Native label-free protein sugars are cleaner and sweeter to identify, quantitate, and taste using CE-MS!
Published in Cancer
In the Ivanov laboratory at Northeastern University, our research is focused on the development of new and innovative sample preparation and nanoflow-based analytical liquid phase separation techniques coupled with mass spectrometry for biomolecular (e.g., proteomic and glycomic) profiling of limited amounts of complex biological and biomedically relevant specimens. By minimizing the number of sample preparation steps and employing more efficient, low volume (nL/sub-nL level)-based analytical techniques, we aim to detect, identify, characterize, and quantify more molecular features (e.g., proteins and N-glycans) compared to conventional techniques for which higher amounts of material are required.
A few years ago, when we started developing high-sensitivity capillary zone electrophoresis-mass spectrometry (CZE-MS) methods for N-glycan profiling of limited and heterogeneous biological samples (e.g., blood-derived extracellular vesicles (EVs) and total human plasma), we used the APTS tag to improve the detectability and separation of N-glycans. The combination of dopant-enriched nitrogen (DEN)-gas introduced into the nanoelectrospray microenvironment with optimized ionization, desolvation, and CZE-MS conditions improved the detection sensitivity up to ~100-fold, as directly compared to the conventional mode of instrument operation through peak intensity measurements. The comparative glycan analysis of IgG and EV isolates, and total plasma was conducted for the first time and resulted in the detection of >200, >400, and >500 N-glycans for injected sample amounts equivalent to <500 nL of blood [1]. Yet, we observed that the yield of APTS-labeling was less than 60%, and that some glycans remained mostly or completely unlabeled, especially highly sialylated ones. The complexity and heterogeneity of the three types of examined blood-derived specimens, and the high density of negatively charged and highly branched moieties on some glycans could explain the inefficient derivatization due to charge repulsion and steric hindrance factors. Nevertheless, such glycans could be detected in their non-labeled state without any further sample preparation and allowed us to increase the coverage of N-glycans detected in the three types of blood-derived isolates.
Based on this observation, we decided to develop an alternative strategy where the glycans would be analyzed in their native underivatized state. As in our previous work, carboxylic-coated magnetic beads were used for the enrichment of N-glycans released from glycoproteins by enzymatic digestion with PNGase F. However, contrary to our previous work, no subsequent derivatization with APTS was performed, and consequently, no cleaning steps were required to remove the excess of the derivatizing reagent (Fig.1). We also used optimized CZE-MS conditions by combining a DEN-gas with optimized levels of ion transfer tube (ITT) temperature and in-source collision-induced dissociation (ISCID). The developed label-free CZE-MS method resulted in a >45-fold increase in signal intensity compared to the conventional CZE-MS approaches used for N-glycan analysis. Strikingly, this non-labeling strategy resulted in the detection and quantitation of an increased number (~1.5-fold) of IgG-derived N-glycans compared to the APTS-labeling approach. Based on a direct and thorough qualitative and quantitative comparison of the N-glycans detected and identified in human serum IgG and human plasma EV isolates with the APTS-labeling and label-free CZE-MS methods using similar amounts of human serum and plasma, we demonstrated that higher levels of fucosylation and sialylation could be detected using the label-free approach due not only to a lesser extent of desialylation/defucosylation and sample loss during sample preparation but also because of lower levels of ESI- or in-source-induced decay.
