Novel enzymatic cascades for C-H-activation of aromatic and hetero-aromatic compounds
Published in Chemistry
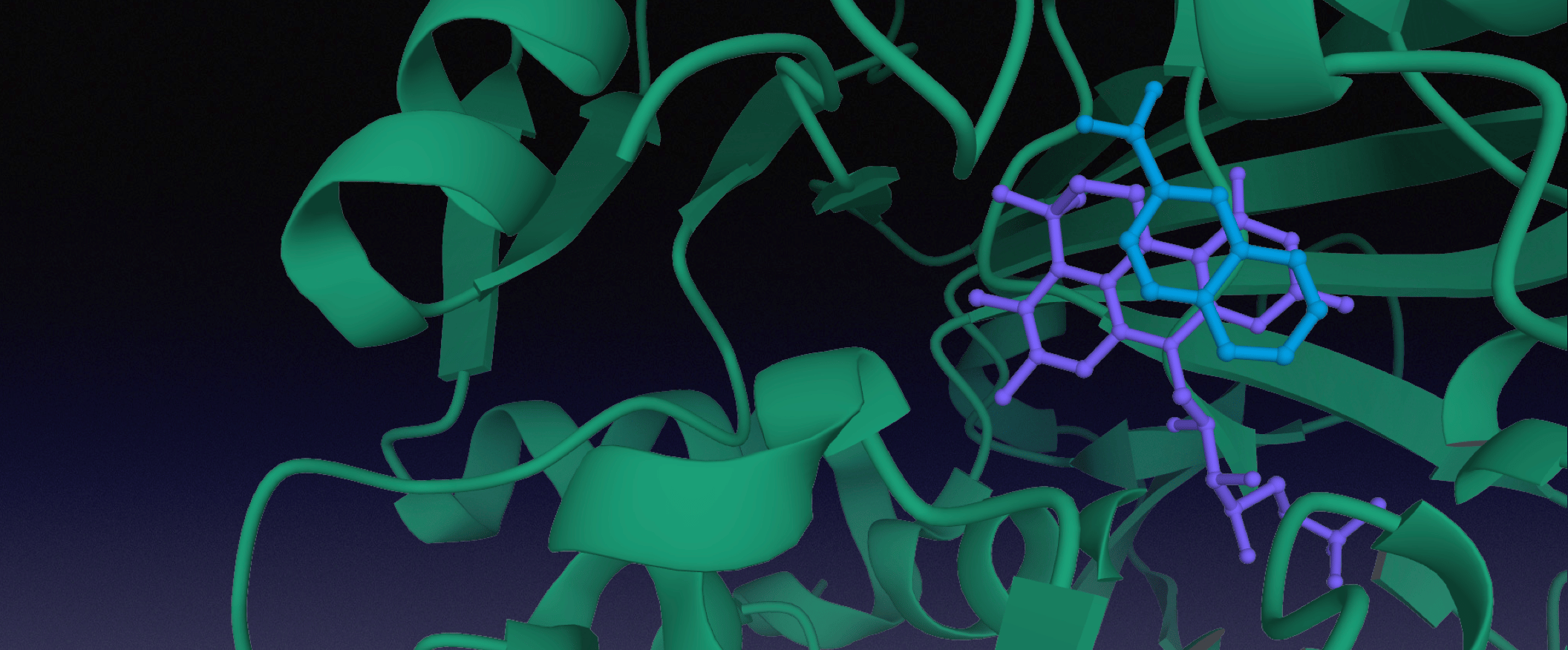
In this paper we look into biocatalytic routes for C-H functionalisation (through CO2-fixation) of a number of aromatic and heteroaromatic compounds. Some of these play part in polluting our natural resources (styrene, naphthalene), while others (N-, O- and S-containing heteroaromatics) serve as important “building blocks” for the chemical and pharmaceutical industries.
