We are living on a hungry planet and securing food supply for the steadily increasing human population is a major challenge. Improving crop productivity is an important strategy to achieve this goal. The productivity of agricultural and horticultural crops is constrained by lack of nutrients, abiotic stressors as well as pests and diseases. Together, all these limitations substantially prevent the accomplishment of the full genetic potential of plants for growth and fitness. The use of pesticides and chemical fertilizers to alleviate restrictions on plant performance causes serious environmental problems, and thus innovative, environmentally friendly solutions need to be developed urgently. Plant growth and health depend to a large extent on soil microbes that are associated with plant roots. These microbes can supply the plant with nutrients, alleviate effects of abiotic stresses, and protect against pathogens and pests. Thus, such beneficial microorganisms may be essential allies to improve crop yields in a sustainable way.
Beneficial symbioses between eukaryotic and prokaryotic organisms have evolved multiple times across the eukaryotic domain. While mutualistic interactions between plants and their associated microbes are common in nature, direct evidence of the evolution of bacterial mutualism is scarce. Although plant-microbe interactions are essential for crop production and ecosystem functionality, little is known about the evolutionary dynamics of plant-microbe interactions in the rhizosphere. In particular, the plant rhizosphere is an open system, free diffusion of plant-derived resources inherently prone to the tragedy of the commons (Figure1).
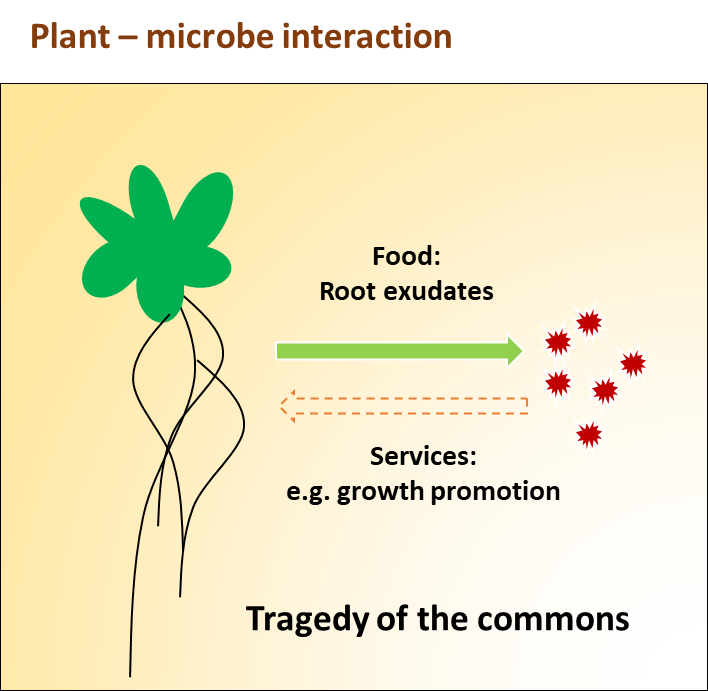
Figure 1. Typical beneficial plant- microbe interaction in the rhizosphere and tragedy of commons. The rhizosphere is a hotspot for interactions between the plant and free-living microorganisms. Plants deposit up to 40% of their photosynthetically fixed carbon into the rhizosphere as root exudates, which is supplied as the food sources of bacteria. Beneficial bacteria in turn, might provide services to their host plant, such as promoting plant growth. However, due to the open nature of the rhizosphere, free diffusion of plant-derived resources could select for increased levels of cheating where mutant bacterial genotypes take advantage of ‘commons’ without contributing to the production of plant growth-promoting compounds. Plants could exert selection against cheating by sanctioning cheating genotypes or positively selecting for mutualist bacterial genotypes.
Here we studied the evolutionary dynamics of plant-microbe interactions for the model bacterium Pseudomonas protegens CHA0 on the roots of model plant Arabidopsis thaliana, with an in vivo experimental evolution design. In our axenic system, bacterial survival and evolution was solely dependent on the presence of the plant-derived resources. After six plant growth cycles (6 months), the performance of evolved bacterial isolates was quantified in comparison with the ancestral bacterial strain. We found that initially plant antagonistic bacterium P. protegens rapidly diversified into five discrete phenotypes, that differed in their mutualism with the plant (Figure 2ac). Interestingly, mutualist genotypes became the dominant strain in 4 out 5 plant replicate lines only in about half year.
