New proteins from nowhere -- how evolution shapes the structure and function of a newly emerged protein in flies
Published in Ecology & Evolution
Many new proteins emerge from existing ones through gene duplication. Over the last decade, however, a new paradigm of protein evolution has come to light: functional proteins can also evolve from DNA sequences that did not previously encode a gene.
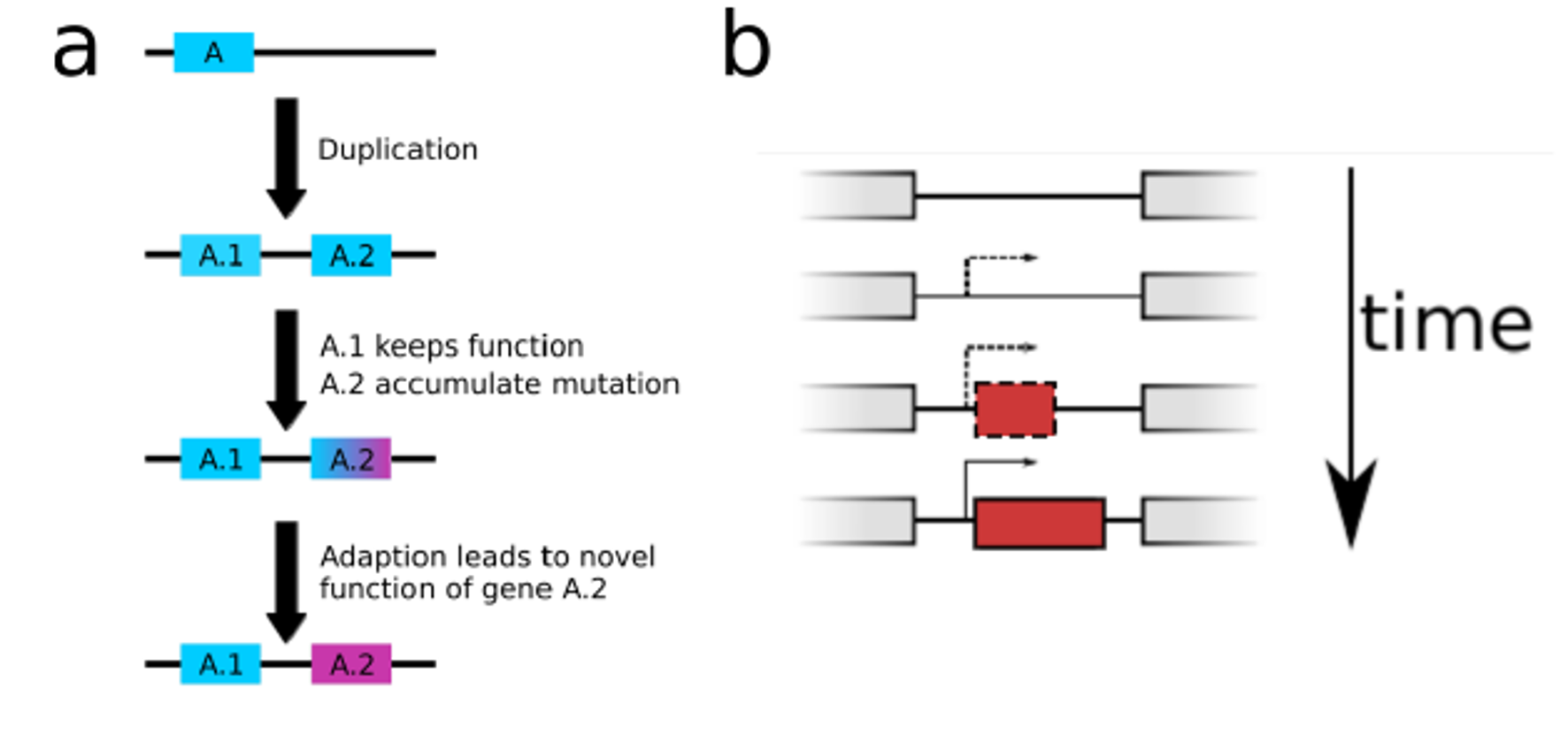
But what do such novel proteins look like upon birth? How do they change, and which functions do they assume as the “new kids on the block”? Three research groups spread over three continents combined their efforts to trace the evolutionary history of a de novo evolved fruit fly protein called Goddard.