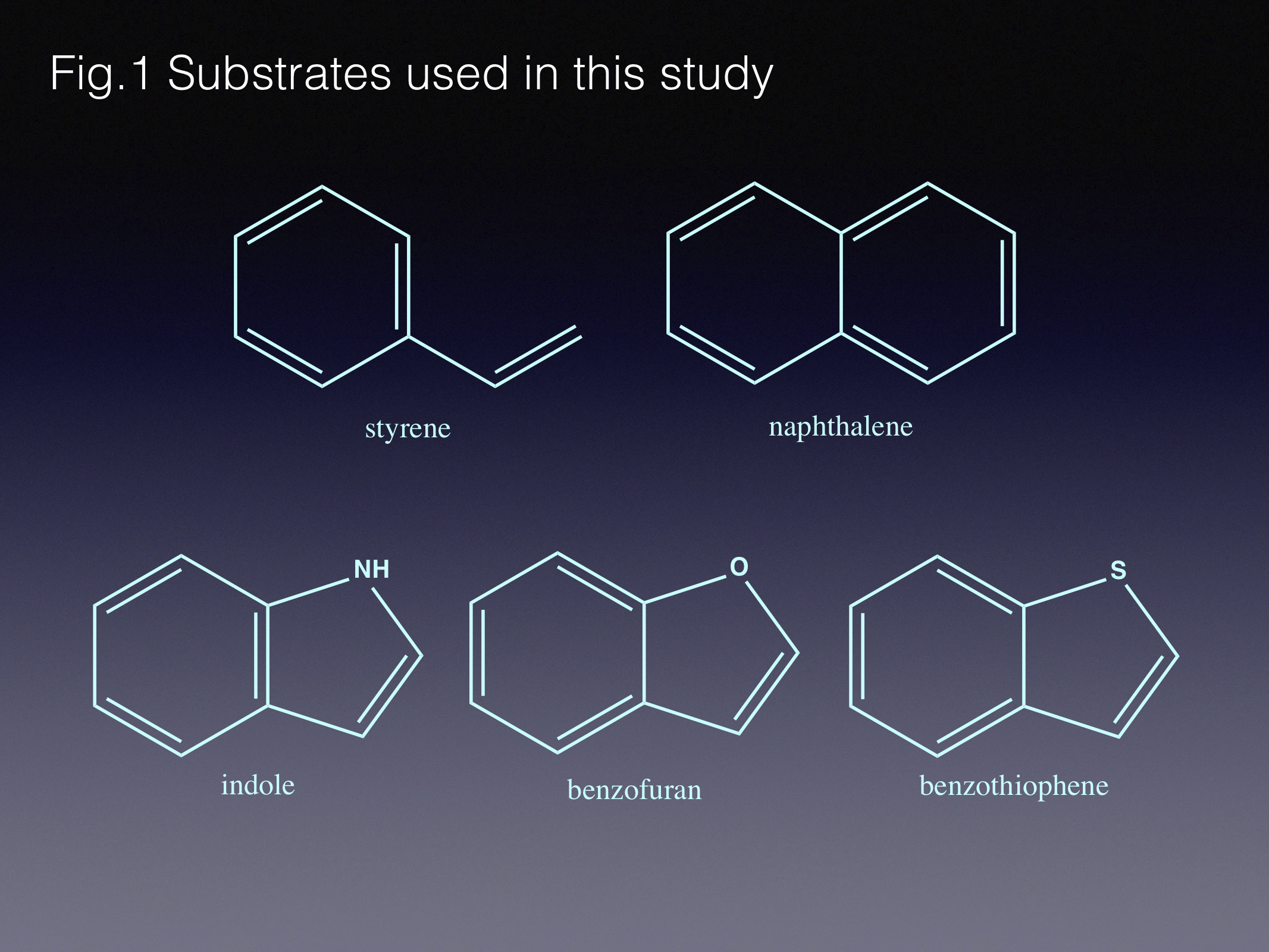
Here we use one of our group’s favourite enzymes, fungal ferulic acid decarboxylase - Fdc1. We have been particularly interested in this and related enzymes, since our lab discovered that they rely on something completely unexpected – a heavily modified flavin, prenylated FMN (prFMN) [1]. The reaction mechanism supported by the prFMN cofactor turned out to be unexpected for an enzymatic process as well: it includes a 1,3-dipolar cycloaddition between substrate and cofactor. While commonly used in organic synthesis, 1,3-dipolar cycloaddition is considered highly unusual for biological systems. Furthermore, the enzyme ensures that this process occurs in a reversible manner [2]. Hence, Fdc1 and related enzymes can also act as carboxylases.
In this current paper we describe how we combined the Fdc1 (de)carboxylase with carboxylic acid reductase, ensuring the reaction equilibrium occurs in the carboxylation direction, achieving ambient CO2 fixation. In addition, we tested the enzyme with new (and more interesting!) substrates, and expanded the biocatalytic cascade to create aldehyde-, alcohol-, amide- and amine- derivatives. As expected, the initial yields of the new reactions were modest at best, and required protein engineering to widen the substrate scope.
