Towards providing physicians with a quantitative tool for optimizing immunotherapy treatment protocols for each individual patient
Published in Bioengineering & Biotechnology
Immunotherapy has begun to make a transformative impact on oncology practice in recent years. However, physicians often find it difficult to predict if immunotherapy drugs will have an eventual effect for a patient or determine how long to keep the patient on immunotherapy drugs. One reason for this problem is the lack of biomarkers—measurable indicators that provide reliable assessment of how well the drug will work or is working—that can provide clear, quantitative signals to physicians as early as possible. Despite significant advances in many research and technological areas, there are still no reliable, broadly capable biomarkers in current clinical practice for determining patient-specific immunotherapy treatment outcome, as to date most biomarkers are often specific to a certain family of drug and disease combinations. In fact, providing reliable biomarkers of immunotherapy response that may be broadly applied across many cancer and treatment types remains an ongoing challenge to frontline researchers.
Reliable, broadly applicable biomarkers of immunotherapy response may provide timely identification of the most effective immunotherapy treatment protocol for an individual patient, which would allow physicians to select or fine-tune the treatment strategy to maximize its effectiveness. This is the fundamental rational behind our development of a mechanistic mathematical model for describing patient tumor burden over time under checkpoint inhibitor-type immunotherapy treatment [1]. We sought to mathematically describe the specific tumor-immune interaction processes or mechanisms underlying immunotherapy intervention as a function of specific, relevant physical and biological processes in order to provide a tool that was able to predict the therapeutic response, as we have done for numerous other cancer and therapy types [2-9]. We have now been successful in developing such a model [1], which is comprised of a set of mathematical equations describing key processes or mechanisms in immunotherapy treatment with checkpoint inhibitors, for example, the kill rate of cancer cells by immune cells (μ), and a measure of the immune state within the tumor under immunotherapy intervention (Λ).
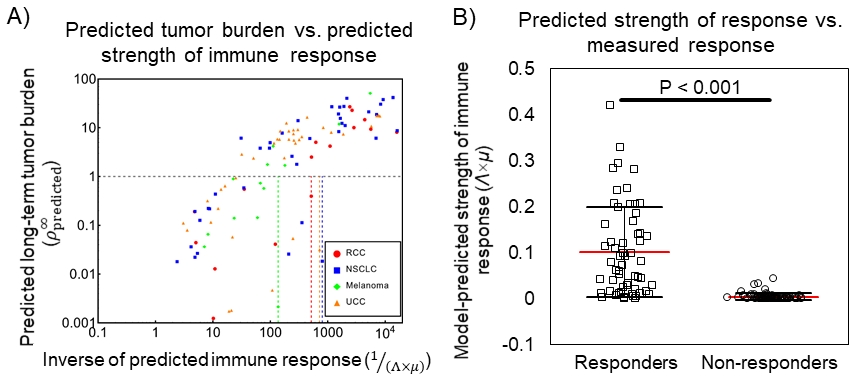
We applied the model to data from 124 patients who received checkpoint inhibitor treatment in order to test its performance in quantifying the strength of immune response. These patients had either renal cell carcinoma (RCC), non-small cell lung cancer (NSCLC), melanoma, or urothelial carcinoma (UCC), and were treated with one of the checkpoint inhibitors nivolumab or atezolizumab, thus representing a diverse sampling of cancer and drug types. The model was also used to predict the strength of treatment response for each drug-cancer type pairing examined. These results were then further validated in 177 additional patients from our in-house clinical trials. Briefly, we observed that the strength of immune response predicted by the model was significantly correlated (P < 0.001) with predicted long-term tumor burden (Figure 1A), which we found to be a reliable predictor of the tumor burden that was measured at last patient follow-up. This allowed us to stratify patients based on their model-predicted clinical response, where we defined responders to have reduced tumor burden at last follow-up, while non-responders had increased tumor burden despite receiving checkpoint inhibitor immunotherapy (Figure 1B). Furthermore, we found that the model predicted patient response with accuracy ranging 77–100%, depending on the drug-tumor combination examined. Using our model, we designed an overarching measure of the strength of immune response, which we calculated by weighting the tumor kill rate by the immune state within the tumor, calculated as Λ×μ.
