Monogenic Diabetes: Precision medicine starts with precision diagnostics
Published in Biomedical Research
Diagnostic genetic testing for monogenic diabetes has been available since the late 1990’s; however, implementation into clinical practice has been patchy, led by enthusiasts and researchers. Access to testing is variable between and within countries and remains a missed opportunity to implement precision medicine for diabetes. Monogenic diabetes accounts for about 1% of diabetes - common enough for most diabetologists to have a few cases on their books, but rare enough that it is difficult to assemble large enough groups of affected individuals for clinical trials or observational reports. Few have heard of monogenic diabetes, and even the name “maturity onset diabetes of the young” (shortened to MODY) is confusing.
Cases are often easy to spot for those in the know. *Hannah was diagnosed with diabetes at age 18 with a couple of Kg of weight loss and some mild thirst. Her mother, who also had diabetes, did a home glucose check and found it raised. Her fasting glucose was 10 mmol/l (180 mg/dl), she was slim with a BMI of 19 kg/m2 and she had no blood ketones. Her astute General Practitioner noted that Hannah didn’t easily fit into either the type 1 (T1D) or type 2 diabetes (T2D) “box” and Hannah was lucky enough to live near one of those enthusiastic diabetologists, who found that in addition to her subacute presentation she was negative for the antibodies associated with Type 1 diabetes. She had a genetic test for diabetes at diagnosis and never needed to inject insulin. Ten years later Hannah is still on a single tablet of gliclazide, an “old fashioned” diabetes tablet that works very well for her particular type of monogenic diabetes due to an HNF1A gene change.
Many people with monogenic diabetes have a less straightforward pathway to their diagnosis. For instance, *Rajiv age 30 was only diagnosed with monogenic diabetes through a research study and it took him 3 years to get on the most appropriate treatment. Despite not having obesity, he was assumed to have T2D because he was of South Asian heritage. Dividing diabetes into T1D and T2D has long been based on simple clinical criteria like age and body mass index and even ancestry. Therefore, many clinicians do not use the non-genetic tests that we have at our disposal like C-peptide or T1D antibodies which can help differentiate between T1D and other common or rare forms of diabetes. Thus, people with monogenic diabetes go unnoticed in our clinics. Diagnostic labs typically report an average 10-15 years between people being diagnosed with diabetes and then finding out they have a monogenic form of diabetes.
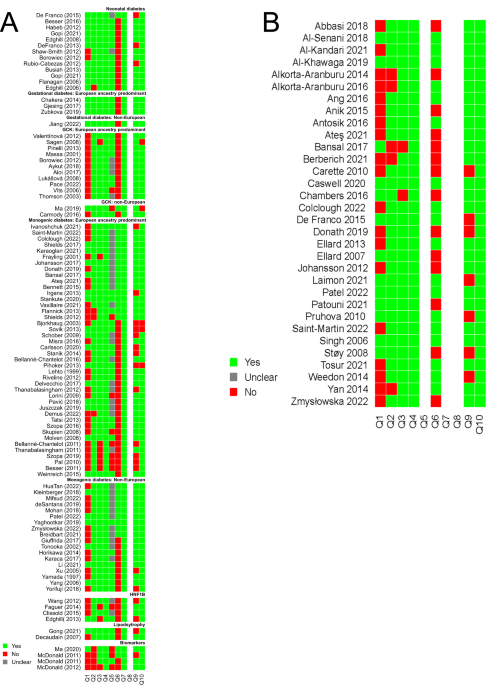
Even after 20 + years of looking after people with monogenic diabetes, there are still a significant number of uncertainties around diagnostic testing which the current review article aimed to address. We assembled an international team with collectively over 200 years of experience in diagnostic testing in monogenic diabetes to evaluate the current evidence and identify the gaps in our knowledge. One of the biggest questions was “Who should be tested?” Performing a systematic review of this question took up a great deal of our efforts. An ideal study would take hundreds of unselected people with diabetes, sequence them for all the known genes and thus extrapolate guidelines with high sensitivity and specificity for finding cases. It’s fair to say such ideal studies did not feature heavily in the literature search, but there were a great many case reports and small studies of highly selected groups of individuals. Our literature search initially produced over 10,000 individual publications, so we quickly had to make pragmatic decisions about which studies to include in our review. We chose to limit the papers included to those where at least 100 individuals had undergone genetic sequencing and used both a quality assessment score for diagnostic accuracy studies (QUADAS-2) and the Grading of Recommendations, Assessment, Development and Evaluation (GRADE) approach for certainty of evidence. We believe this is the first time such a thorough approach has been taken to the literature for monogenic diabetes. The Box shows our recommendations.

Another question we addressed was “How to test?”. We examined the evidence for the best sequencing method to use to diagnose monogenic diabetes. This showed that using Next Generation Sequencing panels simultaneously testing many genes rather than individual gene Sanger sequencing has an appreciably higher pick-up rate of monogenic diabetes due to the unexpected detection of rare subtypes. This was predictable (the wider you cast the net, the more fish you catch) but also demonstrates that rarer subtypes such as mitochondrial diabetes or HNF1B mutations which we expect to have additional syndromic features, may present with just diabetes.
Another question addressed was "What´s next?”, an area of practice which can be variable between clinical centres after a diagnosis of monogenic diabetes. We wrote a narrative review of the literature regarding this. The diagnosis and implications need to be delivered to the patient, and they may need investigation for other co-morbidities and family members may need to be counselled or screened. There may also be uncertainties around interpreting the result which requires testing other family members for co-segregation of diabetes with a variant of unknown significance. Negative genetic testing results may also need careful discussion. Diabetologists will need training in these aspects and/or involve specialist genetic counsellors.
We were on firmer ground when looking at some further questions: for example, whether a specific gene is a cause of monogenic diabetes, whether a variant in that gene is pathogenic or not, and how genetic tests should be reported. These issues have already been thoroughly addressed by expert organisations such as the American College of Medical Genetics & Genomics, Association for Molecular Pathology, Association for Clinical Genomic Science, and the Clinical Genome Resource, and we didn’t need to further review these questions; rather, we summarized the resources produced by these sources.
Finally, we reviewed ongoing challenges in the field of monogenic diabetes. We found several themes here. One of the biggest barriers is lack of access to genetic testing in many countries, which is predictable in a world where we don’t even have equal access to insulin. We also note that variants previously thought to be pathogenic may display reduced penetrance of diabetes particularly in unselected populations, suggesting that other factors are influencing the phenotype. Novel variants are challenging to interpret whether in commonly or rarely implicated genes. We also noted that the increasing volume of genetic sequencing data from large population cohorts with wider ethnic coverage has in many cases made interpreting genetic variation more difficult.
Overall, a huge amount of progress has been made from identifying the first genes for monogenic diabetes in 1992, introducing diagnostic genetic testing in the late 1990’s, and beginning the integration into clinical pathways over the following years. We hope the review will provide useful evidence-based recommendations for diabetologists and geneticists implementing molecular testing into their services and ultimately help more people with diabetes access precision medicine.
*names changed
Follow the Topic
-
Communications Medicine
A selective open access journal from Nature Portfolio publishing high-quality research, reviews and commentary across all clinical, translational, and public health research fields.
Related Collections
With Collections, you can get published faster and increase your visibility.
Public health and health governance in China
Publishing Model: Open Access
Deadline: Jul 31, 2026
Life Course Epidemiology
Publishing Model: Open Access
Deadline: Jun 30, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in