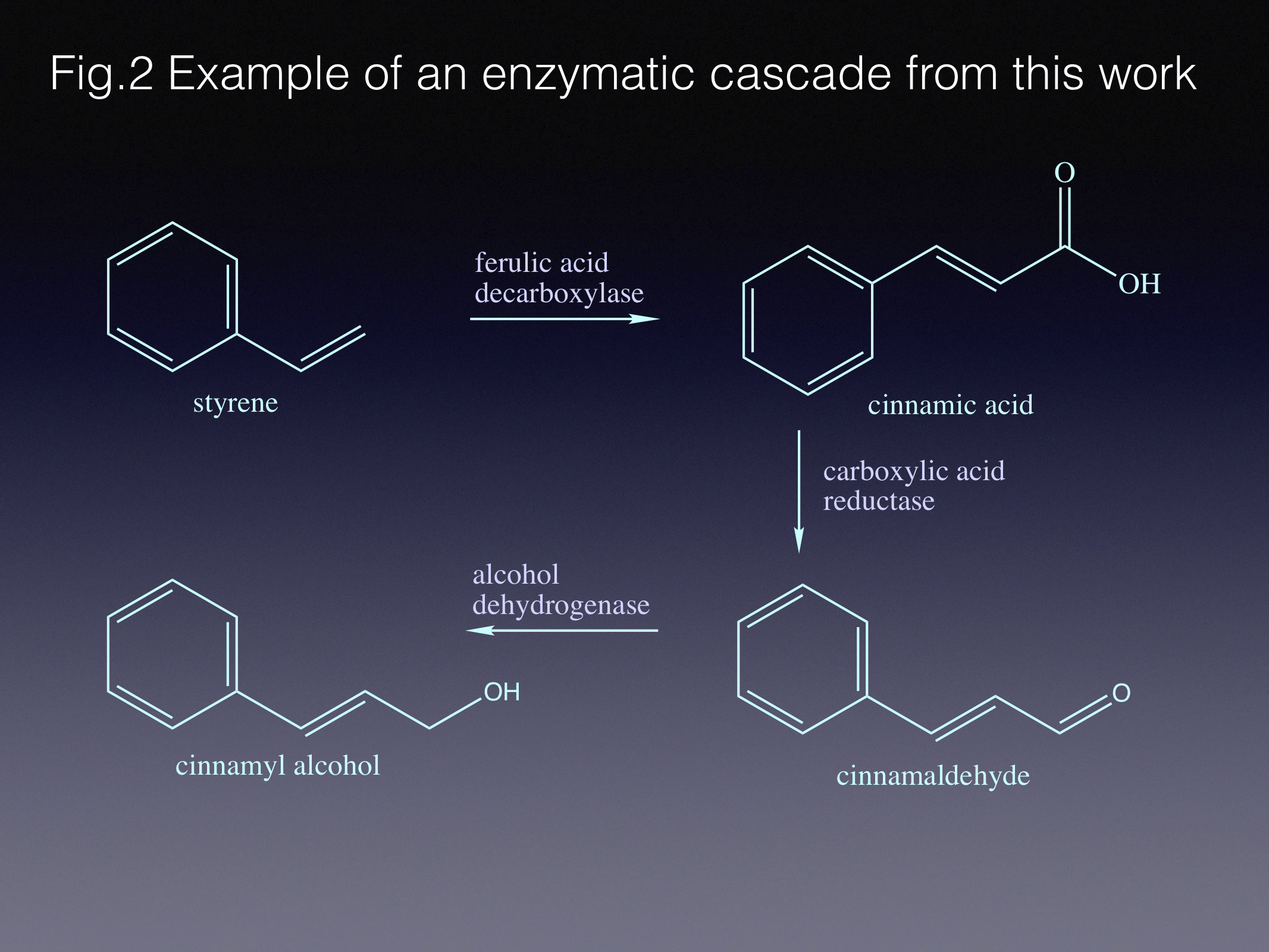
There are different ways of approaching the task of protein engineering. In the course of this work we limited our enzyme library size by getting an insight into how the active site works before creating the mutants – the so-called semi-rational approach. Indeed, our lab heavily relies on structural biology to help us understand enzyme function. In this project we determined that “weaker” substrates adopt slightly different orientations in the active site, when compared to the native pair of substrate/product - styrene and cinnamic acid. That informed us about the mutations that would potentially improve the substrate positioning in the active site. The best variant gave us 150-fold improvement in specific activity compared to the wild type!