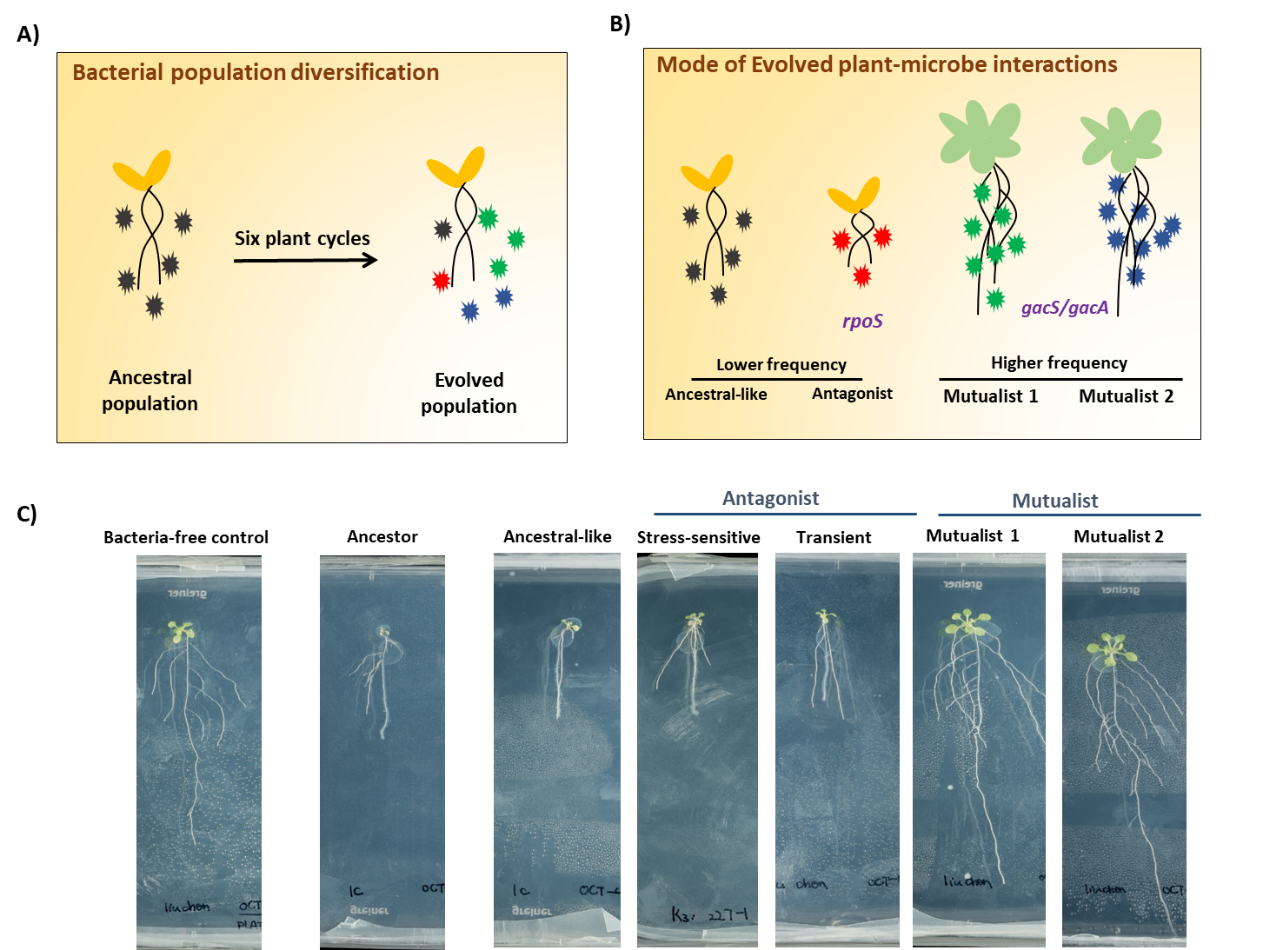
Figure 2. A) Diversification of bacterial population in the rhizosphere. In total, 240 evolved bacterial colonies were randomly selected at the end of the second, fourth and sixth growth cycles and characterized phenotypically by measuring multiple key life-history traits. B) Simplified mode for evolutionary changed plant-microbe interactions. C) Representative pictures of Arabidopsis performance with the effects of ancestral Pseudomonas protegens CHA0 and evolved phenotypes. 10-day-old seedlings (n=3 per side) were lined up in modified Hoagland agar in a two compartments petri dish. The left sides were treated with water (Control) or different CHA0 phenotypes (at root shoot junction). 14 days after digital pictures were taken.
