Rapid prototyping and validation of SARS-CoV-2 clinical diagnostic workflows
Published in Bioengineering & Biotechnology
I woke up on the 29th of January in San Francisco to an email forwarded by my PI, Professor Paul Freemont. I'd almost come to the end of two weeks of training with the Riffyn Science Team and the Global Biofoundry Alliance was trying to coordinate a response to what had quickly become a massive public health threat. With many researchers taking any opportunity to jump on the bandwagon I was at first sceptical. My focus has always been on the translation of research and direct application. Was this just another attempt at publicity, or was this an honest attempt to make a tangible real world impact? With a tongue in cheek response of "a little bit of both" from Paul, we set about developing a plan to help our Chinese colleagues scale up their testing capacity.
My colleagues and I work in the London Biofoundry, a state-of-the-art facility that offers automated high-throughput laboratory equipment to prototype, engineer and validate biological constructs. This equipment is complemented by local expertise in synthetic biology, laboratory automation and programming to enable "full stack" development. And this unique environment provides the perfect platform for agile development, with the relevant expertise and equipment available at short notice for urgent projects.
This was just such an emergency and we mobilised our expertise and focused directly on the problem at hand. However, what had initially been an effort focused on helping our Chinese colleagues quickly morphed into an urgent need to scale up testing capacity in the UK. This came with its own unique challenges because the places where the diagnostic capacity was urgently needed (in hospital diagnostic labs) were somewhat impenetrable to "outsiders" such as ourselves. Diagnostic labs are reticent to spend time validating new diagnostic workflows unless there has already been some form of comprehensive laboratory validation.
However, we had been working on a "secret weapon" for just this purpose. Many viruses and bacteria require BSL2 and BSL3 facilities for the safe handling of clinical samples. This would usually limit diagnostic development efforts to using in vitro transcribed RNA or plasmid targets. These may be suitable for the validation of qPCR workflows, but they are poor process controls for the RNA extraction process which relies on the lysis of viral particles and inactivation of RNases in complex sample matrices. We had previously shown that we could produce MS2 virus-like particles packaged with RNA in-house. These virus-like particles have been shown to be very stable in a variety of clinical matrices and require viral lysis to release the corresponding RNA.
By early March we had a ddPCR quantified MS2 virus-like particle standard packaged with the RNA encoding for the Nucleocapsid Phosphoprotein from SARS-CoV-2, the target for the Center for Disease Control and Prevention's published RT-qPCR Assay. Given the number of reports around reagent shortages we knew that an integral part of this project would be securing a robust supply chain, otherwise there would be no real world impact (or publicity!). In our paper we showed that different qPCR mixes could be used and gave comparable results.
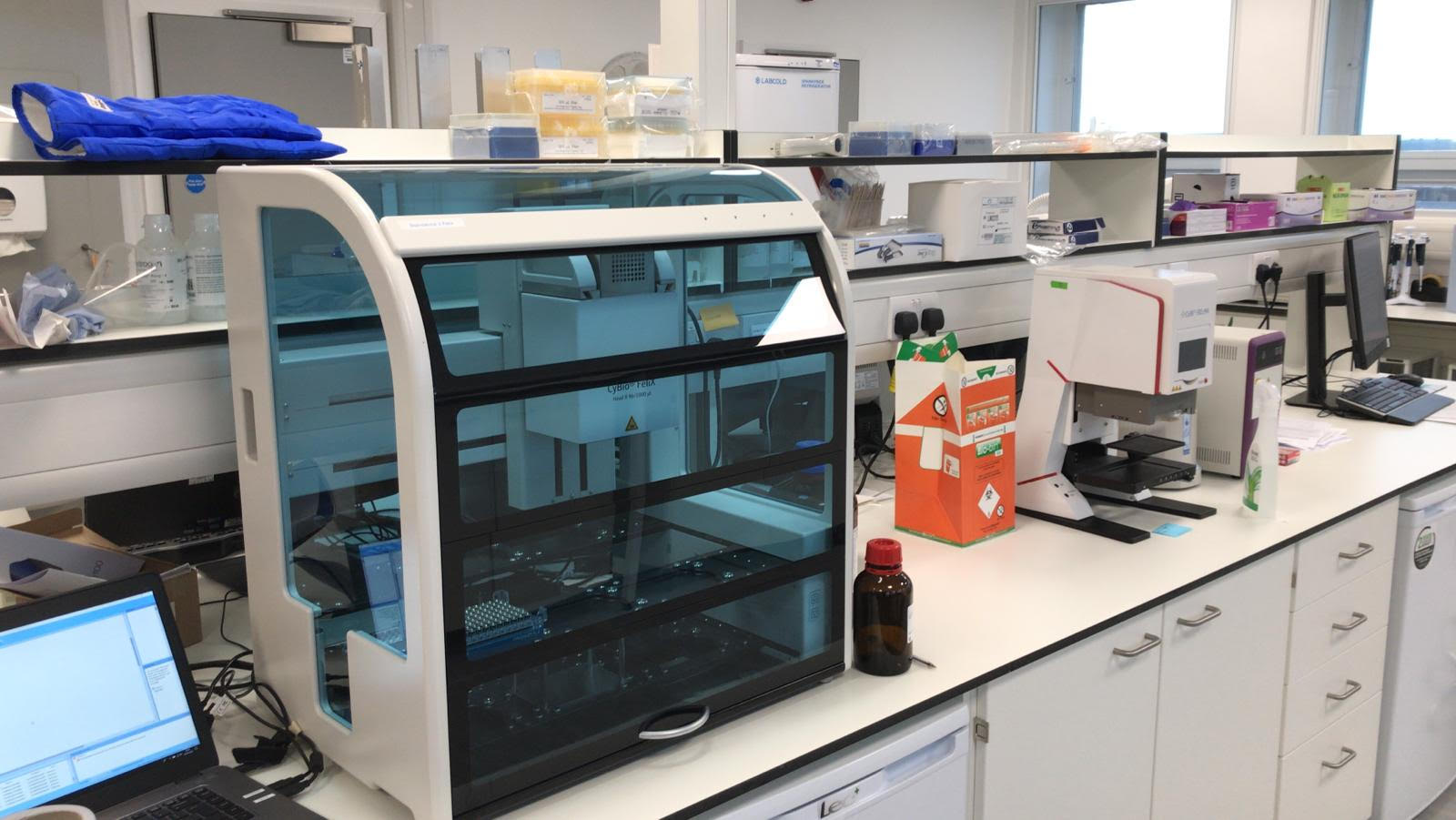
However, RNA extraction had always been the reagent-limiting step and therefore our focus had always been on developing a reagent and plastic consumable agnostic RNA extraction platform. Our liquid handling capabilities in the London Biofoundry are focused around the very flexible Analytik Jena CyBio FeliX. Its compact design made it the perfect platform to be repurposed for RNA extraction due to space limitations in diagnostic labs and its versatile nature would mean that it could be used for other assays once SARS-CoV-2 testing was no longer a focus. In our paper we used our virus-like particles to validate the FeliX as an RNA extraction platform with multiple extraction kits from different suppliers.
