Sailing the high seas of structural genomic variation – A tale from the crow’s nest (literally)
Published in Ecology & Evolution

The human genome project is sometimes called the moonshot of biology, and for a good reason. For the first time in history, the very biological information that makes us human was (more or less) identified. And while before that scientists had obviously made groundbreaking discoveries related to our genome, one could say these studies were resembling first trips to a neighboring village. The technology that came with the human genome project initiated the era of genomic explorers, sailing into the vastness of the genome.
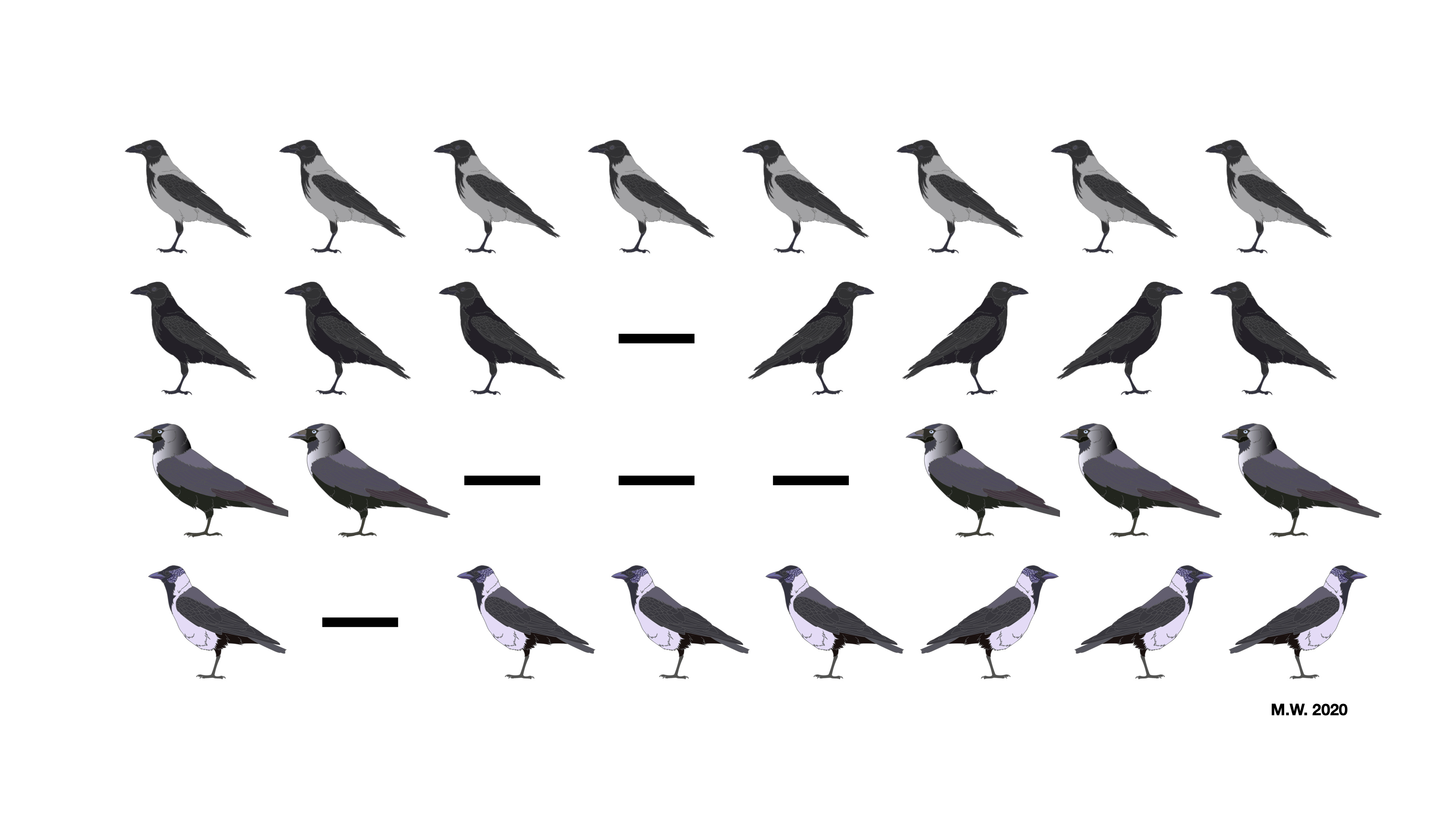
Fast forward about 15 years, and I, a motivated and naïve graduate student, find myself in the next phase of this era: the emergence of long-read sequencing. While I was still learning the basics of speciation and population genetics, I at the same time tried to keep up with the rapid development of new technologies that enabled ever more detailed investigations of the world consisting of A, C, T, and G. In the lab of my PhD advisor Jochen Wolf, endeavors to unravel the mysteries of crow speciation relied on studying single-nucleotide changes (SNPs) thus far, while already giving hints towards the importance of larger rearrangements – a.k.a. structural variation (SV; Poelstra et al. 2014, Vijay et al. 2016, Knief et al. 2019). And soon enough I found myself wrestling with the bioinformatic challenges of a population-scale data set of considerable size, consisting of short-read, long-read and optical mapping data to survey structural variation in this non-model system.
