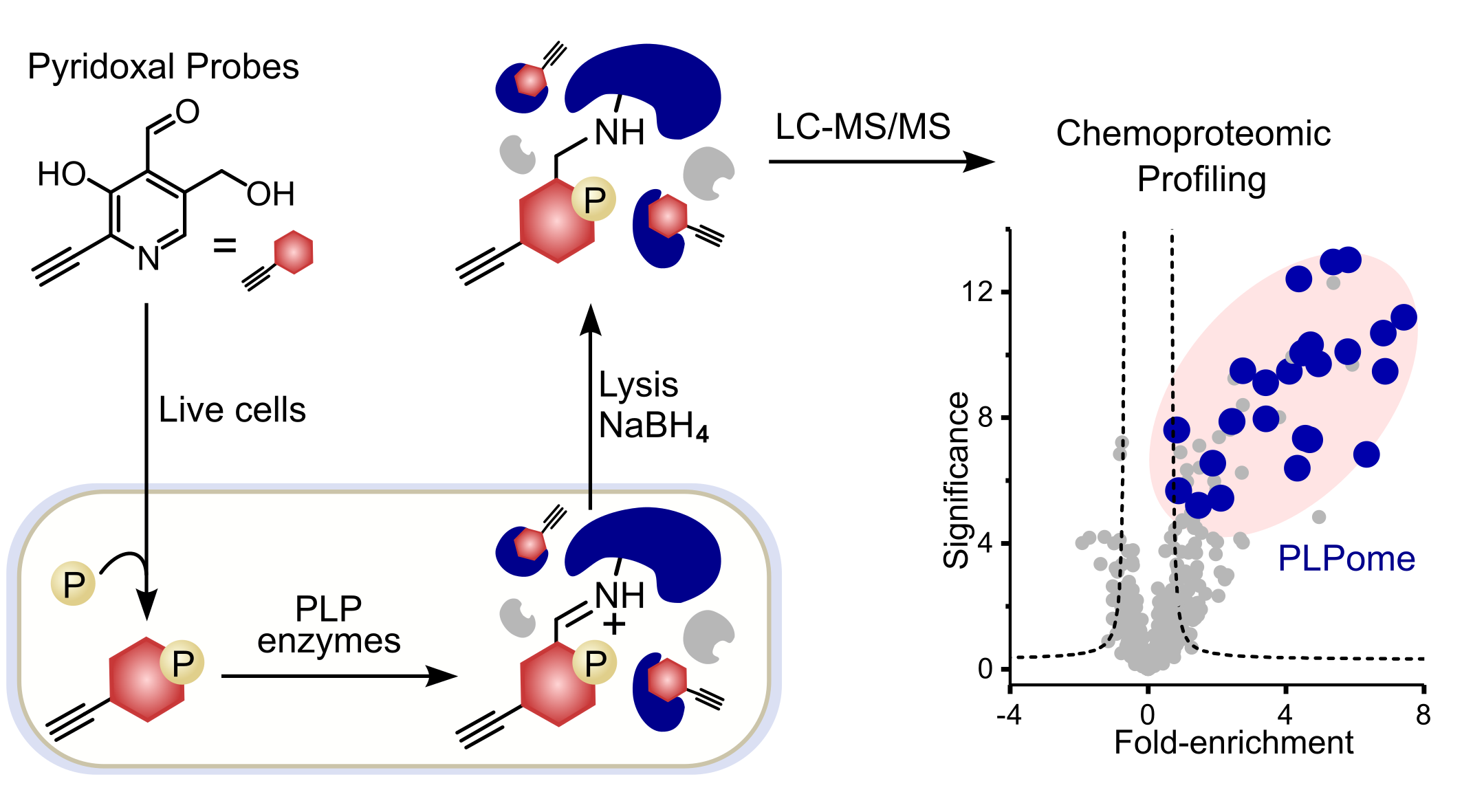
ABPP has evolved as a standard technique to uncover protein targets of small molecules. The workflow requires the modification of the molecule of interest with a terminal alkyne handle to a so-called ABPP-probe. Once the probe has bound to its dedicated protein targets within the cells, the handle can be modified with an azide-bearing biotin tag in a bioorthogonal click-chemistry reaction after cell lysis. Furthermore, using a biotin-avidin pull-down, proteins of interest are enriched among the complex cellular context and can be analyzed via SDS-PAGE or LC‑MS/MS (Figure 1).