And then, we came to the true test. Having built credibility with our virus-like particle validated RNA extraction and qPCR workflows we approached our colleagues at the Imperial College Molecular Diagnostics Unit and Imperial College Healthcare NHS Trust (North West London Pathology). We show in our paper that both kits performed almost equally well when extracting patient samples (even though the Promega kit does not use carrier RNA). This was integral because early on we could no longer source one of our extraction kits (the Analytik Jena innuPREP Virus DNA/RNA Kit) and our approach served as a blueprint to validate other kits from other suppliers.
And that was it, from ordering the DNA for the virus-like particles on the 30th of January, to live patient samples on the 6th of April. We were the "enthusiastic amateurs" that developed a diagnostic workflow from scratch and made it to frontline diagnostic testing in just 9 weeks. And, importantly, we've shown that this platform is truly scalable, achieving 644 samples in 10 hours with just a single operator (in contrast to many other quoted theoretical throughputs or extreme staffing requirements). We currently have two platforms (at two separate sites) performing diagnostic testing, with one performing Imperial College student and staff testing. With these workflows (including our Quality Management System) currently the only custom automated workflows approved for diagnostic testing by the UK Accreditation Service (UKAS) our focus shifted towards validating other high-throughput techniques such as CRISPR and LAMP. The Riffyn Nexus platform was utilised throughout for the tracking of experiments, generation of laboratory automation instruction sets and analysis of data. The ability to track randomised experiments was vital to ensure assay robustness and in our paper we successfully implemented high-throughput CRISPR and LAMP assays and we showed that these assays could be successfully miniaturised for sample testing with a particular focus on potential community testing.

Looking to the future we are now focused on expanding, improving and sharing our approach. Firstly, both our NHS diagnostic platform and Imperial Testing platform are being expanded to 3000 samples a day (4 platforms) and 4000 samples a day (5 platforms) respectively. However, in order to increase capacity further, we are utilising the unparalleled sensitivity of RNA extraction and qPCR workflows to enable sample pooling (where lower sensitivity may preclude pooling with extraction free/LAMP workflows). We are therefore working together with Riffyn and our NHS colleagues to implement and validate a pooling strategy that can be implemented when prevalence rates are low and to help to scale up staff and student testing. We are also actively approaching Low-and Middle-income Countries (LMICs) to help them implement our reagent agnostic platform and our RNA extraction protocols will be made available on the London Biofoundry Github.
In the years to come pandemic preparedness will be an integral focus of governments around the world. We see machines such as the FeliX being an integral part of pandemic resilience strategies because they can form the basis of everyday chemical and biological research, but be repurposed as needed in pandemic scenarios. The London Biofoundry showed that this could be done from scratch in 9 weeks, but, with validated protocols and machinery on hand, in the future we will be able to implement a proactive rather than reactive approach, with a focus on disease containment rather than controlled spread.
This work would not have been possible without my colleagues in the London Biofoundry (Miles Priestman, Marta Ciechonska, Marko Storch and Kirsten Jensen), our project management support (Andrew Griffiths) and funding from the UK Dementia Research Institute, and our North West London Pathology colleagues (Paul Randell, Arthi Anand and Panos Pantelidis).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in