The collaboration began completely by chance, when Prof. Geoff Findlay spoke at a meeting on insect reproduction at the University of Münster in 2015. In the audience was a graduate student from Prof. Erich Bornberg-Bauer’s lab. Conversation over coffee and a lunch with Erich the next day formed the basis for a productive collaboration. The team’s first paper together described a protein called Goddard, which appeared to be de novo evolved and whose depletion by RNA interference abolished the production of mature sperm in D. melanogaster [1].
The current project began in the summer of 2017, when Dr. Prajal Patel, who’d studied developmental biology during his Ph.D., joined Geoff’s group as a research associate. What initially excited him about de novo genes was figuring out how such newly-evolved and simple proteins integrate themselves into incredibly conserved developmental processes like spermatogenesis. Using CRISPR/Cas9 genome editing, Prajal deleted the goddard locus from flies and showed that spermatogenesis in the resulting mutants was arrested in the final stages of the process. He also observed that the protein localizes to the growing sperm tails, suggesting that the protein may play a role in organizing or stabilizing axoneme growth during sperm tail elongation.
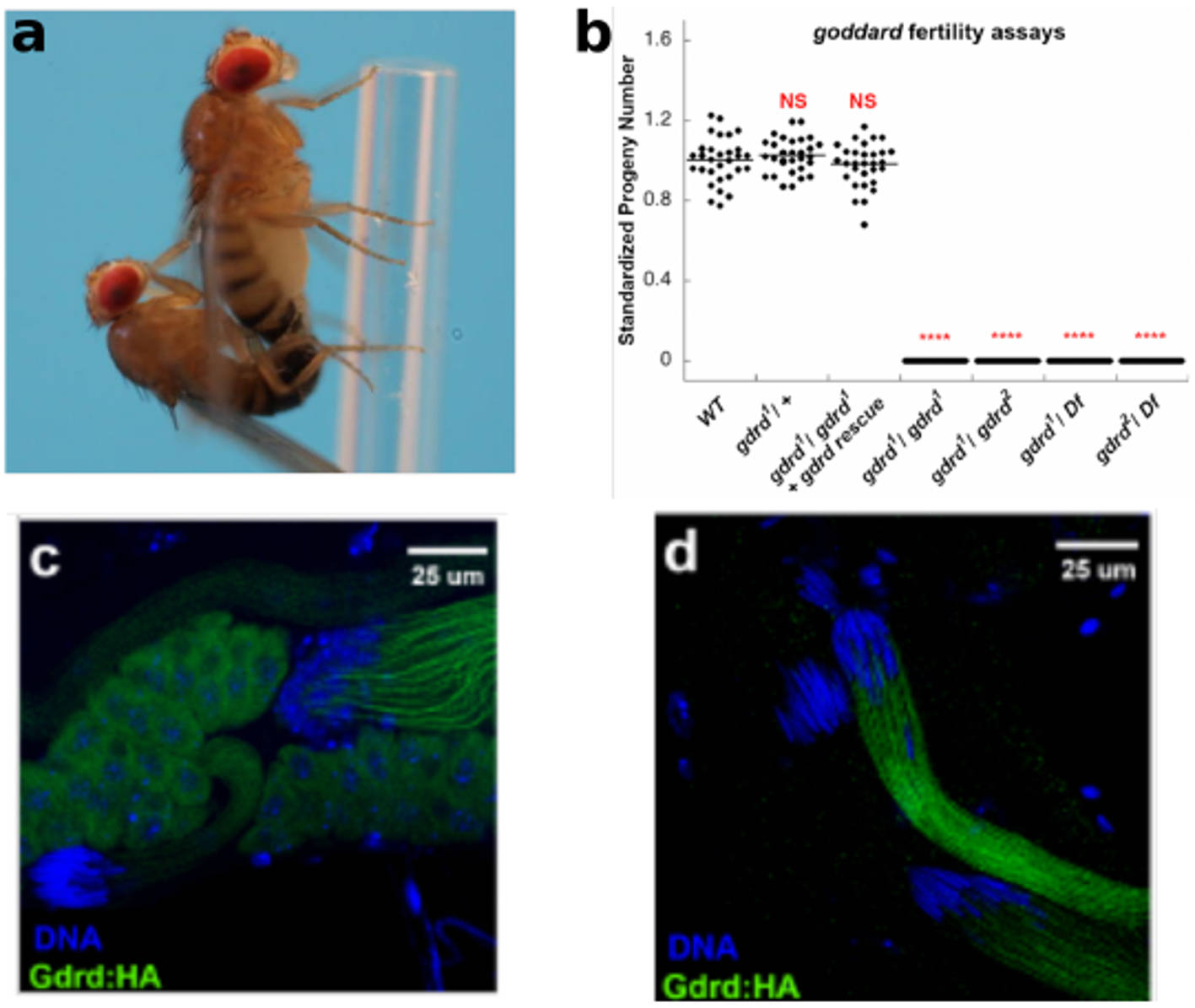
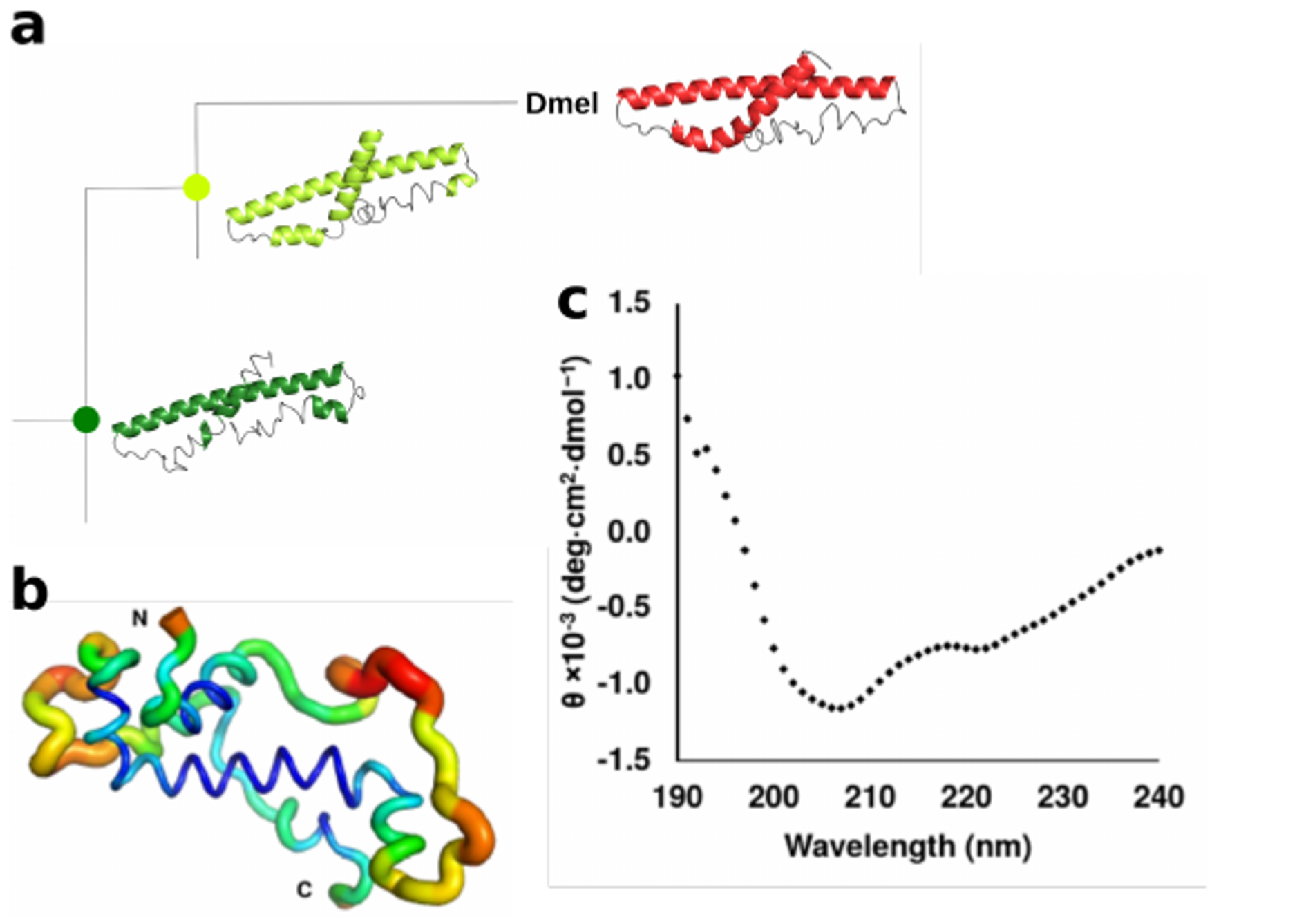