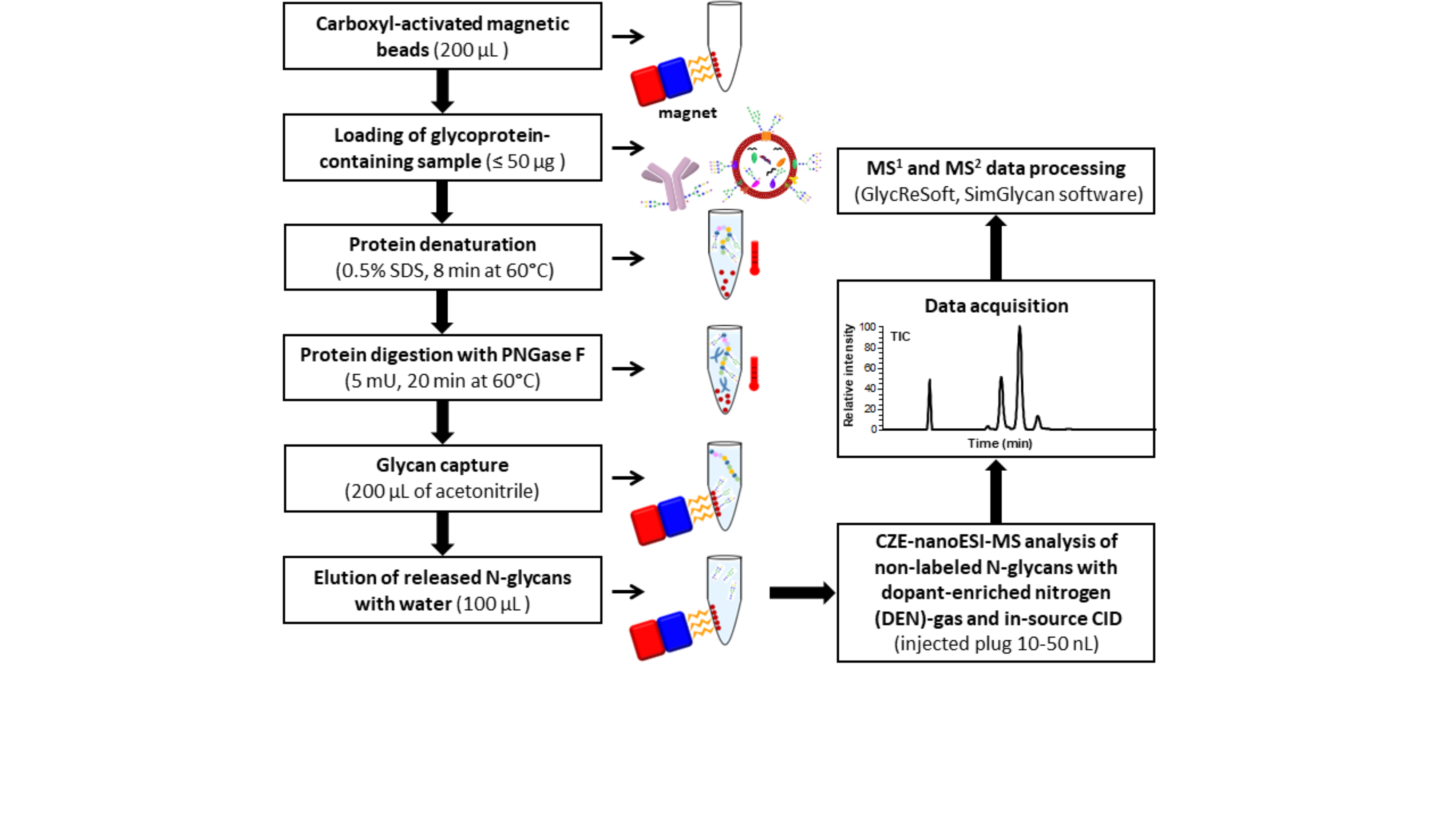 Fig.1: Analytical workflow for CZE-MS-based label-free glycan profiling of N-glycans released from biological sources (e.g., human or bovine sera).
Fig.1: Analytical workflow for CZE-MS-based label-free glycan profiling of N-glycans released from biological sources (e.g., human or bovine sera).
Finally, the high-sensitivity label-free CZE-MS method we developed resulted in the detection of >250, >400, >150, >310, and >520 N-glycans in human serum IgG, bovine serum fetuin, bovine pancreas ribonuclease B, blood-derived EV isolates, and total plasma, respectively, using injected amounts equivalent to <25 ng of model proteins and nL-levels of plasma-derived samples (Fig.2). Compared to reported results for biological samples of similar amounts and complexity, the number of identified N-glycans was increased up to ~15-fold, and unmatched profiling sensitivity was achieved for sub-0.2 nL of plasma volume equivalents. Also, previously undetected highly sialylated (up to 10 SiA residues) N-glycans were identified and structurally characterized, and untreated sialic acid-linkage isomers and positional isomers were resolved in a single CZE-MS analysis (Fig.2). We believe that the developed label-free CZE-MS technique appears as a promising approach for identifying potential glycan biomarkers in cancer or other human pathologies as well as in fundamental glycan profiling biology studies, using minute amounts of biological or clinical samples (i.e., liquid microbiopsies).