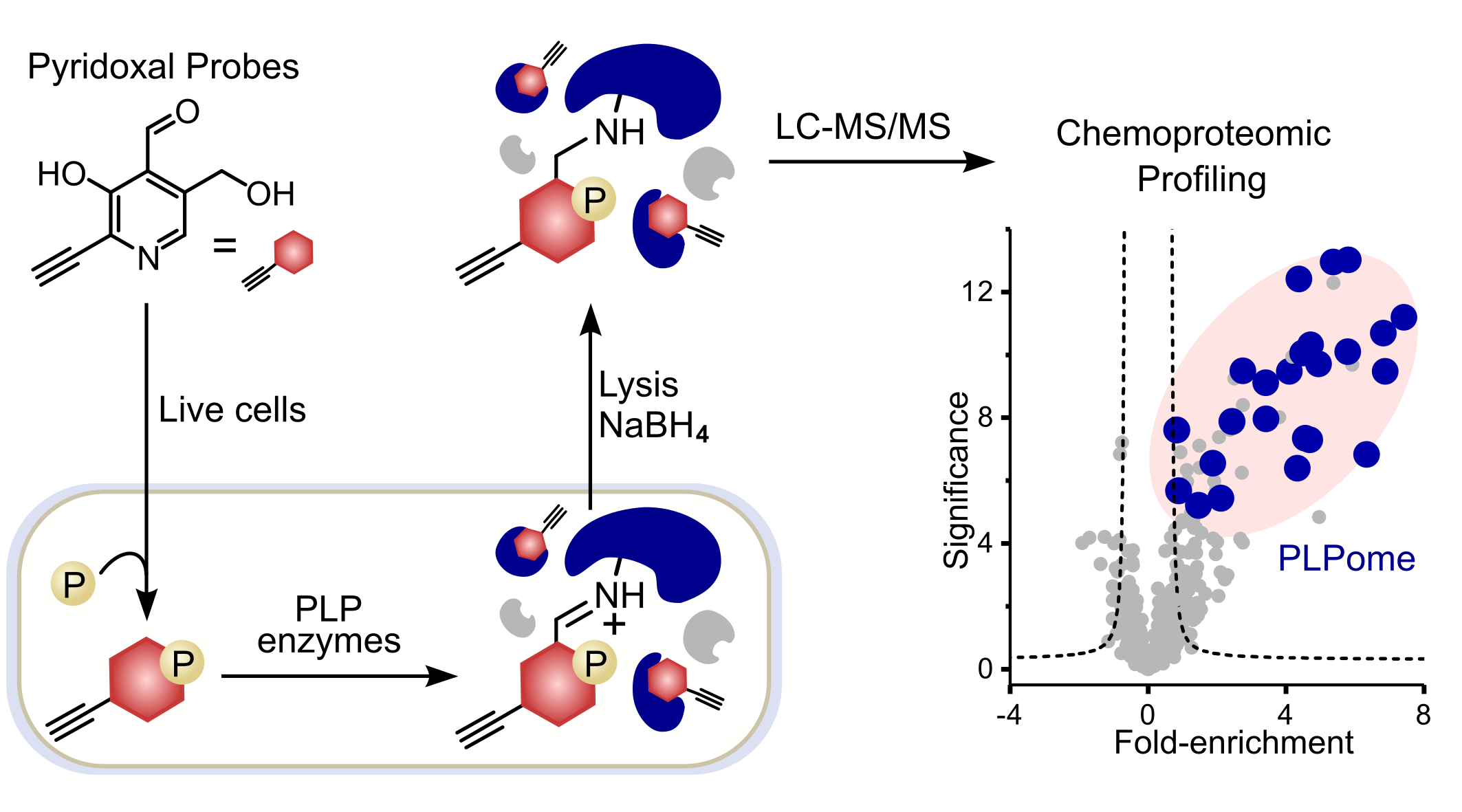
Figure 1: Schematic ABPP-workflow using functionalized PL-derivatives. Probes are metabolized within cells to the active form and covalently attached to protein targets after reduction using NaBH4. Downstream data analysis (two-paired students t‑test) resulted in significantly enriched proteins depicted in the upper right corner of the volcano plot. PLP-DEs are highlighted in blue.