Meanwhile, two members of Prof. Erich Bornberg-Bauer’s group in Muenster, Germany, Dr. Andreas Lange and Brennen Heames, used biochemical and computational techniques to ascertain the shape of the novel protein in present-day flies. Andreas, who studied protein-ligand interactions during his Ph.D. and joined the group in October 2016, worked on both the expression and structural and functional validation of de novo proteins present in Drosophila and humans. He was keen on understanding how these small and young proteins can already have such an impact in different cellular processes. Brennen joined in the spring of 2017 and helped analyze de novo proteins mainly from Drosophila. Interested in how proteins arising from essentially random sequences can acquire folded and functional structures, Brennen used computational and experimental approaches to identify and characterize de novo proteins as part of his PhD. Brennen and Andreas then used evolutionary methods to reconstruct the likely structure of Goddard ~50 million years ago, when the protein first arose. What they found was quite surprising. In contrast to another recent study showing large changes in the sequence and structure of a de novo protein in yeast [2], the ancestral Goddard protein appeared very much like the ones existing in fly species today. From its very beginning, Goddard contained some structural elements, so called alpha-helices, which are believed to be essential for most proteins.
To confirm these findings, the scene shifted to the Australian National University in Canberra, where Dr. Adam Damry and Prof. Colin Jackson used intensive, computational simulations to verify the shape of the Goddard protein predicted by Andreas and Brennen. Colin, who had co-authored a paper based on epistatic networks, visited the Bornberg-Bauer lab in the summer of 2018, where he offered to help analyze the physical characteristics of de novo proteins. Both Adam and Colin validated Andreas’ structural analysis and showed that Goddard, in spite of its young age, is already quite stable – though not quite as stable as highly conserved fly proteins, most of which have been in existence for perhaps hundreds of millions of years. Altogether, Goddard’s structure appears to have been maintained with only minor changes over this long time span.
You can find the whole story on Nature Communications.
Several open questions remain: (i) when and how did Goddard become essential for the process of spermatogenesis, and (ii) how functional are the ancestral Goddard proteins? The collaborators are eager to continue working together to answer these questions.
[1] Gubala AM, Schmitz JF, Kearns MJ, Vinh TT, Bornberg-Bauer E, Wolfner MF & Findlay GD. The goddard and saturn genes are essential for Drosophila male fertility and may have arisen de novo. Mol. Biol. Evol. 34, 1066–1082 (2017).
[2] Vakirlis N, Acar O, Hsu B, Castilho Coelho N, Van Oss SB, Wacholder A, Medetgul-Ernar K, Bowman RW 2nd, Hines CP, Iannotta J, Parikh SB, McLysaght A, Camacho CJ, O'Donnell AF, Ideker T, & Carvunis AR. De novo emergence of adaptive membrane proteins from thymine-rich genomic sequences. Nat. Commun. 11, 781 (2020).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in