Mycofluidics of clocks in single filaments of Neurospora crassa
Published in Cell & Molecular Biology
Mycofluidics of clocks in single filaments of Neurospora crassa
Circadian rhythms are fundamental to many living systems. The clock genes underlying this process have been identified first in the model system, Drosophila melanogaster[1, 2], and then subsequently in other related model systems, such as Neurospora crassa[3, 4]. N. crassa also has the advantage of behavioral screens like yellow, fork to uncover clock-related genes in D. melanogaster, but it also has screens that were developed in parallel to link genes to their biochemistry in the cell[5]. N. crassa is unique in facilitating an understanding of the biochemical, genetic and physical basis of the clock.
While clock models have been developed to link the genes to particular pathways, usually the work is carried out on populations of cells to observe the average behavior of the clock on over 106-7 or more[6]. It would be desirable to understand the collective behavior of individual cellular clocks in cells to explain the origin of clock in tissues and whole organisms[7]. We experience the negative aspects of dis-synchrony among cellular clocks with jet lag, when cells are not in phase with each other. What is the origin of clocks in different cells?[7] This can only be addressed by measurements on single cells or single hyphae.
Even when clocks have been identified and shown to synchronize from the behavior in single cells, most of the work is done on cells not engaged in growth and cell division[8, 9], namely conidial cells. Allowing the cells to grow fundamentally changes the problem and requires the consideration of physical processes of growth, such as diffusion, transvection, and Taylor dispersion[10, 11]. The use of serpentine microfluidic devices enables the measurement of the clock in filaments as well as these physical processes[12].
There has been an exciting connection between cell growth through the TOR pathways and the clock, but the connection between the two is still largely uncharted. Part of this missing connection is connecting the clock with the underlying physical processes responsible for hyphal growth[10] and part with the biochemistry[13].
TOR regulates growth and metabolism through its response to environmental inputs in flies and mice and at the same time regulates circadian rhythms[14]. The exact mechanism of regulation is unknown at this time. TOR also plays a critical role in aging, thereby linking aging and the clock[15].
It would be desirable to know how the clock is tied to the growth of the filaments, the predominant life stage of the organism. The hyphal mat is an interconnected network of filaments that potentially allow a much richer repertoire of mechanisms for synchronization of clocks in the hyphal network.
In this work published today in Communications Biology [16] we link for the first time growth and the clock to underlying physical processes of advection and diffusion in growing filaments through the use of serpentine microfluidic devices. The paper is rather unique in providing a generally accessible model in MATLAB for this clock in a growing filament. Advection is the process of turbulent intake of media into a growing filament. We build on existing filament models[10, 11] and clock models[17, 18] to growth and the clock and show promising support for this model at the single filament level within the serpentine device[16]. In particular, the advection rate and diffusion rates of these models are measured, and the advection rate is an order of magnitude higher than the diffusion rate[16].
One of the outcomes of the model was the empirical identification of an advection zone, where there is substantial movement of the nuclei in a filament near the growing front of the filament; furthermore, one of the remarkable features of the model is its ability to produce stable banding patterns to explain conidiation both at the single filament level and at the macroscopic level of race tubes[16]. The key to making this happen in our model was placing a boundary condition on the size of the advection zone at the beginning and end of the observation period.
In addition to the advection zone and the banding patterns for conidiation, a final key feature predicted by the model was an explanation for long distance synchronization[16]. Advection and diffusion by themselves are not sufficiently high to explain synchronization on a millimeter (mm) scale[16]. The model itself provides a direct connection between the clock model and the physical processes of advection and diffusion so that long distance synchronization occurs through the wave front in the model. The most significant new contribution of the model is an explanation of a growth wave to explain mm scale, long distance synchronization[16].
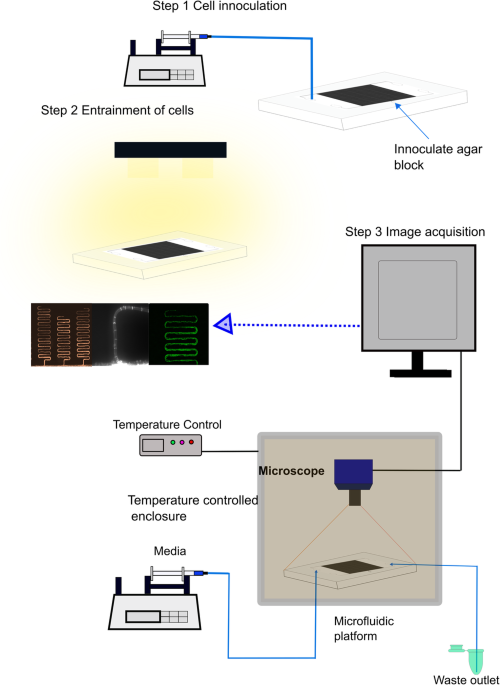
Using a serpentine microfluidics device, the model itself was validated both with frequency and phase data on fluorescent data from a clock promoter (MFNC9)[19]. The experiments with this device validated that single hyphae have clocks in which MFNC9 was shown to be circadian, have light entrainment, and temperature compensation[16]. The ability to see the clock in a single hypha builds on previous designs and is reminiscent of the patterns seen in race tubes on a macroscopic scale (Fig 1).