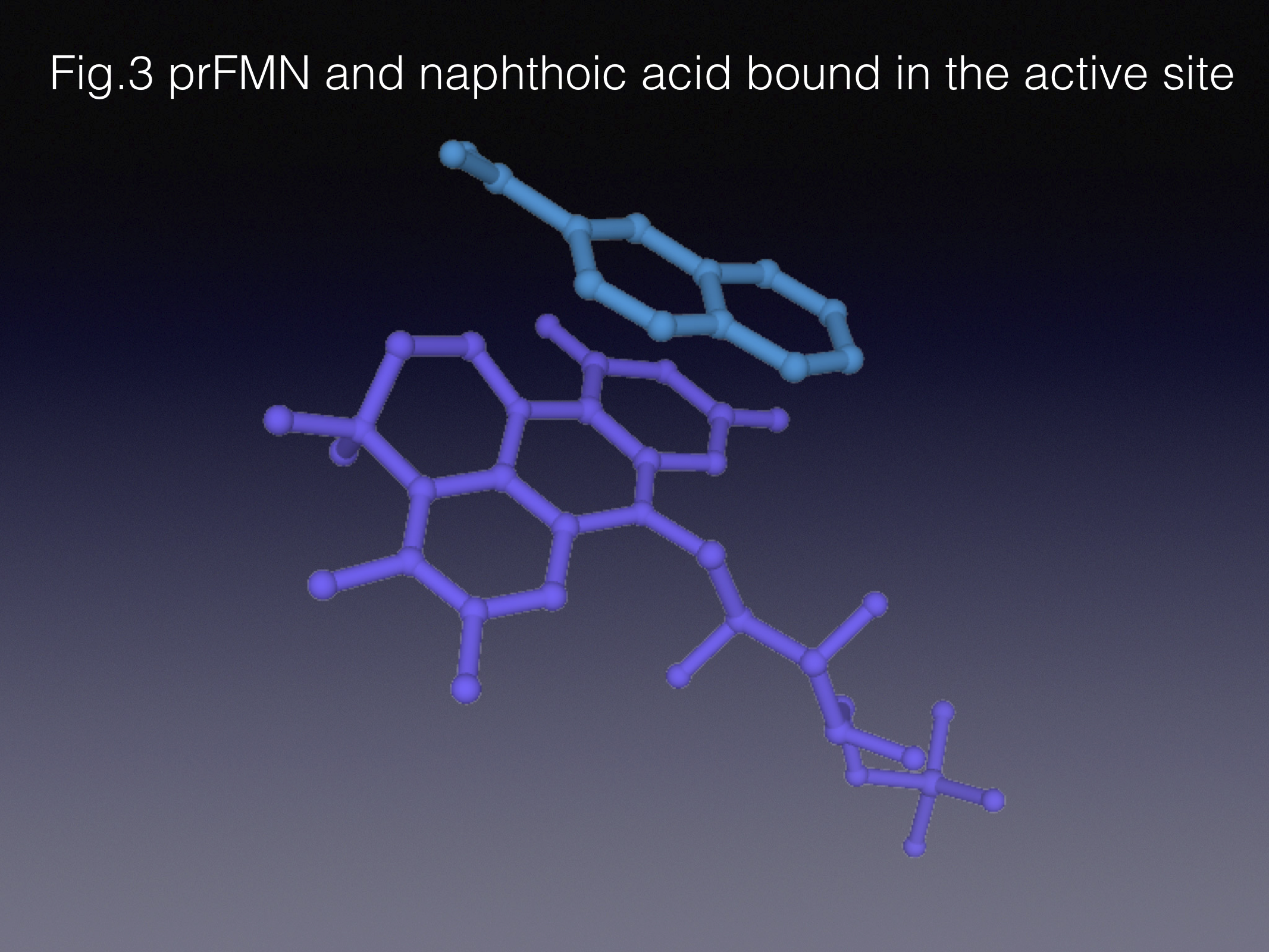
To conclude, this work was inspired by microbiological insights into aromatic degradation, conceived through mechanistic understanding of the key (de)carboxylase enzymes involved, enabled by the use of enzymatic cascades and structure-guided protein engineering techniques, and supported by sophisticated computational studies. Here’s the link to the paper if you are interested in more detail!
Enzymatic C–H activation of aromatic compounds through CO2 fixation Godwin A. Aleku, Annica Saaret, Ruth T. Bradshaw-Allen, Sasha R. Derrington, Gabriel R. Titchiner, Irina Gostimskaya, Deepankar Gahloth, David A. Parker, Sam Hay & David Leys, Nat Chem Biol, Published 27 July 2020.
https://doi.org/10.1038/s41589-020-0603-0
References:
[1] K.A.P. Payne, M.D. White, K. Fisher, B. Khara, S.S. Bailey, D. Parker, N.J.W. Rattray, D.K. Trivedi, R. Goodacre, R. Beveridge, P. Barran, S.E.J. Rigby, N.S. Scrutton, S. Hay and D. Leys (2015). "New cofactor supports α,β-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition." Nature 522 (7557): 497-501.
[2] S.S. Bailey, K.A.P. Payne, A. Saaret, S.A. Marshall, I. Gostimskaya, I. Kosov, K. Fisher, S. Hay and D. Leys (2019). "Enzymatic control of cycloadduct conformation ensures reversible 1,3-dipolar cycloaddition in a prFMN-dependent decarboxylase." Nature Chemistry 11 (11): 1049-1057.
Follow the Topic
-
Nature Chemical Biology
An international monthly journal that provides a high-visibility forum for the chemical biology community, combining the scientific ideas and approaches of chemistry, biology and allied disciplines to understand and manipulate biological systems with molecular precision.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in