In this work, S. aureus cultures were inoculated with pyridoxal-derived ABPP probes. As shown by targeted metabolomics, the bacterial pyridoxal kinase (PLK) accepted the probes as substrates and – comparable to natural pyridoxal (PL) – phosphorylated the primary alcohol group. These findings were additionally confirmed by monitoring the phosphorylation of the probes in kinetics assays using purified PLK.
Selective binding of the phosphorylated PL-mimics was proven by in vitro gel-based labelling experiments using purified enzymes, previously identified as PLP-dependent applying our MS-based approach.
Combining the results of proteomics experiments using our PL-derived probes, we could address 73% of the PLPome of S. aureus, including already known PLP-DEs, e.g. cysteine synthase. Thereby, confirming our method as a powerful tool to investigate the PLPome of living organisms (Figure 2). Furthermore, proteins of unknown function could be assigned as PLP-dependent using additional in vitro assays.
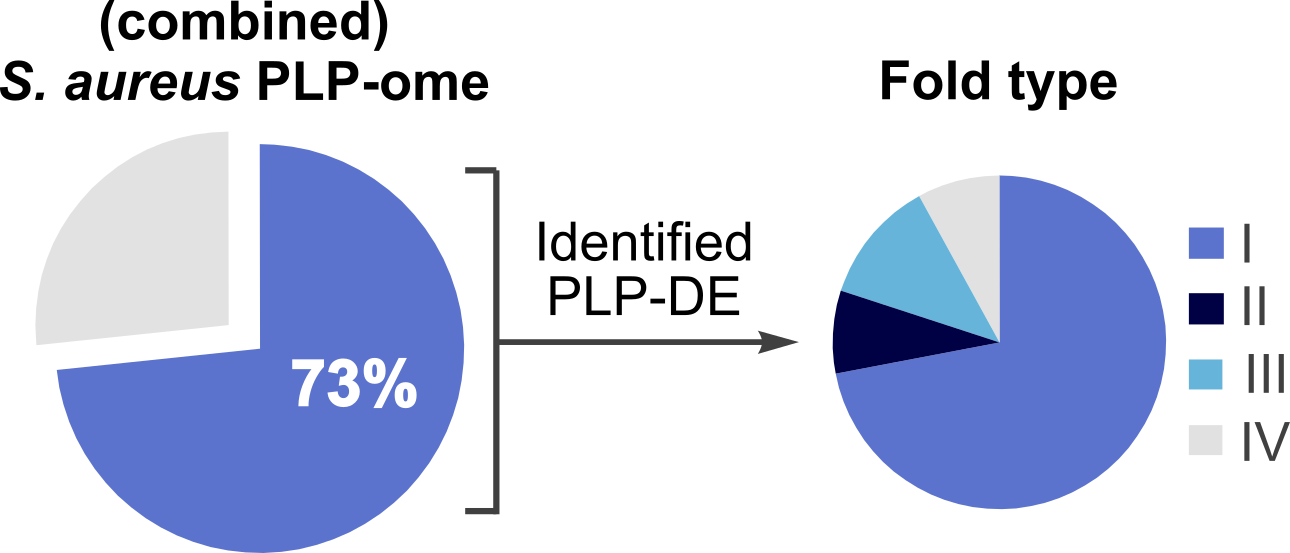
Figure 2: Covered PLPome of S. aureus combining MS-results of all probes. Overall coverage can be divided into the four different fold types as shown above.
You can access the full paper in our Nature Chemistry article here.
Written by Martin Pfanzelt and Volker C. Kirsch.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in