The main plan of the project that followed was actually not that different from the intentions of early explorers: set out, map, measure and describe all things encountered and hopefully bring light into the darkness. This metaphor makes it sound more romantic than it actually was. Wading through enormous amounts of data, I tried to yield sensible results, and one of the main lessons I learned along the way was to never trust initial results right away. In this case that meant staring at tons of alignments on the computer screen to make sure that variants spit out by software were actually real and trustworthy. As one can imagine, this does not scale well, and particularly the reliable genotyping of SV had proven to be difficult. Owing to the wide range of sampling in our data, comprising divergent lineages of the songbird genus Corvus, we could develop a filtering approach: Assuming that after 14 million years of divergence genetic variation was fully sorted, we removed variants that were segregating between those clades. Now, this seems like an overly rough approach, especially considering that SV commonly does not behave like single-nucleotide changes do in a population genetic sense, but given the lack of sophisticated genotyping models for SV, this was our best bet. Judging from the degree of concordance with SNP-inferred population structure analysis, our approach probably was not that far off in the end.
Probably the most exciting result of our study will become yet another example of how repetitive DNA – in this case an LTR retrotransposon – should be considered in all sorts of evolutionary studies. With the help of my co-advisor Alexander Suh, who is an expert for all things jumping and repetitive in the genome, we found out that this mobile genetic element inserted itself nearby a gene called NDP at some point during the evolution of the European crow lineage. In black-and-grey hooded crows the insertion is fixed and associated with reduced expression of NDP in skin tissue, which, together with the results of re-analyzing hybrid individuals, suggests that this structural variant indeed plays a role in maintaining plumage differences of black and pied crow populations.