We found two mechanisms that could explain this evolutionary transition (Figure 3). First, mutualists showed improved ability to compete for root exudates, which likely increased their relative abundance in the rhizosphere. Second, mutualists became better at activating root-specific transcription factor gene MYB72, leading to increased excretion of scopoletin antibacterial, which was relatively more harmful to non-mutualist genotypes. Mechanistically, these adaptations were achieved by key mutations in GacS/GacA two-component global regulator system, which is known to affect bacterial carbon metabolism, host-microbe signaling, and downregulation of phytotoxic secondary metabolites. Crucially, these mutations conferred clear fitness benefits only in the presence of plants, which suggests they were plant-specific adaptations.
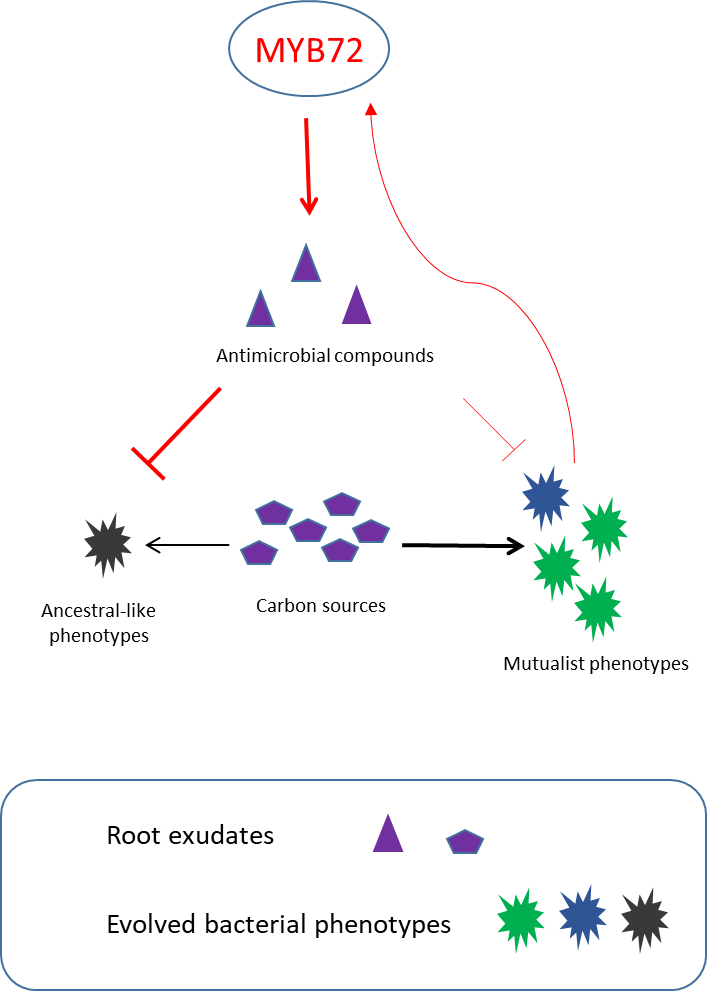
Figure 3. The schematic mechanisms of evolutionary mutualistic transition of bacterial population.
Together, these results show that bacterial mutualism can rapidly evolve in the plant rhizosphere at an agriculturally relevant evolutionary timescale, and that mutations in a single regulatory gene can drive considerable shifts in plant-bacterium interactions. Importantly, our results show that in addition to recruiting beneficial bacteria from multi-species microbial communities, plants can also domesticate their microbiota by imposing strong selection for the evolution of mutualistic bacterial genotypes. We believe that our results will be of broad interest to plant biologists, ecologist, evolutionary biologists, and ever-increasing group of researchers working with host-microbiome interactions across various biological systems. Moreover, due to translational potential, our results will likely interest agri-biotechnology companies as steering bacterial evolution in the rhizosphere could offer a short cut to improve crop yields.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in