Fig. 1. Growth of single hyphae in a microfluidics device over ten days.
[1] A. Patke, M. W. Young, and S. Axelrod, "Molecular mechanisms and physiological importance of circadian rhythms," Nature reviews Molecular cell biology, vol. 21, no. 2, pp. 67-84, 2020.
[2] R. J. Konopka and S. Benzer, "Clock mutants of Drosophila melanogaster," Proceedings of the National Academy of Sciences, vol. 68, no. 9, pp. 2112-2116, 1971.
[3] D. Aronson Benjamin, A. Johnson Keith, J. Loros Jennifer, and C. Dunlap Jay, "Negative Feedback Defining a Circadian Clock: Autoregulation of the Clock Gene frequency," Science, vol. 263, no. 5153, pp. 1578-1584, 1994/03/18 1994, doi: 10.1126/science.8128244.
[4] J. C. Dunlap, "Molecular bases for circadian clocks," (in eng), Cell, vol. 96, no. 2, pp. 271-90, Jan 22 1999. [Online]. Available: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=9988221
[5] G. W. Beadle and E. L. Tatum, "Genetic Control of Biochemical Reactions in Neurospora," (in eng), Proc Natl Acad Sci U S A, vol. 27, no. 11, pp. 499-506, Nov 15 1941. [Online]. Available: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=16588492
[6] S. K. Crosthwaite, J. C. Dunlap, and J. J. Loros, "Neurospora wc-1 and wc-2: transcription, photoresponses, and the origins of circadian rhythmicity," (in eng), Science, vol. 276, no. 5313, pp. 763-9, May 2 1997. [Online]. Available: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=9115195
[7] J. H. Cheong et al., "The macroscopic limit to synchronization of cellular clocks in single cells of Neurospora crassa," Scientific Reports, vol. 12, no. 1, p. 6750, 2022/04/25 2022, doi: 10.1038/s41598-022-10612-2.
[8] Z. Deng et al., "Single cells of Neurospora crassa show circadian oscillations as well light entrainment and temperature compensation," IEEE Access, vol. 7, pp. 49403-49417, 2019.
[9] Z. Deng et al., "Synchronizing stochastic circadian oscillators in single cells of Neurospora crassa," Scientific Reports, vol. 6, no. 1, p. 35828, 2016/10/27 2016, doi: 10.1038/srep35828.
[10] M. Roper, C. Lee, P. C. Hickey, and A. S. Gladfelter, "Life as a moving fluid: fate of cytoplasmic macromolecules in dynamic fungal syncytia," Current Opinion in Microbiology, vol. 26, pp. 116-122, 2015/08/01/ 2015, doi: https://doi.org/10.1016/j.mib.2015.07.001.
[11] M. Roper and A. Seminara, "Mycofluidics: the fluid mechanics of fungal adaptation," Annual Review of Fluid Mechanics, vol. 51, pp. 511-538, 2019.
[12] K. K. Lee, L. Labiscsak, C. H. Ahn, and C. I. Hong, "Spiral-based microfluidic device for long-term time course imaging of Neurospora crassa with single nucleus resolution," Fungal Genetics and Biology, vol. 94, pp. 11-14, 2016.
[13] M. Judge et al., "Continuous in vivo metabolism by NMR," Frontiers in Molecular Biosciences, vol. 6, p. doi:10.3389/fmolb.2019.00026, 2019.
[14] V. A. Acosta-Rodríguez, F. Rijo-Ferreira, C. B. Green, and J. S. Takahashi, "Importance of circadian timing for aging and longevity," Nature Communications, vol. 12, no. 1, p. 2862, 2021/05/17 2021, doi: 10.1038/s41467-021-22922-6.
[15] M. Judge, J. Griffith, and J. Arnold, "Aging and the biological clock," Circadian Rhythms and Their Impact on Aging, pp. 211-234, 2017.
[16] J. H. Cheong et al., "The clock in growing hyphae and their synchronization in Neurospora crassa," Communications Biology, vol. 7, no. 1, p. 735, 2024/06/18 2024, doi: 10.1038/s42003-024-06429-6.
[17] A. M. Al-Omari, J. Griffith, A. Scruse, R. W. Robinson, H. B. Schüttler, and J. Arnold, "Ensemble Methods for Identifying RNA Operons and Regulons in the Clock Network of Neurospora Crassa," IEEE Access, vol. 10, pp. 32510-32524, 2022, doi: 10.1109/ACCESS.2022.3160481.
[18] W. Dong et al., "Systems biology of the clock in Neurospora crassa," (in eng), PloS one, vol. 3, no. 8, p. e3105, 2008. [Online]. Available: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=18769678
[19] E. Castro-Longoria, M. Ferry, S. Bartnicki-Garcia, J. Hasty, and S. Brody, "Circadian rhythms in Neurospora crassa: dynamics of the clock component frequency visualized using a fluorescent reporter," Fungal Genetics and Biology, vol. 47, no. 4, pp. 332-341, 2010.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
Mechanistic insights into human host and microbiome interactions
Publishing Model: Open Access
Deadline: May 31, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in