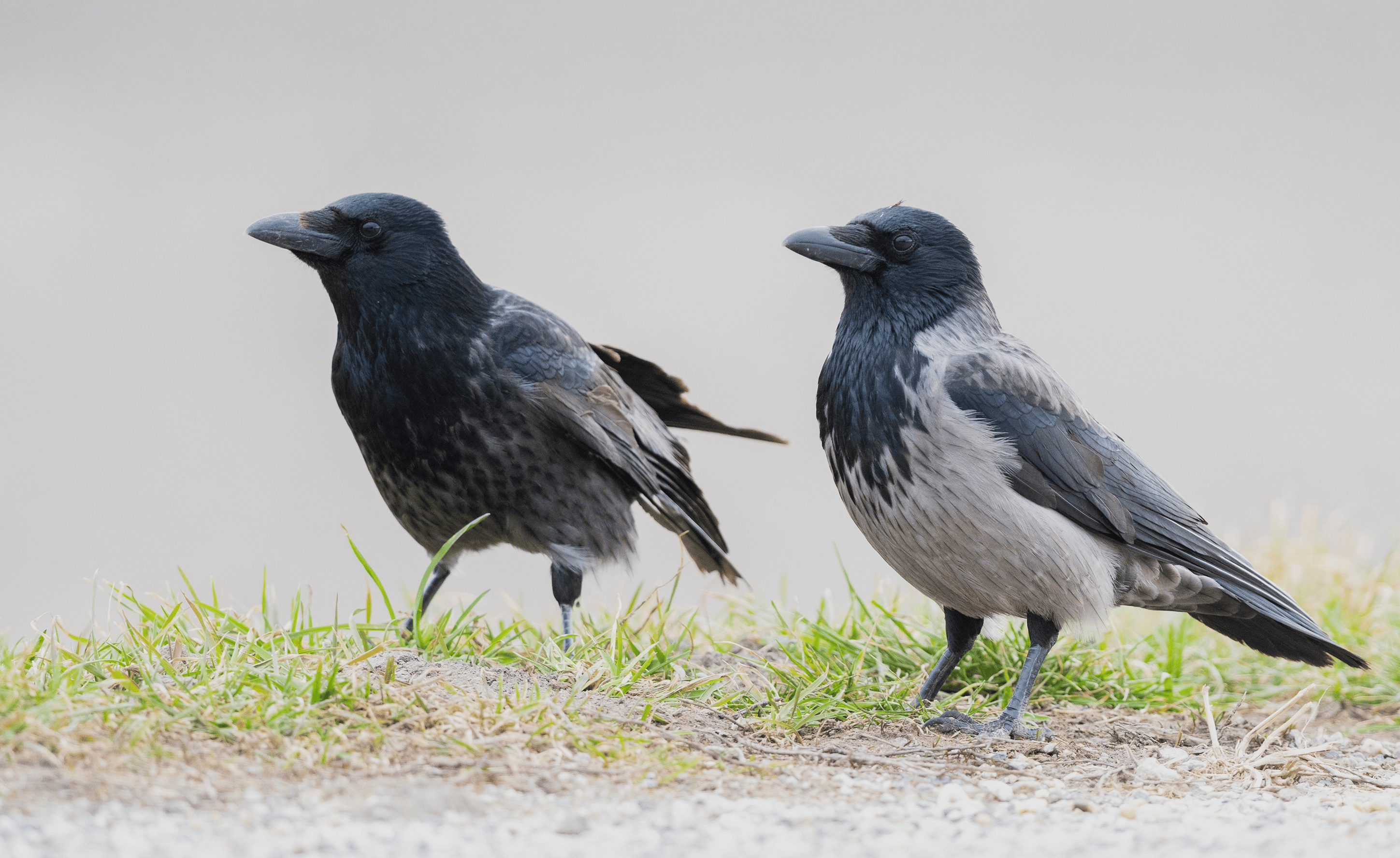

It may well be that in some years from now, at least parts of this study will be smirked at, however I hope that just like the work of early explorers, it may shed some light into uncharted corners of the genomics world. To draw your own conclusion, please have a look at the manuscript:
https://www.nature.com/articles/s41467-020-17195-4
Bibliography
Knief U, Bossu CM, Saino N et al. (2019) Epistatic mutations under divergent selection govern phenotypic variation in the crow hybrid zone. Nature Ecology & Evolution, 3, 570–576.
Poelstra JW, Vijay N, Bossu CM et al. (2014) The genomic landscape underlying phenotypic integrity in the face of gene flow in crows. Science (New York, N.Y.), 344, 1410–1414.
Vijay N, Bossu CM, Poelstra JW et al. (2016) Evolution of heterogeneous genome differentiation across multiple contact zones in a crow species complex. Nature Communications, 7, 13195.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in