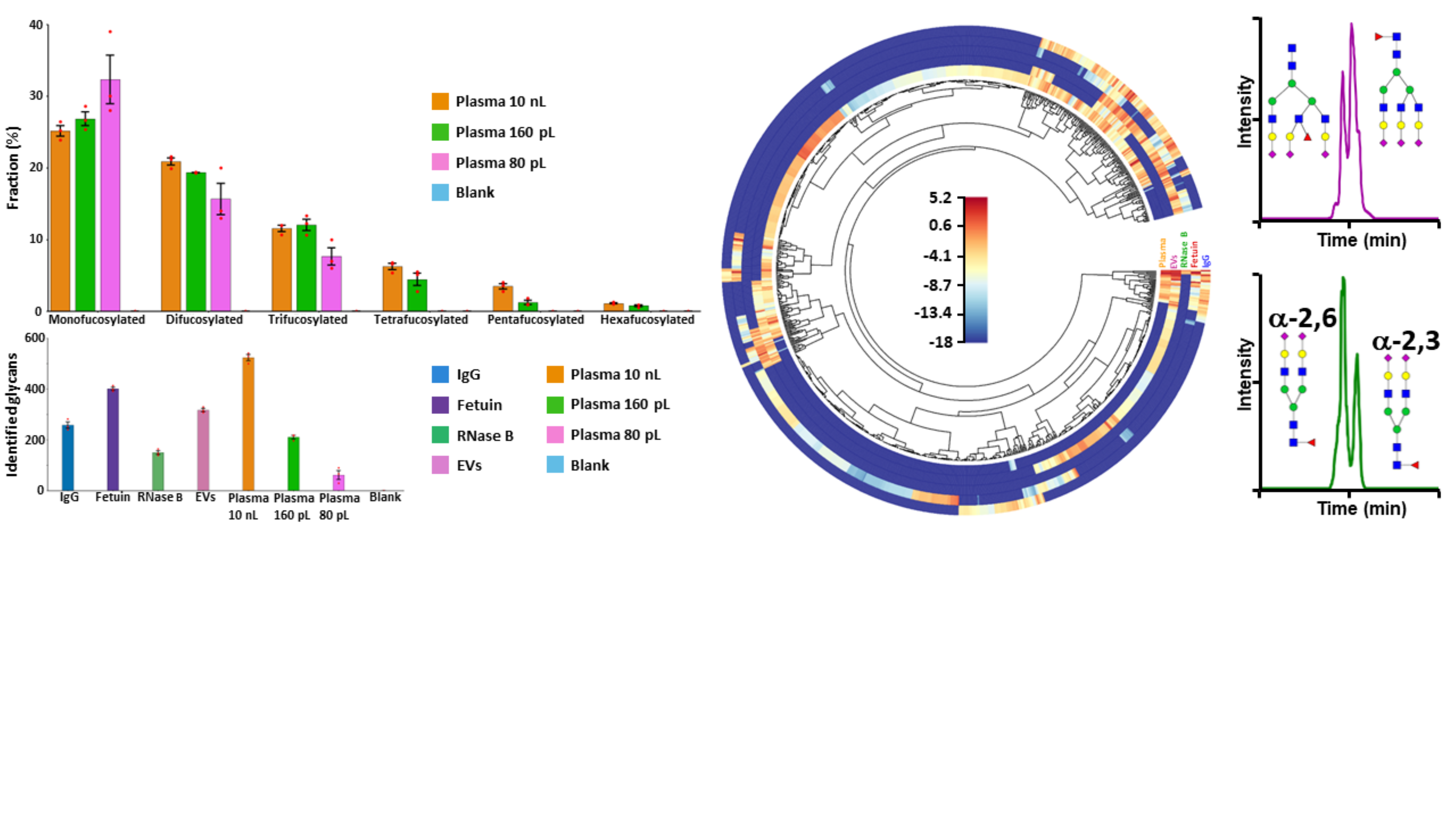
Fig.2: Number of N-glycans identified in the five types of analyzed biological samples using the label-free CZE-MS method, and fractional distributions of the fucosylated glycans detected in total plasma (left panel). Differential qualitative and quantitative N-glycan profiling of the five biological samples (middle panel). Differentiation of positional and linkage glycan isomers (right panel).
Reference:
1-Marie, A. L. et al. High-Sensitivity Glycan Profiling of Blood-Derived Immunoglobulin G, Plasma, and Extracellular Vesicle Isolates with Capillary Zone Electrophoresis-Mass Spectrometry. Anal Chem 93, 1991-2002, doi:10.1021/acs.analchem.0c03102 (2021).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in