Spatial Heterogeneity in Molecular Brightness
Published in Protocols & Methods
Explore the Research
Spatial heterogeneity in molecular brightness - Nature Methods
Nature Methods - Spatial heterogeneity in molecular brightness
Notably, in the field of G Protein Coupled Receptors (GPCRs), one of the most important superfamilies of membrane receptors, the debate had been ongoing over the last 20 years. On the wave of new fluorescence-based methods to assess molecular interactions or proximity, the field has exploded, often with contradictory results. A new influx of observations has been fostered by the development of super-resolution methods starting in 2006, and since then new methods are appearing by the month. For certain well studied receptors there is now an equal number of publications supporting the existence of dimers, against an equal number indicating their monomeric state.
How to reconcile these conflicting observations? Image based methods to assess oligomerisation (i.e. approaches where one actually collects an image of the cell while extracting quantitative data on the oligomerisation state of fluorescently labeled receptors) offer the advantage of introducing spatial selectivity in the measurements. While the common assumption is that all protein of interest already are localized at the plasma membrane, this is often not the case, and the localization of the receptors within the cell plays an important role in influencing the apparent oligomerization state.
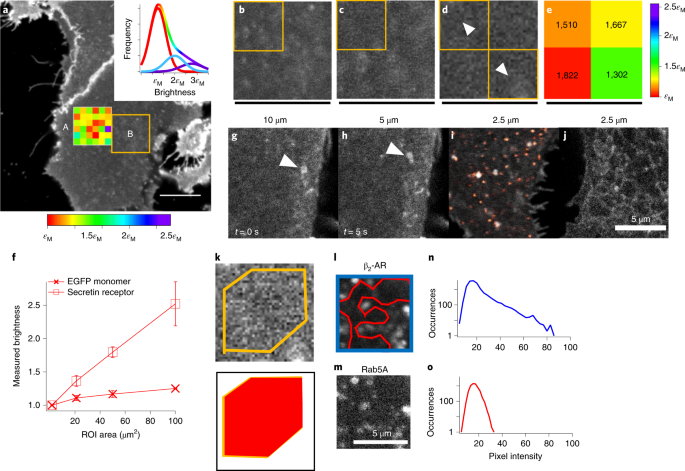
In our contribution we employ an image-based microscopy method, based upon molecular brightness analysis, to investigate the role of spatial heterogeneity in the quantification of GPCR oligomerisation, and we demonstrate that when spatial heterogeneity is properly taken into account, the resulting information can change drastically.
How do we do that? When looking down below from a hot air balloon at motion of a few sparse hikers in a field one can observe them move, form groups, separate.
When the persons are more crowded, e.g. a concert with a large public is taking place in the field, distinguishing the behavior of individual persons becomes much more difficult.
Along this analogy, extracting the behavior of a large number of fluorescently labeled molecules within a cell is problematic, due to inability of resolving them individually. Over the years, mostly thanks to the technical ability to precisely quantitate the number of photons emitted by a fluorescent species, statistical methods have emerged which allow recovering dynamic and kinetic information also from crowded ensembles of molecules.
In particular, so-called molecular brightness methods allow extracting the average number and aggregation state (oligomerization) of molecules present on a certain region of a cell, such as the plasma membrane, by calculating the variance of the photon counts collected from the sample.
In May 2019 we became aware of the work from a research group based in the USA, who was advising an automatic selection algorithm (high-throughput) to perform this type of analysis. We felt the method led to some surprising conclusions, and we challenged some of its assumptions. We further reasoned that by combining two methods to measure the same process (e.g. fluctuations in space as well as in time), one could hope to gain a more insightful and truthful outcome. Our thoughts, are summarized in the linked "Matters Arising" piece!
Follow the Topic
-
Nature Methods
This journal is a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Methods development in Cryo-ET and in situ structural determination
Publishing Model: Hybrid
Deadline: Jul 28, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in