This mechanistic modeling work is a result of integrating the strengths of the diverse fields of mathematical modeling, biology, immunology, and clinical analysis. It provides physicians a predictive tool to estimate patient response early after the start of treatment (for example, 1 month after a patient receives their first treatment), allowing them to refine or adjust treatment strategies to maximize the therapeutic benefit the patient receives. The model can also be used to calculate the specific mechanism(s) causing unsuccessful checkpoint inhibitor immunotherapy intervention, allowing physicians to identify supplemental treatments that may overcome this bottleneck, potentially enabling patients to be transitioned from unfavorable to favorable response categories. In this ongoing project, we are now testing the model in additional cancer-drug combinations to determine if it may be more broadly applied for improved design of treatment strategies based on an individual patient’s needs. We are particularly enthusiastic about the model’s clinical and translational applications as it relies only on standard-of-care imaging measurements as inputs and can thus be implemented directly into current clinical practice.
References
1. Butner, J. D. et al. A mechanistic immunotherapy model provides patient-specific quantification of immune response and associated long-term tumor burden. Nature Biomedical Engineering. 2020. https://www.nature.com/articles/s41551-020-00662-0
2. Cristini, V., Koay, E. & Wang, Z. An Introduction to Physical Oncology: How Mechanistic Mathematical Modeling Can Improve Cancer Therapy Outcomes. (CRC Press, 2017).
3. Brocato, T. A. et al. Predicting breast cancer response to neoadjuvant chemotherapy based on tumor vascular features in needle biopsies. JCI insight 5, doi:10.1172/jci.insight.126518 (2019).
4. Butner, J. D. et al. Mathematical prediction of clinical outcomes in advanced cancer patients treated with checkpoint inhibitor immunotherapy. Science advances 6, eaay6298, doi:10.1126/sciadv.aay6298 (2020).
5. Butner, J. D. et al. A Multiscale Agent-Based Model of Ductal Carcinoma In Situ. IEEE transactions on bio-medical engineering 67, 1450-1461, doi:10.1109/TBME.2019.2938485 (2020).
6. Dogra, P. et al. Establishing the effects of mesoporous silica nanoparticle properties on in vivo disposition using imaging-based pharmacokinetics. Nature communications 9, 4551, doi:10.1038/s41467-018-06730-z (2018).
7. Dogra, P. et al. A mathematical model to predict nanomedicine pharmacokinetics and tumor delivery. Computational and structural biotechnology journal 18, 518-531, doi:10.1016/j.csbj.2020.02.014 (2020).
8. Goel, S. et al. Size-Optimized Ultrasmall Porous Silica Nanoparticles Depict Vasculature-Based Differential Targeting in Triple Negative Breast Cancer. Small 15, e1903747, doi:10.1002/smll.201903747 (2019).
9. Wang, Z. et al. Theory and Experimental Validation of a Spatio-temporal Model of Chemotherapy Transport to Enhance Tumor Cell Kill. PLoS Comput Biol 12, e1004969, doi:10.1371/journal.pcbi.1004969 (2016).
Follow the Topic
-
Nature Biomedical Engineering
This journal aspires to become the most prominent publishing venue in biomedical engineering by bringing together the most important advances in the discipline, enhancing their visibility, and providing overviews of the state of the art in each field.
Related Collections
With Collections, you can get published faster and increase your visibility.
Implantable wireless communication technologies
Publishing Model: Hybrid
Deadline: Nov 28, 2026
Medical Ultrasound: Emerging Techniques and Applications
Publishing Model: Hybrid
Deadline: Jan 29, 2027



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in